Abstract
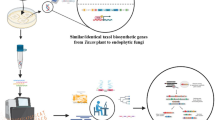
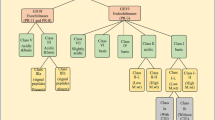
Chitin synthase (CHS) is a glucosyltransferase that converts UDP-N-acetylglucosamine into chitin, one of the main components of fungal cell wall. Class III chitin synthases act directly in the formation of the cell wall. They catalyze the conversion of the immediate precursor of chitin and are responsible for the majority of chitin synthesis in fungi. As such, they are highly specific molecular targets for drugs that can inhibit the growth and development of fungal pathogens. In this work, we have identified and characterized a chitin synthase gene of Moniliophthora perniciosa (Mopchs) by primer walking. The complete gene sequence is 3,443 bp, interrupted by 13 small introns, and comprises a cDNA with an ORF with 2,739 bp, whose terminal region was experimentally determined, encoding a protein with 913 aa that harbors all the motifs and domains typically found in class III chitin synthases. This is the first report on the characterization of a chitin synthase gene, its mature transcription product, and its putative protein in basidioma and secondary mycelium stages of M. perniciosa, a basidiomycotan fungus that causes witches’ broom disease of cacao.
Similar content being viewed by others
References
Aime, M.C. and W. Phillips-Mora. 2005. The causal agents of witches broom and frosty pod rot of cacao (chocolate, Theobroma cacao) form a new lineage of Marasmiaceae. Mycologia 97, 1012–1022.
Altschul, S.F., T.L. Madden, A.A. Schäffer, J. Zhang, Z. Zhang, W. Miller, and D.J. Lipman. 1997. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402.
Bago, B., H. Chamberland, A. Goulet, H. Vierheilig, J.G. Lafontaine, and E. Piche. 1996. Effect of nikkomycin Z, a chitin synthase inhibitor, on hyphal growth and cell wall structure of two arbuscular-mycorrhizal fungi. Protoplasma 192, 80–92.
Behr, J.B. 2003. Chitin synthase as an antifungal target: recent advances. Curr. Med. Chem. 2, 173–189.
Birney, E., M. Clamp, and R. Durbin. 2004. GeneWise and Genome-Wise. Genome Res. 14, 988–995.
Birren, B., E. Lander, J. Galagan, C. Nusbaum, K. Devon, L.J. Ma, D. Jaffe, J. Butler, P. Alvarez, S. Gnerre, M. Grabherr, M. Kleber, E. Mauceli, W. Brockman, S. Rounsley, S. Young, K. LaButti, V. Pushparaj, D. DeCaprio, M. Crawford, et al. 2003. Annotation of the Coprinopsis cinerea genome. The Broad Institute Genome Sequencing Platform. (http://www.ncbi.nlm.nih.gov/entrez/viewer.fcgi?db=protein&id=116504282) and (http://www.ncbi.nlm.nih.gov/entrez/viewer.fcgi?db=116501858) Accessed in: 26 Jul 2007.
Bowman, S.M. and S.J. Free. 2006. The structure and synthesis of the fungal cell wall. Bioessays 28, 799–808.
Broeker, K., S. Fehser, and B.M. Moerschbacher. 2006. Survey and expression analysis of five new chitin synthase genes in the biotrophic rust fungus Puccinia graminis. Curr. Genet. 50, 295–305.
Bulawa, C.E., M. Slater, E. Cabib, J. Au-Young, A. Sburlati, W.L. Adair, and P.W. Robbins. 1986. The S. cerevisiae structural gene for chitin synthase is not required for chitin synthesis in vivo. Cell 46, 213–225.
Burland, T.G. 2000. DNASTAR’s lasergene sequence analysis software. Methods Mol. Biol. 132, 71–91.
Choquer, M., M. Boccara, I.R. Goncalves, M.C. Soulie, and A. Vidal-Cros. 2004. Survey of the Botrytis cinerea chitin synthase multigenic family through the analysis of six euascomycetes genomes. Eur. J. Biochem. 271, 2153–2164.
Doyle, J.J. and J.L. Doyle. 1987. A rapid DNA isolation procedure for small amounts of fresh leaf tissue. Phytochem. Bull. 19, 11–15.
Durán, A., B. Bowers, and E. Cabib. 1975. Chitin synthetase zymogen is attached to the yeast plasma membrane. Proc. Natl. Acad. Set USA 72, 3952–3955.
Fayyad, U.M. 1996. Advances in Knowledge Discovery and Data Mining. AAI Press, Cambridge, UK.
Felsenstein, J. 2005. PHYLIP (Phylogeny Inference Package) version 3.6. Distributed by the author. Department of Genome Sciences, University of Washington, Seattle, USA.
Formighieri, E.F., R.A. Tiburcio, E.D. Armas, F.J. Medrano, H. Shimo, N. Carels, A. Góes-Neto, C. Cotomacci, M.F. Carazzolle, N. Sardinha-Pinto, J. Rincones, L. Digiampietri, D.M. Carraro, A.M. Azeredo-Espin, S.F. Reis, A.C. Deckmann, K. Gramacho, M.S. Gonçalves, J.P. Moura Neto, L.V. Barbosa, et al. 2008. The mitochondrial genome of the phytopathogenic basidiomycete Moniliophthora perniciosa is 109 kb in size and contains a stable integrated plasmid. Mycol. Res. 112, 1136–1152.
Fung, E., R.W. Hyman, D. Rowley, D. Bruno, M. Miranda, M. Fukushima, B.L. Wickes, J. Fu, and R.W. Davis. 2004. Cryptococcus neoformans serotype D sequencing. (http://www.ncbi.nlm.nih.gov/entrez/viewer.fcgi?db=protein&id=50257918). Accessed in: 26 Jul 2007.
Glaser, L. and D.H. Brown. 1957. The synthesis of chitin in cell-free extracts of Neurospora crassa. J. Biol. Chem. 228, 729–742.
Green, P. 2007. Phrap Documentation: Algorithms. Phred/Phrap/Consed System Home Page. (http://www.phrap.org) Accessed in: 2 May 2007.
Griffith, G.W., J. Nicholson, A. Nenninger, and R.N. Birch. 2003. Witches’ brooms and frosty pods: two major pathogens of cacao. New Zeal. J. Bot. 41, 423–435.
James, T.Y., F. Kauff, C.L. Schoch, P.B. Matheny, V. Hofstetter, C.J. Cox, G. Celio, C. Gueidan, E. Fraker, J. Miadlikowska, H.T. Lumbsch, A. Rauhut, V. Reeb, A.E. Arnold, A. Amtoft, J.E. Stajich, K. Hosaka, G.H. Sung, D. Johnson, B. O’Rourke, et al. 2006. Reconstructing the early evolution of Fungi using a six-gene phylogeny. Nature 443, 818–822.
Kamper, J., R. Kahmann, M. Bolker, L.J. Ma, T. Brefort, B.J. Saville, F. Banuett, J.W. Kronstad, S.E. Gold, O. Muller, M.H. Perlin, H.A. Wosten, R. De Vries, J. Ruiz-Herrera, C.G. Reynaga-Pena, K. Snetselaar, M. Mccann, J. Perez-Martin, M. Feldbrugge, C.W. Basse, et al. 2006. Insights from the genome of the biotrophic fungal plant pathogen Ustilago maydis. Nature 444, 97–101.
Krogh, A., B. Larsson, G. von Heijne, and E.L. Sonnhammer. 2001. Predicting transmembrane protein topology with a hidden Markov model: Application to complete genomes. J. Mol. Biol. 305, 567–580.
Latgé, J.P. 2007. The cell wall: A carbohydrate armour for the fungal cell. Mol. Microbiol. 66, 279–290.
Lewis, D.H. 1991. Fungi and sugars — a suite of interactions. Mycol. Res. 95, 897–904.
Loftus, B.J., E. Fung, P. Roncaglia, D. Rowley, P. Amedeo, D. Bruno, J. Vamathevan, M. Miranda, I.J. Anderson, J.A. Fraser, J.E. Allen, I.E. Bosdet, M.R. Brent, R. Chiu, T.L. Doering, M.J. Donlin, C.A. D’souza, D.S. Fox, V. Grinberg, J. Fu, et al. 2005. The genome of the basidiomycetous yeast and human pathogen Cryptococcus neoformans. Science 307, 1321–1324.
Lopes, M.A., D.S. Gomes, M.G.B. Koblitz, C.P. Pirovani, J.C. de M. Cascardo, A. Góes-Neto, and F. Micheli. 2008. Use of response surface methodology to examine chitinase regulation in the basidiomycete Moniliophthora perniciosa. Mycol Res. 112, 399–406.
Merzendorfer, H. 2006. Insect chitin synthases: a review. J. Comp. Physiol. B. 176, 1–15.
Merzendorfer, H. and L. Zimoch. 2003. Chitin metabolism in insects: Structure, function and regulation of chitin synthases and chitinases. J. Exp. Biol. 206, 4393–4412.
Mondego, J.M., M.F. Carazzolle, G.G. Costa, E.F. Formighieri, L.P. Parizzi, J. Rincones, C. Cotomacci, D.M. Carraro, A.F. Cunha, H. Carrer, R.O. Vidal, R.C. Estrela, O. García, D.P. Thomazella, B.V. de Oliveira, A.B. Pires, M.C. Rio, M.R. Araújo, M.H. de Moraes, L.A. Castro, et al. 2008. A genome survey of Moniliophthora perniciosa gives new insights into witches’ broom disease of cacao. BMC Genomics 9, 548.
Mulder, N. and R. Apweiler. 2007. InterPro and InterProScan: Tools for protein sequence classification and comparison. Methods Mol. Biol. 396, 59–70.
Nagahashi, S., M. Sudoh, N. Ono, R. Sawada, E. Yamaguchi, Y. Uchida, T. Mio, M. Takagi, M. Arisawa, and H. Yamada-Okabe. 1995. Characterization of chitin synthase 2 of Saccharomyces cerevisiae. Implication of two highly conserved domains as possible catalytic sites. J. Biol. Chem. 270, 13961–13967.
Nishihara, M., A. Watanabe, and Y. Asada. 2007. Isolation, characterization, and expression analysis of a class IV chitin synthase gene from the edible basidiomycetous mushroom Pleurotus ostreatus. Mycoscience 48, 176–181.
Notredame, C., D. Higgins, and J. Heringa. 2000. T-Coffee: A novel method for multiple sequence alignments. J. Mol. Biol. 302, 205–217.
Page, R.D.M. 1996. Treeview: An application to display phylogenetic trees on personal computers. Comput. Appl. Biosci. 12, 357–358.
Parker, J.D., P.S. Rabinovitch, and G.C. Burmer. 1991. Targeted gene walking polymerase chain reaction. Nucleic Acids Res. 19, 3055–3060.
Pearson, W.R. 1990. Rapid and sensitive sequence comparison with FASTP and FASTA. Methods Enzymol. 183, 63–98.
Pirovani, C.P., B.T. da Hora-Júnior, B.M. Oliveira, M.A. Lopes, C.V. Dias, S.H. Cruz da, A. Schriefer, J.C. de M. Cascardo, G.A.G. Pereira, and A. Góes-Neto. 2005. Knowledge discovery in genome database: the chitin metabolic pathway in Crinipellis perniciosa (Stahel) Singer, vol. 1, p. 122–139. In R. Mondaini (ed.), Proceedings of IV Brazilian Symposium on Mathematical and Computational Biology / I International Symposium on Mathematical and Computational Biology. E-Papers Serviços Editoriais LTDA, Rio de Janeiro, Brazil.
Rincones, J., G.D. Mazotti, G.W. Griffith, A. Pomela, A. Figueira, G.A. Leal, Jr., M.V. Queiroz, J.F. Pereira, R.A. Azevedo, G.A.G. Pereira, and L.W. Meinhardt. 2006. Genetic variability and chromosome-length polymorphisms of the witches’ broom pathogen Crinipellis perniciosa from various plant hosts in South America. Mycol. Res. 110, 821–832.
Roncero, C. 2002. The genetic complexity of chitin synthesis in fungi. Curr. Genet. 41, 367–378.
Ronquist, F.R. and J.P. Huelsenbeck. 2003. MrBayse 3: Bayesian phylogenetic, inference under mixed models. Bioinformatics 19, 1572–1574.
Ruiz-Herrera, J., J.M. González-Prieto, and R. Ruiz-Medrano. 2002. Evolution and phylogenetic relationships of chitin synthases from yeast and fungi. FEMS Yeast Res. 1, 247–256.
Smale, S.T. and J.T. Kadonaga. 2003. The RNA polymerase II core promoter. Annu. Rev. Biochem. 72, 449–479.
Sreenivasaprasad, S., K.S. Burton, and D.A. Wood. 2000. Cloning and characterisation of a chitin synthase gene cDNA from the cultivated mushroom Agaricus bisporus and its expression during morphogenesis. FEMS Microbiol. Lett. 189, 73–77.
Stanke, M. and S. Waack. 2003. Gene prediction with a Hidden-Markov Model and a new intron submodel. Bioinformatics 19,Suppl. 2, ii215–ii225.
Swofford, D.L. 2002. PAUP phylogenetic analysis using parsimony and other methods, version 4.0b 10. Sinauer, Sunderland.
Wang, K., D.W. Ussery, and S. Brunak. 2008. Analysis and prediction of gene splice sites in four Aspergillus genomes. Fungal Genet. Biol., in press.
Weber, I., D. Assmann, E. Thines, and G. Steinberg. 2006. Polar localizing class V myosin chitin synthases are essential during early plant infection in the plant pathogenic fungus Ustilago maydis. Plant Cell 18, 225–242.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Souza, C.S., Oliveira, B.M., Costa, G.G.L. et al. Identification and characterization of a class III chitin synthase gene of Moniliophthora perniciosa, the fungus that causes witches’ broom disease of cacao. J Microbiol. 47, 431–440 (2009). https://doi.org/10.1007/s12275-008-0166-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-008-0166-3