Abstract
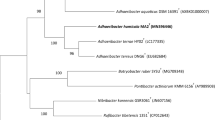
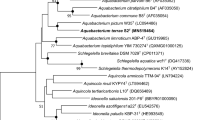
A white-coloured bacterium, SGM1-15T, was isolated from a paddy soil sample from Suwon, Republic of Korea. The cells were strictly aerobic, Gram-negative and curved rod-shaped. A phylogenetic analysis based on 16S rRNA gene sequences revealed that strain SGM1-15T was closely related to Curvibacter delicatus LMG 4328T (97.6% similarity) and Caenimonas koreensis EMB320T (97.5% similarity). The major respiratory quinone system was Q-8 and the predominant cellular fatty acids were C16:0 (39.9%), summed feature 3 (C16:1 ω7c and/or iso-C15:0 2-OH; 24.3%) and C17:0 cyclo (22.7%). The major polar lipids were diphosphatidylglycerol, phosphatidylglycerol and phosphatidylethanolamine. The major polyamines were 2-hydroxypurescine, purescine and spermidine. The DNA G+C content was 68.7 mol%. On the basis of the phylogenetic, physiologicl and chemotaxonomic data, stain SGM1-15T represents a novel species of the genus Caenimonas, for which the name Caenimonas terrae sp. nov. is proposed. The type strain of Caenimonas terrae is SGM1-15T (=KACC 13365T =NBRC 106341T).
Similar content being viewed by others
References
Breznak, J.A. and Costilow, R.N. 1994. Physicochemical factors in growth. Methods for General and Molecular Bacteriology, pp. 137–154. In Gerhardt, P., Murray, R.G.E., Wood, W.A., and Krieg, N.R. (eds.) American Society for Microbiology, Washington, D.C., USA.
Busse, H.J. and Auling, G. 1988. Polyamine pattern as a chemotaxonomic marker within the Proteobacteria. Syst. Appl. Microbiol. 11, 1–8.
Busse, H.J., Bunka, S., Hensel, A., and Lubitz, W. 1997. Discrimination of members of the family Pasteurellaceae based on polyamine patterns. Int. J. Syst. Bacteriol. 47, 698–708.
Chun, J., Lee, J.H., Jung, Y., Kim, M., Kim, S., Kim, B.K., and Lim, Y.W. 2007. EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequences. Int. J. Syst. Evol. Microbiol. 57, 2259–2261.
Groth, I., Schumann, P., Weiss, N., Martin, K., and Rainey, F.A. 1996. Agrococcus jenensis gen. nov., sp. nov., a new genus of actinomycetes with diaminobutyric acid in the cell wall. Int. J. Syst. Bacteriol. 46, 234–239.
Kämpfer, P., Steiof, M., and Dott, W. 1991. Microbiological characterization of a fuel-oil contaminated site including numerical identification of heterotrophic water and soil bacteria. Microb. Ecol. 21, 227–251.
Ludwig, W., Strunk, O., Westram, R., Richter, L., Meier, H., Yadhukumar, Buchner, A., Lai, T., Steppi, S., Jobb, G., and et al. 2004. ARB: a software environment for sequence data. Nucleic Acids Res. 32, 1363–1371.
Mesbah, M., Premachandran, U., and Whitman, W.B. 1989. Precise measurement of the G+C content of deoxyribonucleic-acid by high-performance liquid-chromatography. Int. J. Syst. Bacteriol. 39, 159–167.
Minnikin, D.E., O’Donnell, A.G., Goodfellow, M., Alderson, G., Athalye, M., Schaal, A., and Parlett, J.H. 1984. An integrated procedure for the extraction of isoprenoid quinines and polar lipids. J. Microbiol. Methods 2, 233–241.
Pruesse, E., Quast, C., Knittel, K., Fuchs, B.M., Ludwig, W., Peplies, J., and Glöckner, F.O. 2007. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acid Res. 35, 7188–7196.
Ryu, S.H., Lee, D.S., Park, M., Wang, Q., Jang, H.H., Park, W., and Jeon, C.O. 2008. Caenimonas koreensis gen. nov., sp nov., isolated from activated sludge. Int. J. Syst. Evol. Microbiol. 58, 1064–1068.
Seldin, L. and Dubnau, D. 1985. Deoxyribonucleic acid homology among Bacillus polymyxa, Bacillus macerans, Bacillus azotofixans, and other nitrogen-fixing Bacillus strains. Int. J. Syst. Bacteriol. 35, 151–154.
Smibert, R.M. and Krieg, N.R. 1994. Phenotypic characterization. Methods for General and Molecular Bacteriology, pp. 607–654. In Gerhardt, P., Murray, R.G.E., Wood, W.A., and Krieg, N.R. (eds.) American Society for Microbiology, Washington, D.C., USA.
Stackerbrandt, E, Murray, R.G.E., and Trüper, H.G. 1988. Proteobacteria classis nov., a name for the phylogenetic taxon that includes the “purple bacteria and their relatives”. Int. J. Syst. Bacteriol. 38, 321–325.
Wayne, L.G., Brenner, D.J., Colwell, R.R., Grimont, P.A.D., Kandler, O., Krichevsky, M.I., Moore, L.H., Moore, W.E.C., Murray, R.G.E., Stackebrandt, E., and et al. 1987. International Committee on Systematic Bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int. J. Syst. Bacteriol. 37, 463–464.
Weon, H.Y., Kim, B.Y., Yoo, S.H., Lee, S.Y., Kwon, S.W., Go, S.J., and Stackebrandt, E. 2006. Niastella koreensis gen. nov., sp. nov. and Niastella yeongjuensis sp. nov., novel members of the phylum Bacteroidetes, isolated from soil cultivated with Korean ginseng. Int. J. Syst. Evol. Microbiol. 56, 1777–1782.
Author information
Authors and Affiliations
Corresponding author
Additional information
The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequences of strain SGM1-15T is GU181268.
Rights and permissions
About this article
Cite this article
Kim, SJ., Weon, HY., Kim, YS. et al. Caenimonas terrae sp. nov., isolated from a soil sample in Korea, and emended description of the genus Caenimonas Ryu et al. 2008. J Microbiol. 50, 864–868 (2012). https://doi.org/10.1007/s12275-012-1587-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-012-1587-6