Abstract
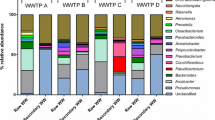
Industrial wastewater effluents present a major source of water pollution, and can potentially alter the microbial ecological landscape. While there are numerous reports on the microbial quality of domestic municipal effluents and their perceived environmental effects, there are limited reports devoted to the study of bacterial diversity of effluents from individual industries before they are mixed up with other sources. This study analyzed both the physicochemical parameters and bacterial community structures of different industrial wastewaters using Illumina high-throughput sequencing platform. Industrial wastewater with temperature ranging from 18.9 to 21.5 °C, and total dissolved solid (TDS) levels at up to 4611 mg/L, appeared to be predominated by Proteobacteria (44.44–75.86%) with the exception of the Capegate sample where Actinobacteria (39.66%) were the highest. Sulfur levels were significantly higher (p < 0.05) in Dixon wastewater constituting higher populations of sulfur reducing bacteria (SRB) compared to the other sites. Diversity index (Shannon-H index) and richness estimator (Chao1 index) ranged from 974 (Capegate) to 4552 (Dixon) and 6.04 (Dixon) to 4.15 (CWI), respectively. Multivariate analysis results highlighted that the bacterial communities were strongly shaped by physicochemical variables. The top 10 operational taxonomic units (OTUs) of each industrial sample had the potential to play important roles in the bioremediation and biodegradation of pollutants. Dominant OTUs belonging to the phyla Planctomyces from the Chemreem sample could not be classified to any genera and are likely to represent novel species.







Similar content being viewed by others
References
Abed RMM, Safi NMD, Köster J et al (2002) Microbial diversity of a heavily polluted microbial mat and its community changes following degradation of petroleum compounds microbial diversity of a heavily polluted microbial mat and its community changes following degradation of petroleum compounds. Appl Environ Microbiol 68:164–1683. https://doi.org/10.1128/AEM.68.4.1674
Abed RMM, Klempova T, Gajdos P, Certik M (2015) Bacterial diversity and fatty acid composition of hypersaline cyanobacterial mats from an inland desert wadi. J Arid Environ 115:81–89. https://doi.org/10.1016/j.jaridenv.2015.01.010
Ahmed W, Staley C, Sidhu J et al (2017) Amplicon-based profiling of bacteria in raw and secondary treated wastewater from treatment plants across Australia. Appl Microbiol Biotechnol 101:1253–1266. https://doi.org/10.1007/s00253-016-7959-9
Albertsen M, McIlroy SJ, Stokholm-Bjerregaard M et al (2016) “Candidatus Propionivibrio aalborgensis”: a novel glycogen accumulating organism abundant in full-scale enhanced biological phosphorus removal plants. Front Microbiol 7:1–17. https://doi.org/10.3389/fmicb.2016.01033
APHA (2001) Standard methods for the examination of water and wastewater, Washington DC
Ballesté E, Blanch AR (2011) Bifidobacterial diversity and the development of new microbial source tracking indicators. Appl Environ Microbiol 77:3518–3525. https://doi.org/10.1128/AEM.02198-10
Bassin JP, Rachid CTCC, Vilela C et al (2017) Revealing the bacterial profile of an anoxic-aerobic moving-bed biofilm reactor system treating a chemical industry wastewater. Int Biodeterior Biodegrad 120:152–160. https://doi.org/10.1016/j.ibiod.2017.01.036
Ben-Dov E, Shapiro OH, Gruber R et al (2008) Changes in microbial diversity in industrial wastewater evaporation ponds following artificial salination. FEMS Microbiol Ecol 66:437–446. https://doi.org/10.1111/j.1574-6941.2008.00549.x
Bronze MR, Vilas-Boas L, Catulo L, Peres C (2008) Use of lactobacillus plantarum in treatments of olive mill wastewater. Acta Hortic 791:637–644. https://doi.org/10.17660/ActaHortic.2008.791.97
Chen C, Khaleel SS, Huang H, Wu CH (2014) Software for pre-processing Illumina next-generation sequencing short read sequences. Source Code Biol Med 9:1–11. https://doi.org/10.1038/nbt1486
Dickman SR, Bray RH (1940) Colorimetric determination of phosphate. Ind Eng Chem Anal Ed 12:665–668. https://doi.org/10.1021/ac50151a013
Edgar RC, Haas BJ, Clemente JC et al (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Felföldi T, Székely AJ, Gorál R et al (2010) Polyphasic bacterial community analysis of an aerobic activated sludge removing phenols and thiocyanate from coke plant effluent. Bioresour Technol 101:3406–3414. https://doi.org/10.1016/j.biortech.2009.12.053
Ferrer M, Guazzaroni ME, Richter M et al (2011) Taxonomic and functional metagenomic profiling of the microbial community in the anoxic sediment of a sub-saline shallow lake (Laguna de Carrizo, Central Spain). Microb Ecol 62:824–837. https://doi.org/10.1007/s00248-011-9903-y
Ford T (1994) Pollutant effects on the microbial ecosystem. Environ Health Perspect 102:45–48
Gao P, Xu W, Sontag P et al (2016) Correlating microbial community compositions with environmental factors in activated sludge from four full-scale municipal wastewater treatment plants in Shanghai, China. Appl Microbiol Biotechnol 100:4663–4673. https://doi.org/10.1007/s00253-016-7307-0
Hammer Ø, Harper DAT, Ryan PD (2001) Paleontological statistics software package for education and data analysis. Palaeontol Electron 4:9–18. https://doi.org/10.1016/j.bcp.2008.05.025
Hülsen T, Hsieh K, Lu Y et al (2018) Simultaneous treatment and single cell protein production from agri-industrial wastewaters using purple phototrophic bacteria or microalgae—a comparison. Bioresour Technol 254:214–223. https://doi.org/10.1016/j.biortech.2018.01.032
Ibarbalz FM, Figuerola ELM, Erijman L (2013) Industrial activated sludge exhibit unique bacterial community composition at high taxonomic ranks. Water Res 47:3854–3864. https://doi.org/10.1016/j.watres.2013.04.010
Jabari L, Gannoun H, Khelifi E et al (2016) Bacterial ecology of abattoir wastewater treated by an anaerobic digestor. Braz J Microbiol 47:73–84. https://doi.org/10.1016/j.bjm.2015.11.029
Johnson M, Zaretskaya I, Raytselis Y et al (2008) NCBI BLAST: a better web interface. Nucleic Acids Res 36:5–9. https://doi.org/10.1093/nar/gkn201
Juretschko S, Loy A, Lehner A, Wagner M (2002) The microbial mommunity momposition of a nitrifying-denitrifying activated sludge from an industrial sewage treatment plant analyzed by the full-cycle rRNA approach. Syst Appl Microbiol 25:84–99. https://doi.org/10.1078/0723-2020-00093
Kamika I, Azizi S, Tekere M (2016) Microbial profiling of South African acid mine water samples using next generation sequencing platform. Appl Microbiol Biotechnol 100:6069–6079. https://doi.org/10.1007/s00253-016-7428-5
Kathiravan V, Krishnani KK (2014) Pseudomonas aeruginosa and Achromobacter sp.: nitrifying aerobic denitrifiers have a plasmid encoding for denitrifying functional genes. World J Microbiol Biotechnol 30:1187–1198. https://doi.org/10.1007/s11274-013-1543-6
Keshri J, Mankazana BBJ, Momba MNB (2015) Profile of bacterial communities in South African mine-water samples using Illumina next-generation sequencing platform. Appl Microbiol Biotechnol 99:3233–3242. https://doi.org/10.1007/s00253-014-6213-6
Kim J, Kim J, Kim H et al (2014) Draft genome sequence of Sphingopyxis sp. strain MWB1, a crude-oil-degrading marine bacterium. Genome Announc 2:2007–2008. https://doi.org/10.1128/genomeA.01256-14.Copyright
Kunito T, Shibata S, Matsumoto S, Oyaizu H (1997) Zinc resistance of Methylobacterium species. Biosci Biotechnol Biochem 61:729–731. https://doi.org/10.1271/bbb.61.729
Lee ZMP, Poret-Peterson AT, Siefert JL et al (2017) Nutrient stoichiometry shapes microbial community structure in an evaporitic shallow pond. Front Microbiol 8:1–15. https://doi.org/10.3389/fmicb.2017.00949
Ma Q, Qu Y, Shen W et al (2015a) Bacterial community compositions of coking wastewater treatment plants in steel industry revealed by Illumina high-throughput sequencing. Bioresour Technol 179:436–443. https://doi.org/10.1016/j.biortech.2014.12.041
Ma Q, Qu Y-Y, Zhang X-W et al (2015b) Identification of the microbial community composition and structure of coal-mine wastewater treatment plants. Microbiol Res 175:1–5. https://doi.org/10.1016/j.micres.2014.12.013
Magnabosco C, Tekere M, Lau MCY et al (2014) Comparisons of the composition and biogeographic distribution of the bacterial communities occupying South African thermal springs with those inhabiting deep subsurface fracture water. Front Microbiol 5:1–17. https://doi.org/10.3389/fmicb.2014.00679
Marchenko AM, Pshinko GN, Demchenko VY, Goncharuk VV (2016) Bioleaching of heavy metals from wastewater sludge by ferrous iron oxidizing bacteria. J Water Chem Technol 38:51–55. https://doi.org/10.3103/S1063455X16010094
Meerbergen K, Van Geel M, Waud M et al (2017) Assessing the composition of microbial communities in textile wastewater treatment plants in comparison with municipal wastewater treatment plants. Microbiology 6:1–13. https://doi.org/10.1002/mbo3.413
Muller EEL, Hourcade E, Louhichi-Jelail Y et al (2011) Functional genomics of dichloromethane utilization in Methylobacterium extorquens DM4. Environ Microbiol 13:2518–2535. https://doi.org/10.1111/j.1462-2920.2011.02524.x
Muyzer G, De Waal E, Uitterlinden A (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Nzila A, Razzak SA, Zhu J (2016) Bioaugmentation: an emerging strategy of industrial wastewater treatment for reuse and discharge. Int J Environ Res Public Health. https://doi.org/10.3390/ijerph13090846
Orisakwe OE, Asomugha R, Afonne OJ et al (2004) Impact of effluents from a car battery manufacturing plant in Nigeria on water, soil, and food qualities. Arch Environ Health 59:31–36. https://doi.org/10.3200/AEOH.59.1.31-36
Pappenhagen JM (1958) Colorimetric determination of nitrates. Anal Chem 30:282–284. https://doi.org/10.1021/ac60134a034
Paul S, Cortez Y, Vera N et al (2016) Metagenomic analysis of microbial community of an Amazonian geothermal spring in Peru. Genomics Data 9:63–66. https://doi.org/10.1016/j.gdata.2016.06.013
Peter H, Sommaruga R (2016) Shifts in diversity and function of lake bacterial communities upon glacier retreat. ISME J 10:1–10. https://doi.org/10.1038/ismej.2015.245
Płaza GA, Jangid K, Łukasik K et al (2008) Reduction of petroleum hydrocarbons and toxicity in refinery wastewater by bioremediation. Bull Environ Contam Toxicol 81:329–333. https://doi.org/10.1007/s00128-008-9411-z
Poi G, Aburto-Medina A, Mok PC et al (2017) Bioremediation of phenol-contaminated industrial wastewater using a bacterial consortium—from laboratory to field. Water Air Soil Pollut. https://doi.org/10.1007/s11270-017-3273-0
Quast C, Pruesse E, Yilmaz P et al (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:590–596. https://doi.org/10.1093/nar/gks1219
Quero GM, Cassin D, Botter M et al (2015) Patterns of benthic bacterial diversity in coastal areas contaminated by heavy metals, polycyclic aromatic hydrocarbons (PAHs) and polychlorinated biphenyls (PCBs). Front Microbiol 6:1–15. https://doi.org/10.3389/fmicb.2015.01053
Rani A, Porwal S, Sharma R et al (2008) Assessment of microbial diversity in effluent treatment plants by culture dependent and culture independent approaches. Bioresour Technol 99:7098–7107. https://doi.org/10.1016/j.biortech.2008.01.003
Ranjard L, Lejon DPH, Mougel C et al (2003) Sampling strategy in molecular microbial ecology: influence of soil sample size on DNA fingerprinting analysis of fungal and bacterial communities. Environ Microbiol 5:1111–1120. https://doi.org/10.1046/j.1462-2920.2003.00521.x
Saiki R, Gelfand D, Stoffel S et al (1988) Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 239:487–491. https://doi.org/10.1126/science.2448875
Sayilgan E, Cakmakci O (2013) Treatment of textile dyeing wastewater by biomass of lactobacillus: lactobacillus 12 and lactobacillus rhamnosus. Environ Sci Pollut Res 20:1556–1564. https://doi.org/10.1007/s11356-012-1009-7
Schloss PD, Westcott SL, Ryabin T et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. https://doi.org/10.1128/AEM.01541-09
Sekar S, Zintchem AAEA, Keshri J et al (2014) Bacterial profiling in brine samples of the Emalahleni water reclamation plant, South Africa, using 454-pyrosequencing method. FEMS Microbiol Lett 359:55–63. https://doi.org/10.1111/1574-6968.12557
Sellami M, Oszako T, Miled N, Ben RF (2015) Industrial wastewater as raw material for exopolysaccharide production by Rhizobium leguminosarum. Braz J Microbiol 46:407–413. https://doi.org/10.1590/S1517-838246220140153
Selvarajan R, Sibanda T, Tekere M (2017a) Thermophilic bacterial communities inhabiting the microbial mats of “indifferent” and chalybeate (iron-rich) thermal springs: diversity and biotechnological analysis. Microbiology:e00560. https://doi.org/10.1002/mbo3.560
Selvarajan R, Sibanda T, Tekere M et al (2017b) Diversity analysis and bioresource characterization of halophilic bacteria isolated from a South African saltpan. Molecules 22:657. https://doi.org/10.3390/molecules22040657
Shchegolkova NM, Krasnov GS, Belova AA et al (2016) Microbial community structure of activated sludge in treatment plants with different wastewater compositions. Front Microbiol 7:1–15. https://doi.org/10.3389/fmicb.2016.00090
Shu D, He Y, Yue H, Wang Q (2015) Microbial structures and community functions of anaerobic sludge in six full-scale wastewater treatment plants as revealed by 454 high-throughput pyrosequencing. Bioresour Technol 186:163–172. https://doi.org/10.1016/j.biortech.2015.03.072
Silva CC, Hayden H, Sawbridge T et al (2012) Phylogenetic and functional diversity of metagenomic libraries of phenol degrading sludge from petroleum refinery wastewater treatment system. AMB Express 2:18. https://doi.org/10.1186/2191-0855-2-18
Somboonna N, Assawamakin A, Wilantho A et al (2012) Metagenomic profiles of free-living archaea, bacteria and small eukaryotes in coastal areas of Sichang island, Thailand. BMC Genomics 13(Suppl 7):S29. https://doi.org/10.1186/1471-2164-13-S7-S29
Tamura K, Stecher G, Peterson D et al (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Tao W, Zhang X-X, Zhao F et al (2016) High levels of antibiotic resistance genes and their correlations with bacterial community and mobile genetic elements in pharmaceutical wastewater treatment bioreactors. PLoS One 11:e0156854. https://doi.org/10.1371/journal.pone.0156854
Tekere M, Sibanda T, Maphangwa KW (2016) An assessment of the physicochemical properties and toxicity potential of carwash effluents from professional carwash outlets in Gauteng Province, South Africa. Environ Sci Pollut Res 23:11876–11884. https://doi.org/10.1007/s11356-016-6370-5
van den Brand TPH, Roest K, Chen GH et al (2015) Potential for beneficial application of sulfate reducing bacteria in sulfate containing domestic wastewater treatment. World J Microbiol Biotechnol 31:1675–1681. https://doi.org/10.1007/s11274-015-1935-x
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Native Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. https://doi.org/10.1128/AEM.00062-07
Wang X, Hu M, Xia Y et al (2012) Pyrosequencing analysis of bacterial diversity in 14 wastewater treatment systems in China. Appl Environ Microbiol 78:7042–7047. https://doi.org/10.1128/AEM.01617-12
Xie W, Wang F, Guo L et al (2011) Comparative metagenomics of microbial communities inhabiting deep-sea hydrothermal vent chimneys with contrasting chemistries. ISME J 5:414–426. https://doi.org/10.1038/ismej.2010.144
Yan QX, Wang YX, Li SP et al (2010) Sphingobium qiguonii sp. nov., a carbaryl-degrading bacterium isolated from a wastewater treatment system. Int J Syst Evol Microbiol 60:2724–2728. https://doi.org/10.1099/ijs.0.020362-0
Yan P, Wang J, Chen YP et al (2015) Investigation of microbial community structure in an advanced activated sludge side-stream reactor process with alkaline treatment. Int Biodeterior Biodegrad 104:356–362. https://doi.org/10.1016/j.ibiod.2015.07.003
Yao X, Zhang J, Tian L, Guo J (2017) The effect of heavy metal contamination on the bacterial community structure at Jiaozhou Bay, China. Braz J Microbiol 48:71–78. https://doi.org/10.1016/j.bjm.2016.09.007
Yu K, Zhang T (2012) Metagenomic and metatranscriptomic analysis of microbial community structure and gene expression of activated sludge. PLoS One. https://doi.org/10.1371/journal.pone.0038183
Zhang T, Shao MF, Ye L (2012) 454 pyrosequencing reveals bacterial diversity of activated sludge from 14 sewage treatment plants. ISME J 6:1137–1147. https://doi.org/10.1038/ismej.2011.188
Zhang Y, Chen L, Sun R et al (2015) Temporal and spatial changes of microbial community in an industrial effluent receiving area in Hangzhou Bay. J Environ Sci 44:57–68. https://doi.org/10.1016/j.jes.2015.11.023
Zhao Y, Ren N, Wang A (2008) Contributions of fermentative acidogenic bacteria and sulfate-reducing bacteria to lactate degradation and sulfate reduction. Chemosphere 72:233–242. https://doi.org/10.1016/j.chemosphere.2008.01.046
Acknowledgements
The authors are thankful for the University of South Africa (UNISA) research fund. Also, the authors are grateful to the industrial owners for allowing the researchers to sample their effluent tanks. Sincere thanks to Mrs. Eunice Iloms for collecting sample.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This study does not contain any study with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(DOCX 39 kb)
Rights and permissions
About this article
Cite this article
Selvarajan, R., Sibanda, T., Venkatachalam, S. et al. Industrial wastewaters harbor a unique diversity of bacterial communities revealed by high-throughput amplicon analysis. Ann Microbiol 68, 445–458 (2018). https://doi.org/10.1007/s13213-018-1349-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13213-018-1349-8