Abstract
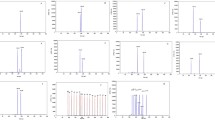
Frequent outbreaks of avian influenza viruses (AIV), which are influenza A viruses, result in heavy economic and national damages. However, current diagnostic methods require about 5–7 days for AIV to be confirmed, which allows enough time for the widespread dissemination of the virus. Therefore, there is a need for rapid and accurate diagnosis of AIV. Here, we developed a diagnostic method using chip-based quantitative reverse transcription-polymerase chain reaction (RT-qPCR) for the rapid identification of the major subtypes H5, H7, and H9 of AIV that influence the pathogenicity of the virus. Specific primer pairs and probes were designed to target the highly conserved Matrix (M) gene regions in influenza A viruses. Specific primer pairs and probes for subtyping were developed by targeting the conserved sequences in the haemagglutinin (HA) gene specific to each subtype. For pathotyping, specific primer pairs and probes for the highly pathogenic avian influenza viruses (HPAIV) were generated by targeting the HA cleavage site, which is highly related to the pathogenicity of the virus. Using the primer pairs and probe sets, we synthesized the major sequence of each type and constructed a standard curve to confirm their detection limit. In addition, we used a chip-based RT-qPCR method to test the samples collected from the faeces of wild birds, cloacal and oropharyngeal swab samples of infected chicken, and allantoic fluids samples incubated with inoculated eggs. As a result, AIV was accurately detected in all samples. Our study shows that by using a single DNA chip in a portable chip-based RT-qPCR device and specific primer pairs and probes, rapid and accurate AIV detection and subtyping and pathotyping is possible, which will reduce the spread of the virus, thereby reducing the economic and national damages.




Similar content being viewed by others
References
Hause, B.M., Ducatez, M., Collin, E.A., Ran, Z., Liu, R., Sheng, Z., Armien, A., Kaplan, B., Chakravarty, S., Hoppe, A.D., Webby, R.J., Simonson, R.R. & Li, F. Isolation of a novel swine influenza virus from Oklahoma in 2011 which is distantly related to Human Influenza C viruses. PLoS Pathog.9, e100 3176 (2013).
Perdue, M.L. & Swayne, D.E. Public health risk from avian influenza viruses. Avian Dis.49, 317–327 (2005).
Senne, D., Panigrahy, B., Kawaoka, Y., Pearson, J. E., Süss, J., Lipkind, M., Kida, H. & Webster, R.G. Survey of the hemagglutinin (HA) cleavage site sequence of H5 and H7 avian influenza viruses: amino acid sequence at the HA cleavage site as a marker of pathogenicity potential. Avian Dis.40, 425–437 (1996).
Koopmans, M., Wilbrink, B., Conyn, M., Natrop, G., van der Nat, H., Vennema, H., Meijer, A., van Steenbergen, J., Fouchier, R. & Osterhaus, A. Transmission of H7N7 avian influenza A virus to human beings during a large outbreak in commercial poultry farms in the Netherlands. Lancet363, 587–593 (2004).
Swayne, D.E. 2016. Animal Influenza, Second Edition, in: Suarez, D.L. (Eds.), Influenza A virus. John Wiley & Sons, Inc., Hobokenk, pp. 1–30.
Kim, H.-R., Lee, Y.-J., Park, C.-K., Oem, J.-K., Lee, O.-S., Kang, H.-M., Choi, J.-G. & Bae, Y.-C. Highly pathogenic avian influenza (H5N1) outbreaks in wild birds and poultry, South Korea. Emerging Infect. Dis.18, 480–483 (2012).
Hoffmann, E., Stech, J., Guan, Y., Webster, R.G. & Perez, D.R. Universal primer set for the full-length amplification of all influenza A viruses. Arch. Virol.146, 2275–2289 (2001).
Behlke, M.A., Huang, L., Bogh, L., Rose, S. & Devor, E. J. Fluorescence and fluorescence applications. Integr. DNA Technol. 1–13. (2005).
Marras, S.A.E., Kramer, F.R. & Tyagi, S. Efficiencies of fluorescence resonance energy transfer and contact-mediated quenching in oligonucleotide probes. Nucleic Acids Res.30, e122 (2002).
Atkinson, C., Emery, V. & Griffiths, P. Development of a novel single tube nested PCR for enhanced detection of cytomegalovirus DNA from dried blood spots. J. Virol. Methods196, 40–44 (2014).
Koshkin, A.A., Singh, S.K., Nielsen, P., Rajwanshi, V.K., Kumar, R., Meldgaard, M., Olsen, C.E. & Wengel, J. LNA (Locked Nucleic Acids): Synthesis of the adenine, cytosine, guanine, 5-methylcytosine, thymine and uracil bicyclonucleoside monomers, oligomerisation, and unprecedented nucleic acid recognition. Tetrahedron54, 3607–3630 (1998).
Astakhova, K. Toward non-enzymatic ultrasensitive identification of single nucleotide polymorphisms by optical methods. Chemosensors2, 193–206. (2014).
Ugozzoli, L.A., Latorra, D., Pucket, R. Arar, K. & Hamby, K. Real-time genotyping with oligonucleotide probes containing locked nucleic acids. Anal. Biochem.324, 143–152 (2004).
Ahn, J.J., Kim, Y., Hong, J.Y., Kim, G.W., Kim, S. Y. & Hwang, S.Y. Probe-based fluorescence melting curve analysis for differentiating Larimichthys polyactis and Larimichthys crocea. Food Anal. Methods9, 2036–2041 (2016).
Braasch, D.A. & Corey, D.R. Locked nucleic acid (LNA): fine-tuning the recognition of DNA and RNA. Chem. Biol.8, 1–7 (2001).
Schmittgen, T.D. & Livak, K.J. Analyzing real-time PCR data by the comparative C T method. Nat. Protoc.3, 1101–1108 (2008).
Bustin, S.A., Benes, V., Garson, J.A., Hellemans, J., Huggett, J., Kubista, M., Mueller, R., Nolan, T., Pfaffl, M.W. & Shipley, G.L. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin. Chem.55, 611–622 (2009).
Schefe, J.H., Lehmann, K.E., Buschmann, I.R., Unger, T. & Funke-Kaiser, H. Quantitative real-time RT-PCR data analysis: current concepts and the novel “gene expression’s C T difference” formula. J. Mol. Med.84, 901–910 (2006).
Holland, P.M., Abramson, R.D., Watson R. & Gelfand, D.H. Detection of specific polymerase chain reaction product by utilizing the 5′—3′exonuclease activity of Thermus aquaticus DNA polymerase. Proc. Natl. Acad. Sci. U. S. A.88, 7276–7280 (1991).
Arya, M., Shergill, I. S., Williamson, M., Gommersall, L., Arya, N. & Patel, H. R. Basic principles of real-time quantitative PCR. Expert Rev. Mol. Diagn.5, 209–219 (2005).
Hall, T.A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser.41, 95–98 (1999).
Kwon, S.H., Lee, S., Jang, J., Seo, Y. & Lim, H.Y. A point-of-care diagnostic system to influenza viruses using chip-based ultra-fast PCR. J. Med. Virol.90, 1019–1102. (2018).
Schrader, C., Schielke, A., Ellerbroek, L. & Johne, R. PCR inhibitors-occurrence, properties and removal. J. Appl. Microbiol.113, 1014–1026 (2012).
Acknowledgements
This work was supported by Korea Institute of Planning and Evaluation for Technology in Food, Agriculture, Forestry (IPET) through Animal Disease Management Technology Development Program, funded by Ministry of Agriculture, Food and Rural Affairs (MAFRA) (116103032HD020).
Author information
Authors and Affiliations
Corresponding author
Additional information
Conflict of Interests
The authors declare no competing financial interests.
These authors contrilbuted equally.
Rights and permissions
About this article
Cite this article
Kwon, N.y., Ahn, J.J., Kim, JH. et al. Rapid Subtyping and Pathotyping of Avian Influenza Virus using Chip-based RT-PCR. BioChip J 13, 333–340 (2019). https://doi.org/10.1007/s13206-019-3405-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13206-019-3405-2