Abstract
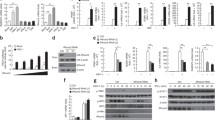
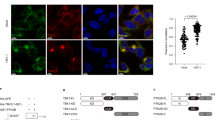
We previously reported that human cytomegalovirus (HCMV) 86 kDa immediate-early 2 gene product (IE86) promotes proteasome-dependent degradation of STING. In the present study, we determined the specific residues of IE86 responsible for STING degradation using a STING-firefly luciferase fusion protein expression system for quantitative meas-urement of STING protein levels. IE86 amino acids (aa) 136-289 were sufficient to promote STING degradation and further induced down-regulation of 2′3′-cyclic GMP-AMP (cGAMP)-mediated IFN-β promoter activation. Interestingly, transactivation domains (TAD) of the IE86 protein located at the N- and C-termini were required for down-regulation of Toll/interleukin-1 receptor (TIR) domain-containing adaptor-inducing interferon β (IFN-β) (TRIF)-mediated IFN-β-and p65/RelA-induced NF-κB-dependent promoter activation while amino acids (aa) 136-289 had no significant effects. Our collective data suggest that the IE86 protein utilizes the aa 136-289 region to promote STING degradation and inhibit the cGAS-STING pathway.
Similar content being viewed by others
References
Ablasser, A., Goldeck, M., Cavlar, T., Deimling, T., Witte, G., Rohl, I., Hopfner, K.P., Ludwig, J., and Hornung, V. 2013. cGAS produces a 2′–5′-linked cyclic dinucleotide second messenger that activates STING. Natured498, 380–384.
Ahn, J.H., Jang, W.J., and Hayward, G.S. 1999. The human cytomegalovirus IE2 and UL112-113 proteins accumulate in viral DNA replication compartments that initiate from the periphery of pro-myelocytic leukemia protein-associated nuclear bodies (PODs or ND10). J. Virol.d73, 10458–10471.
Choi, H.J., Park, A., Kang, S., Lee, E., Lee, T.A., Ra, E.A., Lee, J., Lee, S., and Park, B. 2018. Human cytomegalovirus-encoded US9 targets MAVS and STING signaling to evade type I interferon immune responses. Nat. Commun.9, 125.
Civril, F., Deimling, T., de Oliveira Mann, C.C., Ablasser, A., Moldt, M., Witte, G., Hornung, V., and Hopfner, K.P. 2013. Structural mechanism of cytosolic DNA sensing by cGAS. Natured498, 332–337.
Diner, E.J., Burdette, D.L., Wilson, S.C., Monroe, K.M., Kellenberger, C.A., Hyodo, M., Hayakawa, Y., Hammond, M.C., and Vance, R.E. 2013. The innate immune DNA sensor cGAS produces a noncanonical cyclic dinucleotide that activates human STING. Cell Rep.d3, 1355–1361.
Fu, Y.Z., Su, S., Gao, Y.Q., Wang, P.P., Huang, Z.F., Hu, M.M., Luo, W.W., Li, S., Luo, M.H., Wang, Y.Y.,et al. 2017. Human cytomegalovirus tegument protein UL82 inhibits STING-mediated signaling to evade antiviral immunity. Cell Host Microbe21, P231–P243.
Giorgi, C,, Ito, K., Lin, H.K., Santangelo, C., Wieckowski, M.R., Lebiedzinska, M., Bononi, A., Bonora, M., Duszynski, J., Bernardi, R.,et al. 2010. PML regulates apoptosis at endoplasmic reticulum by modulating calcium release. Scienced330, 1247–1251.
Huang, Z.F., Zou, H.M., Liao, B.W., Zhang, H.Y., Yang, Y., Fu, Y.Z., Wang, S.Y., Luo, M. H., and Wang, Y.Y. 2018. Human cytomegalovirus protein UL31 inhibits DNA sensing of cGAS to mediate immune evasion. Cell Host Microbed24, 69–80.e64.
Kalamvoki, M. and Roizman, B. 2014. HSV-1 degrades, stabilizes, requires, or is stung by STING depending on ICP0, the US3 protein kinase, and cell derivation. Proc. Natl. Acad. Sci. USA111, E611–E617.
Kim, J.E., Kim, Y.E., Stinski, M.F., Ahn, J.H., and Song, Y.J. 2017. Human cytomegalovirus IE2 86 kDa protein induces STING degradation and inhibits cGAMP-mediated IFN-β induction. Front. Microbiol.8, 1854.
Konno, H., Konno, K, and Barber, G.N. 2013. Cyclic dinucleotides trigger ULK1 (ATG1) phosphorylation of STING to prevent sustained innate immune signaling. Celld155, 688–698.
Li, Q., Lin, L., Tong, Y., Liu, Y., Mou, J., Wang, X., Wang, X., Gong, Y., Zhao, Y, Liu, Y,et al. 2018. TRIM29 negatively controls antiviral immune response through targeting STING for degradation. Cell Discov.4, 13.
Lio, C.W., McDonald, B., Takahashi, M., Dhanwani, R., Sharma, N., Huang, J., Pham, E., Benedict, C.A., and Sharma, S. 2016. cGASSTING signaling regulates initial innate control of cytomegalo-virus infection. J. Virol.d90, 7789–7797.
Liu, S., Cai, X., Wu, J., Cong, Q., Chen, X., Li, T., Du, F., Ren, J., Wu, Y.T., Grishin, N.V.,et a.l 2015. Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science347, aaa2630.
Ma, Z. and Damania, B. 2016. The cGAS-STING defense pathway and its counteraction by viruses. Cell Host Microbed19, 150–158.
Motwani, M., Pesiridis, S., and Fitzgerald, K.A. 2019. DNA sensing by the cGAS-STING pathway in health and disease. Nat. Rev. Genet.d20, 657–674.
Paijo, J., Doring, M., Spanier, J., Grabski, E., Nooruzzaman, M., Schmidt, T., Witte, G., Messerle, M., Hornung, V., Kaever, V.,et a.l 2016. cGAS senses human cytomegalovirus and induces type I interferon responses in human monocyte-derived cells. PLoS Pathog.12, e1005546.
Prabakaran, T., Bodda, C., Krapp, C., Zhang, B.C., Christensen, M.H., Sun, C., Reinert, L., Cai, Y., Jensen, S.B., Skouboe, M.K.,et al. 2018. Attenuation of cGAS-STING signaling is mediated by a p62/SQSTM1-dependent autophagy pathway activated by TBK1. EMBO J.37, e97858.
Sanchez, V., Clark, C.L., Yen, J.Y., Dwarakanath, R., and Spector, D.H. 2002. Viable human cytomegalovirus recombinant virus with an internal deletion of the IE2 86 gene affects late stages of viral re-plication. J. Virol.d76, 2973–2989.
Sommer, M.H., Scully, A.L., and Spector, D.H. 1994. Transactivation by the human cytomegalovirus IE2 86-kilodalton protein requires a domain that binds to both the TATA box-binding protein and the retinoblastoma protein. J. Virol.d68, 6223–6231.
Srikanth, S., Woo, J.S., Wu, B., El- Sherbiny, Y.M., Leung, J., Chupradit, K., Rice, L., Seo, G.J., Calmettes, G., Ramakrishna, C.,et al. 2019. The Ca2+ sensor STIM1 regulates the type I interferon response by retaining the signaling adaptor STING at the endo-plasmic reticulum. Nat. Immunol.d20, 152–162.
Stinski, M.F. and Petrik, D.T. 2008. Functional roles of the human cytomegalovirus essential IE86 protein. Curr. Top. Microbiol. Immunol.d325, 133–152.
Taylor, R.T. and Bresnahan, W.A. 2005. Human cytomegalovirus immediate-early 2 gene expression blocks virus-induced beta interferon production. J. Virol.d79, 3873–3877.
Taylor, R.T. and Bresnahan, W.A. 2006a. Human cytomegalovirus IE86 attenuates virus- and tumor necrosis factor alpha-induced NFκB-dependent gene expression. J. Virol.d80, 10763–10771.
Taylor, R.T. and Bresnahan, W.A. 2006b. Human cytomegalovirus immediate-early 2 protein IE86 blocks virus-induced chemokine expression. J. Virol.d80, 920–928.
Wang, Y., Lian, Q., Yang, B., Yan, S., Zhou, H., He, L., Lin, G., Lian, Z., Jiang, Z., and Sun, B. 2015. TRIM30α is a negative-feedback regulator of the intracellular DNA and DNA virus-triggered response by targeting STING. PLoS Pathog.11, e1005012.
Weekes, M.P., Tomasec, P., Huttlin, E.L., Fielding, C.A., Nusinow, D., Stanton, R.J., Wang, E.C., Aicheler, R., Murrell, I., Wilkinson, G.W.,et a.l 2014. Quantitative temporal viromics: an approach to investigate host-pathogen interaction. Celld157, 1460–1472.
Yu, C.Y., Chang, T.H., Liang, J.J., Chiang, R.L., Lee, Y.L., Liao, C.L., and Lin, Y.L. 2012. Dengue virus targets the adaptor protein MITA to subvert host innate immunity. PLoS Pathog.8, e1002780.
Zhong, B., Zhang, L., Lei, C., Li, Y., Mao, A.P., Yang, Y., Wang, Y.Y., Zhang, X.L., and Shu, H.B. 2009. The ubiquitin ligase RNF5 regulates antiviral responses by mediating degradation of the adaptor protein MITA. Immunityd30, 397–407.
Acknowledgments
This research was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education, Science and Tech-nology (NRF-2017R1D1A1B03032494). This work was also supported by the Gachon University research fund of 2018 (GCU-2018-0362).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lee, JK., Kim, JE., Park, B.J. et al. Human cytomegalovirus IE86 protein aa 136–289 mediates STING degradation and blocks the cGAS-STING pathway. J Microbiol. 58, 54–60 (2020). https://doi.org/10.1007/s12275-020-9577-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-020-9577-6