Abstract
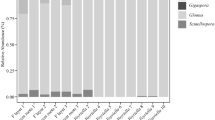
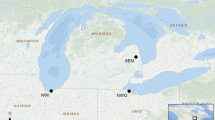
Symbioses between plants and nitrogen-fixing cyanobacteria are benchmark biological systems to understand mutualism and patterns of coevolution. The diversity of cyanobacteria associated to non-vascular land plants is still being unraveled, with a huge gap in knowledge from tropical areas, especially in hornworts. We focus on the Neotropical hornwort Leiosporoceros dussii, which has a semi-permanent association with cyanobacteria. Unlike other hornworts, the peculiar Nostoc canals suggest cyanobiont specificity and selectivity. Here, we characterize the first hornwort bacteriome and focus on the endophytic cyanobacteria by describing its morphology, metagenomic diversity and phylogeny between two sampling sites in Panama. First, we use a metagenomic analyses (16S) from gametophytes from Río El Guayabo and Río Indio in Panama to identify the endophytic bacteria community of L. dussii. To provide more information on the cyanobacterial endophytes, we have extracted cyanobacterial endophytes to measure cell size variation and use the markers trnL and rbcLX to determine their phylogenetic relationships. Finally, we use stable isotope δ 15N to assess nitrogen flow in the plants. We find little diversity in Leiosporoceros bacteriome but observe significant community variations between sites. Results indicate a lack of cyanobacterial specificity of cyanobacteria associated to Leiosporoceros, with six unrelated clades of Nostoc forming a polyphyletic assemblage. Four clades are nested within bacteria collected as free-living taxa suggesting that typically non-symbiotic cyanobacteria can be associated with this hornwort. Finally, the little nitrogen fractioning of the plant suggests the role of the cyanobacterial symbionts in nitrogen fixation. This study provides the first hornwort bacteriome and is an important step in the characterization of tropical bryophyte symbioses.





Similar content being viewed by others
Data availability
The dataset generated and analysed during the current study is available from GenBank and DRYAD (https://doi.org/10.5061/dryad.sxksn030k).
References
Adams DG (2002) Cyanobacteria in Symbiosis with hornworts and liverworts. In: Rai AN, Bergman B, Rasmussen U (eds) Cyanobacteria in Symbiosis. Springer, Dordrecht, pp 117–135. https://doi.org/10.1007/0-306-48005-0_7
Adams DG, Duggan PS (2008) Cyanobacteria-bryophyte symbioses. J Exp Bot 59:1047–1058. https://doi.org/10.1093/jxb/ern005
Alcaraz LD, Peimbert M, Barajas HR, Dorantes-Acosta AE, Bowman JL, Arteaga-Vázquez MA (2018) Marchantia liverworts as a proxy to plants’ basal microbiomes. Sci Rep 8:1–12. https://doi.org/10.1038/s41598-018-31168-0
Arróniz-Crespo M, Pérez-Ortega S, De Los RA et al (2014) Bryophyte-cyanobacteria associations during primary succession in recently deglaciated areas of Tierra del Fuego (Chile). PLoS One 9:15–17. https://doi.org/10.1371/journal.pone.0096081
Barrow JR, Lucero ME, Reyes-Vera I, Havstad KM (2008) Do symbiotic microbes have a role in regulating plant performance and response to stress? Commun Integr Biol 1:69–73. https://doi.org/10.4161/cib.1.1.6238
Bell-Doyon P, Laroche J, Saltonstall K, Villarreal Aguilar JC (2020) Specialized bacteriome uncovered in the coralloid roots of the epiphytic gymnosperm, Zamia pseudoparasitica. Environmental DNA:1–11. https://doi.org/10.1002/edn3.66
Brookes PC, Powlson DS, Jenkinson DS (1982) Measurement of microbial biomass phosphorus in soil. Soil Biol Biochem 14:319–329. https://doi.org/10.1016/0038-0717(82)90001-3
Callahan B, McMurdie P, Rosen M et al (2016a) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Callahan B, Sankaran K, Fukuyama J et al (2016b) Bioconductor workflow for microbiome data analysis: from raw reads to community analyses [version 1; referees: 3 approved]. F1000Research 5:1–49. https://doi.org/10.12688/F1000RESEARCH.8986.1
Callahan B, Sankaran K, Julia A, McMurdie P, Holmes S (2017) Workflow for microbiome data analysis: from raw reads to community analyses. IOP Publishing. https://bioconductor.org/help/course-materials/2017/BioC2017/Day1/Workshops/Microbiome/MicrobiomeWorkflowII.html#acknowledgments. Accessed 20 May 2019
Cernava T, Erlacher A, Aschenbrenner IA, Krug L, Lassek C, Riedel K, Grube M, Berg G (2017) Deciphering functional diversification within the lichen microbiota by meta-omics. Microbiome 5:82. https://doi.org/10.1186/s40168-017-0303-5
Costa J-L, Paulsrud P, Rikkinen J, Lindblad P (2001) Genetic diversity of Nostoc Symbionts Endophytically associated with two bryophyte species. Appl Environ Microbiol 67:4393–4396. https://doi.org/10.1128/AEM.67.9.4393-4396.2001
Costa JL, Paulsrud P, Lindblad P (2002) The cyanobacterial tRNALeu (UAA) intron: evolutionary patterns in a genetic marker. Mol Biol Evol 19:850–857. https://doi.org/10.1093/oxfordjournals.molbev.a004142
Costa JL, Romero EM, Lindblad P (2004) Sequence based data supports a single Nostoc strain in individual coralloid roots of cycads. FEMS Microbiol Ecol 49:481–487. https://doi.org/10.1016/j.femsec.2004.05.001
Darnajoux R, Constantin J, Miadlikowska J, Lutzoni F, Bellenger JP (2014) Is vanadium a biometal for boreal cyanolichens? New Phytol 202:765–771. https://doi.org/10.1111/nph.12777
DeLuca T, Zackrisson O, Nilsson M, Sellstedt A (2002) Quantifying nitrogen-fixation in feather moss carpets of boreal forests. Nature 419:917–920. https://doi.org/10.1038/nature01051
DeLuca T, Zackrisson O, Gentili F et al (2007) Ecosystem controls on nitrogen fixation in boreal feather moss communities. Oecologia 152:121–130. https://doi.org/10.1007/s00442-006-0626-6
Desiró A, Duckett JG, Pressel S Villarreal JC, Bidartondo MI (2013) Fungal symbioses in hornworts: a chequered history. Proc R Soc B-Biol Sci 280:20130207. https://doi.org/10.1098/Rspb.2013.0207
Dixon P (2003) Computer program review VEGAN , a package of R functions for community ecology. J Veg Sci 14:927–930
Fedrowitz K, Kaasalainen U, Rikkinen J (2011) Genotype variability of Nostoc symbionts associated with three epiphytic Nephroma species in a boreal forest landscape. Bryologist 114:220–230. https://doi.org/10.1639/0007-2745-114.1.220
Gentili F, Nilsson M, Zackrisson O et al (2005) Physiological and molecular diversity of feather moss associative N 2-fixing cyanobacteria. J Exp Bot 56:3121–3127. https://doi.org/10.1093/jxb/eri309
Gotelli NJ, Colwell RK (2001) Quantifying biodiversity: procedures and pitfalls in the measurement and comparison of species richness. Ecol Lett 4:379–391. https://doi.org/10.1046/j.1461-0248.2001.00230.x
Gutiérrez-García K, Bustos-Dıaz ED, Corona-Gomez JA et al (2019) Cycad coralloid roots contain bacterial communities including cyanobacteria and Caulobacter spp. that encode niche-specific biosynthetic gene clusters. Genome Biol Evol 11:319–334. https://doi.org/10.1093/gbe/evy266
Holland-Moritz H, Stuart J, Lewis LR, Miller S, Mack MC, McDaniel SF, Fierer N (2018) Novel bacterial lineages associated with boreal moss species. Environ Microbiol 20:2625–2638. https://doi.org/10.1111/1462-2920.14288
Hughes JB, Hellmann JJ, Ricketts TH, Bohannan BJM (2001) Counting the uncountable: statistical approaches to estimating microbial diversity. Appl Environ Microbiol 10:4399–4406. https://doi.org/10.1128/AEM.67.10.4399
Joneson S, O’Brien H (2017) A molecular investigation of free-living and lichenized Nostoc sp. and symbiotic lifestyle determination. Bryologist 120:371–381. https://doi.org/10.1639/0007-2745-120.4.371
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glöckner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41:1–11. https://doi.org/10.1093/nar/gks808
Knack JJ, Wilcox LW, Delaux P-M, Ané JM, Piotrowski MJ, Cook ME, Graham JM, Graham LE (2015) Microbiomes of Streptophyte algae and bryophytes suggest that a functional suite of microbiota fostered plant colonization of land. Int J Plant Sci 176:405–420. https://doi.org/10.1086/681161
Lavoie C, Renaudin M, Troy McMullin R, Gagnon J, Roy C, Beaulieu M-E, Bellenger JP, Villarreal JC (2020) Extremely low genetic diversity of Stigonema associated with Stereocaulon in eastern Canada. Bryologist 123:188–203. https://doi.org/10.1639/0007-2745-123.2.188
LeBauer DS, Treseder KK (2008) Nitrogen limitation of net primary productivity in terrestrial ecosystems is globally distributed. Ecology 89:371–379. https://doi.org/10.1890/06-2057.1
Lindo Z, Nilsson MC, Gundale MJ (2013) Bryophyte-cyanobacteria associations as regulators of the northern latitude carbon balance in response to global change. Glob Chang Biol 19:2022–2035. https://doi.org/10.1111/gcb.12175
Lodge DJ, McDowell WH, McSwiney CP (1994) The importance of nutrient pulses in tropical forests. Trends Ecol Evol 9:384–387. https://doi.org/10.1016/0169-5347(94)90060-4
Magain N, Miadlikowska J, Goffinet B, Sérusiaux E, Lutzoni F (2016) Macroevolution of specificity in cyanolichens of the genus Peltigera section Polydactylon (Lecanoromycetes, Ascomycota). Syst Biol 66:74–99. https://doi.org/10.1093/sysbio/syw065
McMurdie PJ, Holmes S (2013) Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8:e61217. https://doi.org/10.1371/journal.pone.0061217
Meeks JC (2007) Physiological adaptations in nitrogen-fixing Nostoc–plant symbiotic associations. In: Prokaryotic symbionts in plants. Springer-Verlag, Berlin Heidelberg, pp 181–205
Nelson J, Shaw AJ (2019) Exploring the natural microbiome of the model liverwort: fungal endophyte diversity in Marchantia polymorpha L. Symbiosis. 78:45–59. https://doi.org/10.1007/s13199-019-00597-4
Nelson JM, Hauser DA, Hinson R, Shaw AJ (2018) A novel experimental system using the liverwort Marchantia polymorpha and its fungal endophytes reveals diverse and context-dependent effects. New Phytol 218:1217–1232. https://doi.org/10.1111/nph.15012
Nelson JM, Hauser DA, Gudiño JA, Guadalupe YA, Meeks JC, Salazar Allen N, Villarreal JC, Li FW (2019) Complete genomes of symbiotic cyanobacteria clarify the evolution of vanadium-Nitrogenase. Genome Biol Evol 11:1959–1964. https://doi.org/10.1093/gbe/evz137
O’Brien H, Miadlikowska J, Lutzoni F (2005) Assessing host specialization in symbiotic cyanobacteria associated with four closely related species of the lichen fungus Peltigera. Eur J Phycol 40:363–378. https://doi.org/10.1080/09670260500342647
O’Brien HE, Miadlikowska J, Lutzoni F (2013) Assessing population structure and host specialization in lichenized cyanobacteria. New Phytol 198:557–566. https://doi.org/10.1111/nph.12165
Oksanen J, Blanchet FG, Friendly M, et al (2019) Package ‘vegan.’ Community Ecology Package. R package version 2.5–6
Pardo-De la Hoz CJ, Magain N, Lutzoni F et al (2018) Contrasting symbiotic patterns in two closely related lineages of Trimembered lichens of the genus Peltigera. Front Microbiol 9:1–14. https://doi.org/10.3389/fmicb.2018.02770
R Core Team R (2017) R core team. R: A Language and Environment for Statistical Computing [Internet]
Remy W, Taylor T, Hass H, Kerp H (1994) Four hundred-million-year-old vesicular arbuscular mycorrhizae. Proc Natl Acad Sci U S A 91:11841–11843. https://doi.org/10.1073/pnas.91.25.11841
Renzaglia KS (1978) A comparative morphology and developmental anatomy of the Anthocerotophyta. J Hattori Bot Lab 44:31–90
Renzaglia KS, Schuette S, Duff J et al (2007) Bryophyte phylogeny: advancing the molecular and morphological frontiers. Bryologist 110:179–213. https://doi.org/10.1639/0007-2745(2007)110[179:BPATMA]2.0.CO;2
Rikkinen J, Virtanen V (2008) Genetic diversity in cyanobacterial symbionts of thalloid bryophytes. J Exp Bot 59:1013–1021. https://doi.org/10.1093/jxb/ern003
Rousk K, Jones DL, DeLuca TH (2013) Moss-cyanobacteria associations as biogenic sources of nitrogen in boreal forest ecosystems. Front Microbiol 4:1–10. https://doi.org/10.3389/fmicb.2013.00150
RStudio Team (2015) RStudio: Integrated Development for R. RStudio, Inc., Boston, MA. http://www.rstudio.com/
Rudi K, Skulberg O, Jakobsen K (1998) Evolution of cyanobacteria by exchange of genetic material among Phyletically related strains. J Bacteriol 180:3453–3461. https://doi.org/10.1128/JB.180.13.3453-3461.1998
Schirrmeister B, de Vos J, Antonelli A, Bagheri H (2013) Evolution of multicellularity coincided with increased diversification of cyanobacteria and the great oxidation event. PNAS 110:1791–1796. https://doi.org/10.1073/pnas.1209927110
Schlesinger W, Klein EM, Vengosh A (2017) Global biogeochemical cycle of vanadium. Proc Natl Acad Sci U S A 114:E11092–E11100. https://doi.org/10.1073/pnas.1715500114
Sierra MA, Danko DC, Sandoval TA, Pishchany G, Moncada B, Kolter R, Mason CE, Zambrano MM (2020) The microbiomes of seven lichen genera reveal host specificity, a reduced Core Community and potential as source of antimicrobials. Front Microbiol 11:1–12. https://doi.org/10.3389/fmicb.2020.00398
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313
Villarreal AJC, Renzaglia KS (2006) Structure and development of Nostoc strands in Leiosporoceros dussii (Anthocerotophyta): a novel symbiosis in land plants. Am J Bot 93:693–705. https://doi.org/10.3732/ajb.93.5.693
Vincent AT, Derome N, Boyle B, Culley AI, Charette SJ (2017) Next-generation sequencing (NGS) in the microbiological world: how to make the most of your money. J Microbiol Methods 138:60–71. https://doi.org/10.1016/j.mimet.2016.02.016
Ward LM, Cardona T, Holland-Moritz H (2019) Evolutionary implications of Anoxygenic Phototrophy in the bacterial phylum Candidatus Eremiobacterota (WPS-2). Front Microbiol 10:1–19. https://doi.org/10.3389/fmicb.2019.01658
West NJ, Adams DG (1997) Phenotypic and genotypic comparison of symbiotic and free-living cyanobacteria from a single field site. Appl Environ Microbiol 63:4479–4484
Wickham H, Chang W (2009) ggplot2: an implementation of the grammar of graphics. R package version 0.8. 3
Wong FCY, Meeks JC (2002) Establishment of a functional symbiosis between the cyanobacterium Nostoc punctiforme and the bryophyte Anthoceros punctatus requires genes involved in nitrogen control and initiation of heterocyst differentiation the GenBank accession numbers for the cod. Microbiology 148:315–323. https://doi.org/10.1099/00221287-148-1-315
Wright D, Prickett T, Helm R, Potts M (2001) Form species Nostoc commune (cyanobacteria). Int J Syst Evol Microbiol 51:1839–1852. https://doi.org/10.1099/00207713-51-5-1839
Zheng Y, Chiang T-Y, Huang C-L, Gong X (2018) Highly diverse endophytes in roots of Cycas bifida (Cycadaceae), an ancient but endangered gymnosperm. J Microbiol 56:337–345. https://doi.org/10.1007/s12275-018-7438-3
Funding
Établissement de nouveaux chercheurs universitaires FRQNT- 206943 to JCVA.
Earl S. Tupper Fellowship 2015 to JCVA.
National Science Foundation Dimensions of Biodiversity grant (1831428) to F.-W.L.
Author information
Authors and Affiliations
Contributions
RB and JCV conceived the study and performed the phylogenetic work and wrote the first version of the manuscript; GPB did the metagenomic analysis; ST performed the analysis of isotope data, YG did the morphological measurements, JG and NSA provided logistic support in the field and funding; FWL contributed with logistic support and funding; RB, GPB, NS, FWL, ST and JCV wrote and edited the manuscript.
Corresponding author
Ethics declarations
Conflicts of interest/competing interests
Authors have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Bouchard, R., Peñaloza-Bojacá, G., Toupin, S. et al. Contrasting bacteriome of the hornwort Leiosporoceros dussii in two nearby sites with emphasis on the hornwort-cyanobacterial symbiosis. Symbiosis 81, 39–52 (2020). https://doi.org/10.1007/s13199-020-00680-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13199-020-00680-1