Abstract
Background
GDP-D-mannose pyrophosphorylase (GMP) is one of the key enzymes determining ascorbic acid (AsA) biosynthesis. However, little information about GMP genes is currently available for the Rosaceae species, especially in the AsA-riched cultivated octoploid strawberry (Fragaria × ananassa).
Objective
To identify the all the GMP genes in Rosaceae, as well as the predominant homologues and the role of GMP genes in strawberry AsA accumulation.
Methods
In the present study, we performed genome-wide identification and comprehensive analysis of the duplicated GMP genes in strawberry and other Rosaceae species by bioinformatics methods, the expression of the GMP genes from cultivated strawberry (Fragaria × ananassa, FaGMP) was specifically analyzed by qPCR. Finally, the FaGMP4 was transiently overexpressed in strawberry to estimate the role of GMP in regulating AsA accumulation in strawberry.
Results
As results, a total of 28 GMP genes were identified in the five Rosaceae species. The origins of duplication events analysis suggested that most GMP duplications in Rosaceae species were generated from whole genome duplication (WGD). The Ka/Ks ratio suggested that FaGMP genes underwent a stabilization selection. qPCR based expression analysis showed different patterns of FaGMP paralogs during fruit ripening, while FaGMP4 expressed higher in the variety containing higher AsA. Overexpression of FaGMP4 in strawberry significantly enhanced AsA accumulation. Furthermore, the expression of FaGMP4 under the treatment of blue and red light was largely increased in leaves while significantly inhibited in fruit. These results revealed the vital role of FaGMP4 in regulating AsA in strawberry.






Similar content being viewed by others
References
Agius F, González-Lamothe R, Caballero JL, Muñoz-Blanco J, Botella MA, Valpuesta V (2003) Engineering increased vitamin C levels in plants by overexpression of a D-galacturonic acid reductase. Nat Biotechnol 21(2):177–181
Aron MB, Shennan L, Anderson JB, Farideh C, Derbyshire MK, Carol DWS, Fong JH, Geer LY, Geer RC, Gonzales NR (2011) CDD: a conserved domain database for the functional annotation of proteins. Nucleic Acids Res 39(Database issue):225–229
Badejo AA, Jeong ST, Goto-Yamamoto N, Esaka M (2007) Cloning and expression of GDP-D-mannose pyrophosphorylase gene and ascorbic acid content of acerola (Malpighia glabra L.) fruit at ripening stages. Plant Physiol Biochem 45(9):665–672
Badejo A, Tanaka N, Esaka M (2008) Analysis of GDP-D-mannose pyrophosphorylase gene promoter from acerola (Malpighia glabra) and increase in ascorbate content of transgenic tobacco expressing the acerola gene. Plant Cell Physiol 49(1):126–132
Blaschke K, Ebata KT, Karimi MM, Zepedamartínez JA, Goyal P, Mahapatra S, Tam A, Laird DJ, Hirst M, Rao A (2013) Vitamin C induces Tet-dependent DNA demethylation and a blastocyst-like state in ES cells. Nature 500(7461):222–226
Bulley SM, Rassam M, Hoser D, Otto W, Schünemann N, Wright M, Macrae E, Gleave A, Laing W (2009) Gene expression studies in kiwifruit and gene over-expression in Arabidopsis indicates that GDP-L-galactose guanyltransferase is a major control point of vitamin C biosynthesis. J Exp Bot 60(3):765–778
Chen Q, Yu H, Wang X, Xie X, Yue X, Tang H (2012) An alternative cetyltrimethylammonium bromide-based protocol for RNA isolation from blackberry (Rubus L.). Genet Mol Res 11(2):1773–1782
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13(8):1194–1202
Clarke SG (2008) L-Ascorbate biosynthesis in higher plants: the role of VTC2. Trends Plant Sci 13(11):567–573
Conklin PL, Williams EH, Last RL (1996) Environmental stress sensitivity of an ascorbic acid-deficient Arabidopsis mutant. Proc Nat Acad Sci USA 93(18):9970–9974
Conklin PL, Norris SR, Wheeler GL, Williams EH, Smirnoff N, Last RL (1999) Genetic evidence for the role of GDP-mannose in plant ascorbic acid (vitamin C) biosynthesis. Proc Natl Acad Sci USA 96(7):4198–4203
Conklin PL, Saracco SA, Norris SR, Last RL (2000) Identification of ascorbic acid-deficient Arabidopsis thaliana mutants. Genet 154(2):847–856
Cruz-Rus E, Amaya I, Sánchez-Sevilla JF, Botella MA, Valpuesta V (2011) Regulation of L-ascorbic acid content in strawberry fruits. J Exp Bot 62(12):4191–4201
Davey MW, Montagu MV, Inzé D, Sanmartin M, Kanellis A, Smirnoff N, a x JI, Benzie J, Strain JJ, Favell D, Fletcher J (2000) Plant L-ascorbic acid: chemistry, function, metabolism, bioavailability and effects of processing. J Sci Food Agric 80(7):825–860
Edger PP, Poorten TJ, VanBuren R et al (2019) Origin and evolution of the octoploid strawberry genome. Nat Genet 51(3):541–547
Emms DM, Kelly S (2019) OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol 20(1):238
Finn RD, Coggill P, Eberhardt RY, Eddy SR, Mistry J, Mitchell AL, Potter SC, Punta M, Qureshi M, Sangradorvegas A (2016) The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res 44(Database issue):D279–D285
Fukunaga K, Fujikawa Y, Esaka M (2010) Light Regulation of Ascorbic Acid Biosynthesis in Rice via Light Responsive cis-Elements in Genes Encoding Ascorbic Acid Biosynthetic Enzymes. J Agric Chem Soci Jpn 74(4):888–891
Imai T, Ban Y, Terakami S, Yamamoto T, Moriguchi T (2009) L-Ascorbate biosynthesis in peach: cloning of six L-galactose pathway-related genes and their expression during peach fruit development. Physiol Plant 136(2):139–149
Ioannidi E, Kalamaki MSE, Pateraki C, Alexandrou I, Mellidou D, Giovannonni I, Kanellis J AK (2009) Expression profiling of ascorbic acid-related genes during tomato fruit development and ripening and in response to stress conditions. J Exp Bot 60(2):663–678
Jiang ZY, Zhong Y, Zheng J, Ali M, Liu GD, Zheng XL (2018) L-ascorbic acid metabolism in an ascorbate-rich kiwifruit (Actinidia. Eriantha Benth.) cv. ‘White’ during postharvest. Plant Physiol Biochem 12420–28
Keller R, Renz FS, Kossmann J (2010) Antisense inhibition of the GDP-mannose pyrophosphorylase reduces the ascorbate content in transgenic plants leading to developmental changes during senescence. Plant J 19(2):131–141
Koonin EV (2005) Orthologs, paralogs, and evolutionary genomics. Annu Rev Genet 39309–39338
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874
Li M, Chen X, Wang P, Ma F (2011) Ascorbic acid accumulation and expression of genes involved in its biosynthesis and recycling in developing apple fruit. J Am Soc Hortic Sci 136(4):231–238
Liu JH, Peng T, Dai W (2014) Critical cis-acting elements and interacting transcription factors: key players associated with abiotic stress responses in plants. Plant Mol Biol Rep 32(2):303–317
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2 –∆∆ C T method. Methods 25(4):402–408
Magadum S, Murugan P, Gangapur D, Ravikesavan R (2013) Gene duplication as a major force in evolution. J Genet 92(1):155–161
Nishikimi M, Fukuyama R, Minoshima S, Shimizu N, Yagi K (1994) Cloning and chromosomal mapping of the human nonfunctional gene for L-gulono-gamma-lactone oxidase, the enzyme for L-ascorbic acid biosynthesis missing in man. J Biol Chem 269(18):13685–13688
Panchy N, Lehti-Shiu M, Shiu SH (2016) Evolution of gene duplication in plants. Plant Physiol 171(4):2294–2316
Proulx SR (2012) Multiple routes to subfunctionalization and gene duplicate specialization. Genet 190(2):737–751
Qin H, Deng Z, Zhang C, Wang Y, Wang J, Liu H, Zhang Z, Huang R, Zhang Z (2016) Rice GDP-mannose pyrophosphorylase OsVTC1-1 and OsVTC1-3 play different roles in ascorbic acid synthesis. Plant Mol Biol 90(3):317–327
Qin H, Deng Z, Zhang C, Wang Y, Wang J, Liu H, Zhang Z, Huang R, Zhang Z (2021) Rice GDP-mannose pyrophosphorylase OsVTC1-1 and OsVTC1-3 play different roles in ascorbic acid synthesis. Plant Mol Biol 90(3):317–327
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4):406–425
Smirnoff N (2000) Ascorbic acid: metabolism and functions of a multi-facetted molecule. Curr Opin Plant Biol 3(3):229–235
Smirnoff N, Conklin PL, Loewus FA (2001) Biosynthesis of ascorbic acid in plants: a renaissance. Annu Rev Plant Physiol Plant Mol Biol 52(52):437–467
Sook J, Taein L, Chun-Huai C, Katheryn B, Ping Z, Jing Y, Jodi H, Ksenija PFS, Kristin G (2018) 15 years of GDR: new data and functionality in the genome database for Rosaceae. Nucleic Acids Res D1:D1137–D1145
Spolaore S, Trainotti L, Casadoro G (2001) A simple protocol for transient gene expression in ripe fleshy fruit mediated by Agrobacterium. J Exp Bot 52(357):845–850
Tabata K, Takaoka T, Esaka M (2002) Gene expression of ascorbic acid-related enzymes in tobacco. Phytochem 61(6):631–635
Velde FVD, Pirovani ME, Cámara MS, Güemes DR, Bernardi CMDH (2012) Optimization and validation of a UV–HPLC method for vitamin C determination in strawberries (Fragaria ananassa Duch.), using experimental designs. Food Analyt Methods 5(5):1097–1104
Wheeler GL, Jones MA, Smirnoff N (1998) The biosynthetic pathway of vitamin C in higher plants. Nature 393(6683):365–369
Xue CC, Jin-Yan XU, Wang C, Guo N, Hou JF, Dong X, Zhao JM, Han X (2018) Molecular cloning and functional characterization of a soybean GmG MP1 gene reveals its involvement in ascorbic acid biosynthesis and multiple abiotic stress tolerance in transgenic plants. J Integr Agric 17(3):539–553
Yabuta Y, Mieda T, Rapolu M, Nakamura A, Motoki T, Maruta T, Yoshimura K, Ishikawa T, Shigeoka S (2007) Light regulation of ascorbate biosynthesis is dependent on the photosynthetic electron transport chain but independent of sugars in Arabidopsis. J Exp Bot 58(10):2661–2671
Zhang Y (2013) Biological role of ascorbate in plants. In: Zhang Y (ed) Ascorbic acid in plants, biosynthesis, regulation and enhancement. Springer, New York, pp 7–33
Zhang Y (2013) Enzymes involved in ascorbate biosynthesis and metabolism in plants. Ascorbic acid in plants,biosynthesis, regulation and enhancement. Springer, New York, pp 57–86
Zhang Z, Li J, Zhao XQ, Wang J, Wong KS, Yu J (2006) KaKs_Calculator: calculating Ka and Ks throughmodel selection and model averaging. Genom Proteom Bioinform 4(4):259–263
Zhang Q, Chen W, Sun L et al (2012) The genome of Prunus mume. Nat Commun 31318 – 1318
Zhang C, Ouyang B, Yang C, Zhang X, Liu H, Zhang Y, Zhang J, Li H, Ye Z (2013) Reducing AsA leads to leaf lesion and defence response in knock-down of the AsA biosynthetic enzyme GDP-D-mannose pyrophosphorylase gene in tomato plant. Plos One 8(4):e61987
Acknowledgements
We would like to thank the Institute of Pomology and Olericulture in Sichuan Agricultural University for providing the HPLC system to determine the AsA content.
Author information
Authors and Affiliations
Contributions
Conceptualization, HT; Methodology, YL, JZ, and LW; Software, YL, ML and QC; Validation, YL, JZ, and LW; Formal analysis, YL; Data curation, YL; Writing—original draft preparation, YL; Writing—review and editing, YL, YZ, YZ, YW and XW; Supervision, HT.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Figure S2.
Multiple alignment of GMP amino acids sequences from Rosaceae and Arabidopsis. The secondary structure was constructed by the pattern of T. maritima GMP protein (accession: 2 × 5s in protein data bank). Red text represents the core for metal binding site. Black box indicates substrate binding sites (SBS); red box represents Hexapeptide repeat; blue box indicates transmembrane motif (TM).The graph above the sequences indicates the secondary structure of the proteins (pdf 560 KB)
Figure S2.
Multiple alignment of GMP amino acids sequences from Rosaceae and Arabidopsis. The secondary structure was constructed by the pattern of T. maritima GMP protein (accession: 2 × 5s in protein data bank). Red text represents the core for metal binding site. Black box indicates substrate binding sites (SBS); red box represents Hexapeptide repeat; blue box indicates transmembrane motif (TM).The graph above the sequences indicates the secondary structure of the proteins (pdf 2789 KB)
Figure S3.
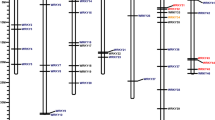
The chromosome to chromosome synteny relationship between Arabidopsis and five Rosaceae species. The location of GMP gene on the chromone was represented by red triangle. The black line indicated the collinear relationship between Arabidopsis and five other Rosaceae species. The red, yellow, blue and purple lines indicated the collinear relationship between octoploid strawberry and apple, peach, European pear, woodland strawberry respectively (pdf 370 KB)
Table S1
. The correspondence of gene ID and gene name in this study (xlsx 11 KB)
Table S2.
Collinear gene pairs of GMP in Arabidopsis and Rosaceae (xlsx 2641 KB)
Table S3.
The duplication types of GMP in Rosaceae (xlsx10 KB)
Table S4
. Information of motifs in the promoter region of FaGMP genes (xlsx 16 KB)
Table S5.
Primers used in this study (xlsx 10 KB)
Rights and permissions
About this article
Cite this article
Lin, Y., Zhang, J., Wu, L. et al. Genome-wide identification of GMP genes in Rosaceae and functional characterization of FaGMP4 in strawberry (Fragaria × ananassa). Genes Genom 43, 587–599 (2021). https://doi.org/10.1007/s13258-021-01062-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-021-01062-7