Abstract
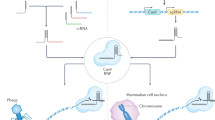
Genome editing is a useful tool in basic and clinical research. Among the several approaches used in genome editing, the CRISPR–Cas9 system using clustered regularly interspaced short palindromic repeats (CRISPR) and CRISPR-associated protein 9 (Cas9) along with a guide RNA has been developed recently. The CRISPR/Cas9 system induces site-specific double-stranded DNA breaks, which result in DNA repair via non-homologous end joining (NHEJ) or homology-directed repair (HDR). However, HDR efficiency is lower than that of NHEJ and accordingly poses a challenge in genome modification studies. Several chemical compounds including RS-1 have been shown to enhance the HDR knock-in process by two- to six-fold in HEK 293 cells and rabbit embryos. Based on this finding, we developed an antibiotic resistance system to screen RS-1 chemical derivatives, which may promote efficient HDR. In this study, we report several chemical compounds with high knock-in efficiency at the ATG5 gene locus, using HeLa cell-based assays.



Similar content being viewed by others
References
Cho SW, Kim S, Kim JM, Kim JS (2013) Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat Biotechnol 31:230–232. https://doi.org/10.1038/nbt.2507
Hsu PD, Lander ES, Zhang F (2014) Development and applications of CRISPR-Cas9 for genome engineering. Cell 157:1262–1278. https://doi.org/10.1016/j.cell.2014.05.010
Hu Z, Shi Z, Guo X, Jiang BS, Wang G, Luo DX, Chen YL, Zhu YS (2018) Ligase IV inhibitor SCR7 enhances gene editing directed by CRISPR-Cas9 and ssODN in human cancer cells. Cell Biosci 8:12. https://doi.org/10.1186/s13578-018-0200-z
Jayathilaka K, Sheridan SD, Bold TD, Bochenska K, Logan HL, Weichselbaum RR, Bishop DK, Connell PP (2008) A chemical compound that stimulates the human homologous recombination protein RAD51. Proc Natl Acad Sci 105:15848–15853. https://doi.org/10.1073/pnas.0808046105
Jeon IS, Kim HR, Shin EY, Kim EG, Han HS, Hong JT, Lee HK, Song KD, Choi JK (2018) Modulation of store-operated calcium entry and nascent adhesion by p21-activated kinase 1. Exp Mol Med 50:65. https://doi.org/10.1038/s12276-018-0093-2
Kim HJ, Lee HJ, Kim H, Cho SW, Kim JS (2009) Targeted genome editing in human cells with zinc finger nucleases constructed via modular assembly. Genome Res 19:1279–1288. https://doi.org/10.1101/gr.089417.108
Knott GJ, Doudna JA (2018) CRISPR-Cas guides the future of genetic engineering. Science 361:866–869. https://doi.org/10.1126/science.aat5011
Lee SH, Kim S, Hur JK (2018) CRISPR and target-specific DNA endonucleases for efficient DNA knock-in in eukaryotic genomes. Mol Cells 41:943–952. https://doi.org/10.14348/molcells.2018.0408
Li GL, Zhang XW, Zhong CL, Mo JX, Quan R, Yang J, Liu DW, Li ZC, Yang HQ, Wu ZF (2017) Small molecules enhance CRISPR/Cas9-mediated homology-directed genome editing in primary cells. Sci Rep 7:8943. https://doi.org/10.1038/s41598-017-09306-x
Maruyama T, Dougan SK, Truttmann MC, Bilate AM, Ingram JR, Ploegh HL (2015) Increasing the efficiency of precise genome editing with CRISPR-Cas9 by inhibition of nonhomologous end joining. Nat Biotechnol 33:538–542. https://doi.org/10.1038/nbt.3190
Mason JM, Logan HL, Budke B, Wu M, Pawlowski M, Weichselbaum RR, Kozikowski AP, Bishop DK, Connell PP (2014) The RAD51-stimulatory compound RS-1 can exploit the RAD51 overexpression that exists in cancer cells and tumors. Cancer Res 74:3546–3555. https://doi.org/10.1158/0008-5472.CAN-13-3220
O’Brien KA, Atkinson RA, Richardson L, Koulman A, Murray AJ, Harridge SDR, Martin DS, Levett DZH, Mitchell K, Mythen MG, Montgomery HE, Grocott MW, Griffin JL, Edwards LM (2019) Metabolomic and lipidomic plasma profile changes in human participants ascending to Everest Base Camp. Sci Rep 9:2297. https://doi.org/10.1038/s41598-019-38832-z
Paquet D, Kwart D, Chen A, Sproul A, Jacob S, Teo S, Olsen KM, Gregg A, Noggle S, Tessier-Lavigne M (2016) Efficient introduction of specific homozygous and heterozygous mutations using CRISPR/Cas9. Nature 533:125–129. https://doi.org/10.1038/nature17664
Pinder J, Salsman J, Dellaire G (2015) Nuclear domain ‘knock-in’ screen for the evaluation and identification of small molecule enhancers of CRISPR-based genome editing. Nucleic Acids Res 43:9379–9392. https://doi.org/10.1093/nar/gkv993
Rouet P, Smih F, Jasin M (1994) Expression of a site-specific endonuclease stimulates homologous recombination in mammalian cells. Proc Natl Acad Sci 91:6064–6068. https://doi.org/10.1073/pnas.91.13.6064
Song J, Yang DS, Xu J, Zhu TQ, Chen YE, Zhang JF (2016) RS-1 enhances CRISPR/Cas9- and TALEN-mediated knock-in efficiency. Nat Commun 7:10548. https://doi.org/10.1038/ncomms10548
Yu C, Liu YX, Ma TH, Liu K, Xu SH, Zhang Y, Liu HL, La Russa ML, Xie M, Ding S, Lei SQ (2015) Small molecules enhance CRISPR genome editing in pluripotent stem cells. Cell Stem Cell 16:142–147. https://doi.org/10.1016/j.stem.2015.01.003
Zhang JP, Li XL, Li GH, Chen W, Arakaki C, Botimer GD, Baylink D, Zhang L, Wen W, Fu YW, Xu J, Chun N, Yuan WP, Cheng T, Zhang XB (2017) Efficient precise knockin with a double cut HDR donor after CRISPR/Cas9-mediated double-stranded DNA cleavage. Genome Biol 18(1):35. https://doi.org/10.1186/s13059-017-1164-8
Acknowledgements
This work was supported by (1) Next Generation BioGreen21 project (Project Nos. PJ01334802 & PJ01315101) of the Rural Development Administration, Korea, (2) Ministry of Education and the National Research Foundation of Korea (NRF-2017S1A5B8059946), and (3) Korea Ministry of Environment (MOE) and Korea Environment Industry & Technology Institute (KEITI) as a "Technology Program for establishing biocide safety management” [2018002490003]. The chemical library used in this study was kindly provided by Korea Chemical Bank (http://www.chembank.org/) of KRICT.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
We declare no actual or potential conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jeon, IS., Shin, JC., Kim, S.R. et al. Role of RS-1 derivatives in homology-directed repair at the human genome ATG5 locus. Arch. Pharm. Res. 43, 639–645 (2020). https://doi.org/10.1007/s12272-020-01226-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12272-020-01226-1