Abstract
Purpose
To find immune-related genes with prognostic value in breast cancer, and construct a prognostic risk assessment model to make a more accurate assessment. Moreover, looking for potential immune markers for breast cancer immunotherapy.
Methods
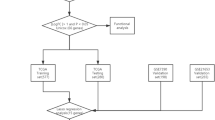
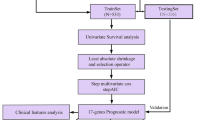
The breast cancer (BC) data were retrieved from The Cancer Genome Atlas (TCGA) database as a training set. Through the Weighted gene co-expression network analysis (WGCNA), Kaplan–Meier (KM) analysis, lasso regression analysis and stepwise backward Cox regression analysis, screening for prognosis-related immune genes, a prognostic index was built, and external validation with two data sets of Gene Expression Omnibus (GEO) database was performed. Transcription factor (TF) regulatory network was constructed to identify key transcription factors that regulate prognostic immune genes. Gene set enrichment analysis (GSEA) was used to explore the signal pathways differences between high and low-risk groups, estimate package and TIMER database were used to evaluate the relationship between risk score and tumor immune microenvironment.
Results
We obtained 10 prognosis-related immune genes, and the index showed accurate prognostic value. We also identified 7 prognostic transcription factors. Multiple signaling pathways that inhibit tumor progression were enriched in the low-risk group, and risk score was significantly negatively related to the degree of immune infiltration and the expression level of immune checkpoint genes.
Conclusion
We successfully constructed an independent prognostic index, which not only has a stronger predictive ability than the tumor pathological stage, but also can reflect the immune infiltration of breast cancer patients.







Similar content being viewed by others
Abbreviations
- BC:
-
Breast cancer
- TCGA:
-
The cancer genome atlas
- GEO:
-
Gene expression omnibus
- WGCNA:
-
Weighted gene co-expression network analysis
- GSEA:
-
Gene set enrichment analysis
- TME:
-
Tumor microenvironment
- IRGs:
-
Immune-related genes
- TF:
-
Transcription factor
- FPKM:
-
Fragments per kilobase million
- TPM:
-
Transcripts per million
- GS:
-
Gene significance
- MS:
-
Module significance
- KM:
-
Kaplan–Meier
- RS:
-
Risk score
- OS:
-
Overall survival
- HR:
-
Hazard ratio
References
Bray F, Ferlay J, Soerjomataram I et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA. 2018; 68:394–424.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA. 70:7–30.
Yeo SK, Guan JL. Breast Cancer: Multiple Subtypes within a Tumor? Trends Cancer. 2017;3:753–60.
Emens LA, Ascierto PA, Darcy PK et al. Cancer immunotherapy: opportunities and challenges in the rapidly evolving clinical landscape. European journal of cancer (Oxford, England: 1990) 2017; 81:116–129.
Kalbasi A, Ribas A. Tumour-intrinsic resistance to immune checkpoint blockade. Nat Rev Immunol. 2020;20:25–39.
Tray N, Taff J, Adams S. Therapeutic landscape of metaplastic breast cancer. Cancer Treat Rev. 2019;79:101888.
Emens LA. Breast Cancer Immunotherapy: Facts and Hopes. Clin Cancer Res. 2018;24:511–20.
Mittal S, Brown NJ, Holen I. The breast tumor microenvironment: role in cancer development, progression and response to therapy. Expert Rev Mol Diagn. 2018;18:227–43.
Savas P, Salgado R, Denkert C, et al. Clinical relevance of host immunity in breast cancer: from TILs to the clinic. Nat Rev Clinical Oncol. 2016;13:228–41.
Ernst B, Anderson KS. Immunotherapy for the treatment of breast cancer. Curr Oncol Rep. 2015;17:5.
Kroemer G, Senovilla L, Galluzzi L, et al. Natural and therapy-induced immunosurveillance in breast cancer. Nat Med. 2015;21:1128–38.
Denkert C, von Minckwitz G, Darb-Esfahani S, et al. Tumour-infiltrating lymphocytes and prognosis in different subtypes of breast cancer: a pooled analysis of 3771 patients treated with neoadjuvant therapy. Lancet Oncol. 2018;19:40–50.
Burugu S, Asleh-Aburaya K, Nielsen TO. Immune infiltrates in the breast cancer microenvironment: detection, characterization and clinical implication. Breast Cancer. 2017;24:3–15.
Stanton SE, Disis ML. Clinical significance of tumor-infiltrating lymphocytes in breast cancer. J Immunotherapy Cancer. 2016;4:59.
Asano Y, Kashiwagi S, Goto W, et al. Tumour-infiltrating CD8 to FOXP3 lymphocyte ratio in predicting treatment responses to neoadjuvant chemotherapy of aggressive breast cancer. Br J Surg. 2016;103:845–54.
Bhattacharya S, Andorf S, Gomes L, et al. ImmPort: disseminating data to the public for the future of immunology. Immunol Res. 2014;58:234–9.
Tang Q, Chen Y, Meyer C, et al. A comprehensive view of nuclear receptor cancer cistromes. Can Res. 2011;71:6940–7.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics. 2008;9:559.
Zhou Y, Zhou B, Pache L, et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun. 2019;10:1523–1523.
Rizvi AA, Karaesmen E, Morgan M, et al. gwasurvivr: an R package for genome-wide survival analysis. Bioinformatics (Oxford, England). 2019;35:1968–70.
Friedman J, Hastie T, Tibshirani R. Regularization paths for generalized linear models via coordinate descent. J Stat Softw. 2010;33:1–22.
Shannon P, Markiel A, Ozier O, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–504.
Chin CH, Chen SH, Wu HH, et al. cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol. 2014;8(Suppl 4):S11.
Uno H, Cai TX, Tian L, Wei LJ. Evaluating prediction rules for t-year survivors with censored regression models. J Am Stat Assoc. 2007;102:527–37.
Park SY. Nomogram: an analogue tool to deliver digital knowledge. J Thoracic Cardiovasc Surg. 2018;155:1793.
Subramanian A, Tamayo P, Mootha VK, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Liberzon A, Birger C, Thorvaldsdóttir H, et al. The molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst. 2015;1:417–25.
Yoshihara K, Shahmoradgoli M, Martínez E, et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4:2612.
Li T, Fan J, Wang B, et al. TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Can Res. 2017;77:e108–10.
Disis ML, Stanton SE. Immunotherapy in breast cancer: an introduction. Breast (Edinburgh, Scotland). 2018;37:196–9.
Cassetta L, Pollard JW. Repolarizing macrophages improves breast cancer therapy. Cell Res. 2017;27:963–4.
Ban Y, Mai J, Li X, et al. Targeting autocrine CCL5-CCR5 axis reprograms immunosuppressive myeloid cells and reinvigorates antitumor immunity. Can Res. 2017;77:2857–68.
Dangaj D, Bruand M, Grimm AJ, et al. Cooperation between constitutive and inducible chemokines enables T cell engraftment and immune attack in solid tumors. Cancer Cell. 2019;35(885–900):e810.
Rizeq B, Malki MI. The Role of CCL21/CCR7 chemokine axis in breast cancer progression. cancers (basel). 2020;12:1036.
Klimczak M, Biecek P, Zylicz A, Zylicz M. Heat shock proteins create a signature to predict the clinical outcome in breast cancer. Sci Rep. 2019;9:7507.
Wu J, Wan F, Sheng H, et al. NR1H3 expression is a prognostic factor of overall survival for patients with muscle-invasive bladder cancer. J Cancer. 2017;8:852–60.
Yen MC, Huang YC, Kan JY, et al. S100B expression in breast cancer as a predictive marker for cancer metastasis. Int J Oncol. 2018;52:433–40.
Charmsaz S, Hughes É, Bane FT, et al. S100β as a serum marker in endocrine resistant breast cancer. BMC Med. 2017;15:79.
Saha SK, Yin Y, Chae HS, Cho SG. Opposing Regulation of cancer properties via KRT19-Mediated Differential Modulation Of Wnt/β-catenin/notch signaling in breast and colon cancers. Cancers (Basel) 2019; 11.
Shao L, Hou W, Scharping NE, et al. IRF1 inhibits antitumor immunity through the upregulation of PD-L1 in the tumor cell. Cancer Immunol Res. 2019;7:1258–66.
Kim GC, Kwon HK, Lee CG, et al. Upregulation of Ets1 expression by NFATc2 and NFKB1/RELA promotes breast cancer cell invasiveness. Oncogenesis. 2018;7:91.
Chen N, Zhao G, Yan X, et al. A novel FLI1 exonic circular RNA promotes metastasis in breast cancer by coordinately regulating TET1 and DNMT1. Genome Biol. 2018;19:218.
Zhao X, Liu J, Ge S, et al. Saikosaponin A inhibits breast cancer by regulating Th1/Th2 balance. Front Pharmacol. 2019;10:624.
Peck AR, Witkiewicz AK, Liu C, et al. Low levels of Stat5a protein in breast cancer are associated with tumor progression and unfavorable clinical outcomes. Breast Cancer Res. 2012;14:R130.
Berraondo P, Sanmamed MF, Ochoa MC, et al. Cytokines in clinical cancer immunotherapy. Br J Cancer. 2019;120:6–15.
Braumüller H, Wieder T, Brenner E, et al. T-helper-1-cell cytokines drive cancer into senescence. Nature. 2013;494:361–5.
Müller-Hermelink N, Braumüller H, Pichler B, et al. TNFR1 signaling and IFN-gamma signaling determine whether T cells induce tumor dormancy or promote multistage carcinogenesis. Cancer Cell. 2008;13:507–18.
Pagès F, Galon J, Dieu-Nosjean MC, et al. Immune infiltration in human tumors: a prognostic factor that should not be ignored. Oncogene. 2010;29:1093–102.
Saleh R, Toor SM, Khalaf S, Elkord E. Breast Cancer Cells and PD-1/PD-L1 Blockade Upregulate the Expression of PD-1, CTLA-4, TIM-3 and LAG-3 Immune Checkpoints in CD4(+) T Cells. Vaccines 2019; 7.
Zhu B, Tse LA, Wang D, et al. Immune gene expression profiling reveals heterogeneity in luminal breast tumors. Breast Cancer Res. 2019;21:147.
Acknowledgements
The authors would like to thank the TCGA, GEO, ImmPort, cBioPortal, and Cistrome cancer databases for the availability of the data needed for analyses. This work was supported by grants from National Natural Science Foundation of China (81272372 and 30873044) and by Zhongnan Hospital of Wuhan University Science, Technology and Innovation Seed Fund (znpy2016033). The Project-sponsored by SRF for ROCS, SEM.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There authors declare no conflict of interest in relation to the submission.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
About this article
Cite this article
Yu, X., Guo, J., Zhou, Q. et al. A novel immune-related prognostic index for predicting breast cancer overall survival. Breast Cancer 28, 434–447 (2021). https://doi.org/10.1007/s12282-020-01175-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12282-020-01175-z