Abstract
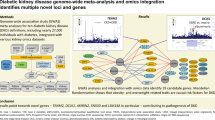
Diabetic kidney disease (DKD) is a microvascular complication of type 1 and 2 diabetes with a devastating impact on individuals with the disease, their families, and society as a whole. DKD is the single most frequent cause of incident chronic kidney disease cases and accounts for over 40 % of the population with end-stage renal disease. Contributing factors for the high prevalence are the increase in obesity and subsequent diabetes combined with an improved long-term survival with diabetes. Environment and genetic variations contribute to DKD susceptibility and progressive loss of kidney function. How the molecular mechanisms of genetic and environmental exposures interact during DKD initiation and progression is the focus of ongoing research efforts. The development of standardized, unbiased high-throughput profiling technologies of human DKD samples opens new avenues in capturing the multiple layers of DKD pathobiology. These techniques routinely interrogate analytes on a genome-wide scale generating comprehensive DKD-associated fingerprints. Linking the molecular fingerprints to deep clinical phenotypes may ultimately elucidate the intricate molecular interplay in a disease stage and subtype-specific manner. This insight will form the basis for accurate prognosis and facilitate targeted therapeutic interventions. In this review, we present ongoing efforts from large-scale data integration translating “-omics” research efforts into improved and individualized health care in DKD.

Similar content being viewed by others
Abbreviations
- GWAS:
-
Genome-wide association study
- SNP:
-
Single nucleotide polymorphism
- CNV:
-
Copy number variation
- NF-κB:
-
Nuclear factor κ-light chain enhancer of activated B cells
- JAK/STAT:
-
Janus kinase/signal transducer and activator of transcription
- MCP-1/CCL2:
-
Macrophage chemoattractant protein 1
- FDA:
-
Food and Drug Administration
- DCCT/EDIC:
-
The Diabetes Control and Complications Trial/Epidemiology of Diabetes Interventions and Complications Study
- UKPDS:
-
The UK Prospective Diabetes Study
- ERCB:
-
European Renal cDNA Bank–Kröner–Fresenius Biopsy Bank
- MIAME:
-
Minimum Information About a Microarray Experiment
References
Wild, S., Roglic, G., Green, A., Sicree, R., & King, H. (2004). Global prevalence of diabetes. Diabetes Care, 27(5), 1047–1053.
Shaw, J. E., Sicree, R. A., & Zimmet, P. Z. (2010). Global estimates of the prevalence of diabetes for 2010 and 2030. Diabetes Res Clin Pract, 87(1), 4–14.
Coresh, J., Selvin, E., Stevens, L. A., Manzi, J., Kusek, J. W., Eggers, P., Van Lente, F., & Levey, A. S. (2007). Prevalence of chronic kidney disease in the United States. JAMA, 298(17), 2038–2047.
Foley, R., & Collins, A. (2009). The growing economic burden of diabetic kidney disease. Curr Diab Rep, 9(6), 460–465.
The Australia and New Zealand Dialysis and Transplant Registry (2011) The 34th Annual ANZDATA Report 2011—Data to 2010. At: http://wwwanzdataorgau/v1/report_2011html. Accessed March 30, 2012.
European Renal Association—European Dialysis and Transplant (ERA-EDTA) Registry (2011) ERA-EDTA registry annual report 2009. At: http://wwwera-edta-regorg/files/annualreports/pdf/AnnRep2009_newpdf. Accessed March 30, 2012
US Renal Data System (2011) USRDS 2011 annual data report: atlas of chronic kidney disease and end-stage renal disease in the United States. National Institutes of Health, National Institute of Diabetes and Digestive and Kidney Diseases, Bethesda, MD. At: http://www.usrds.org. Accessed 22 March 2012.
de Boer, I. H., Rue, T. C., Hall, Y. N., Heagerty, P. J., Weiss, N. S., & Himmelfarb, J. (2011). Temporal trends in the prevalence of diabetic kidney disease in the United States. JAMA, 305(24), 2532–2539.
Haffner, S. M., Lehto, S., Rönnemaa, T., Pyörälä, K., & Laakso, M. (1998). Mortality from coronary heart disease in subjects with type 2 diabetes and in nondiabetic subjects with and without prior myocardial infarction. New Engl J Med, 339(4), 229–234.
Valmadrid, C. T., Klein, R., Moss, S. E., & Klein, B. E. K. (2000). The risk of cardiovascular disease mortality associated with microalbuminuria and gross proteinuria in persons with older-onset diabetes mellitus. Arch Intern Med, 160(8), 1093–1100.
Sarnak, M. J., Levey, A. S., Schoolwerth, A. C., Coresh, J., Culleton, B., Hamm, L. L., McCullough, P. A., Kasiske, B. L., Kelepouris, E., Klag, M. J., Parfrey, P., Pfeffer, M., Raij, L., Spinosa, D. J., & Wilson, P. W. (2003). Kidney disease as a risk factor for development of cardiovascular disease. Hypertension, 42(5), 1050–1065.
Brownlee, M. (2001). Biochemistry and molecular cell biology of diabetic complications. Nature, 414(6865), 813–820.
Forbes, J. M., Fukami, K., & Cooper, M. E. (2007). Diabetic nephropathy: where hemodynamics meets metabolism. Exp Clin Endocrinol Diabetes, 115(2), 69–84.
Figueroa-Romero, C., Sadidi, M., & Feldman, E. (2008). Mechanisms of disease: the oxidative stress theory of diabetic neuropathy. Rev Endocr Metab Disord, 9(4), 301–314.
Berthier, C. C., Zhang, H., Schin, M., Henger, A., Nelson, R. G., Yee, B., Boucherot, A., Neusser, M. A., Cohen, C. D., Carter-Su, C., Argetsinger, L. S., Rastaldi, M. P., Brosius, F. C., & Kretzler, M. (2009). Enhanced expression of Janus kinase and signal transducer and activator of transcription pathway members in human diabetic nephropathy. Diabetes, 58(2), 469–477.
Navarro, J. F., & Mora-Fernández, C. (2006). The role of TNF-α in diabetic nephropathy: pathogenic and therapeutic implications. Cytokine Growth Factor Rev, 17(6), 441–450.
Williams, M. E. (2005). Diabetic nephropathy: the proteinuria hypothesis. Am J Nephrol, 25(2), 77–94.
Schena, F. P., & Gesualdo, L. (2005). Pathogenetic mechanisms of diabetic nephropathy. J Am Soc Nephrol, 16(3_suppl_1), S30–S33.
Galkina, E., & Ley, K. (2006). Leukocyte recruitment and vascular injury in diabetic nephropathy. J Am Soc Nephrol, 17(2), 368–377.
Nawroth, P. P., & Isermann, B. (2010). Mechanisms of diabetic nephropathy—old buddies and newcomers part 2. Exp Clin Endocrinol Diabetes, 118(10), 667–672.
Nawroth, P. P., & Isermann, B. (2010). Mechanisms of diabetic nephropathy—old buddies and newcomers part 1. Exp Clin Endocrinol Diabetes, 118(9), 571–576.
Kanwar, Y. S., Sun, L., Xie, P., Liu, F.-Y., & Chen, S. (2011). A glimpse of various pathogenetic mechanisms of diabetic nephropathy. Annu Rev Pathol, 6(1), 395–423.
DiBona, G. F., & Kopp, U. C. (1997). Neural control of renal function. Physiol Rev, 77(1), 75–197.
Augustyniak, R. A., Tuncel, M., Zhang, W., Toto, R. D., & Victor, R. G. (2002). Sympathetic overactivity as a cause of hypertension in chronic renal failure. J Hypertens, 20(1), 3–9.
Grotendorst, G. R. (1997). Connective tissue growth factor: a mediator of TGF-β action on fibroblasts. Cytokine Growth Factor Rev, 8(3), 171–179.
Böttinger, E. P., & Bitzer, M. (2002). TGF-ß signaling in renal disease. J Am Soc Nephrol, 13(10), 2600–2610.
Liu, Y. (2006). Renal fibrosis: new insights into the pathogenesis and therapeutics. Kidney Int, 69(2), 213–217.
Froissart, M., Rossert, J., Jacquot, C., Paillard, M., & Houillier, P. (2005). Predictive performance of the modification of diet in renal disease and Cockcroft-Gault equations for estimating renal function. J Am Soc Nephrol, 16(3), 763–773.
Stevens, L. A., Coresh, J., Greene, T., & Levey, A. S. (2006). Assessing kidney function—measured and estimated glomerular filtration rate. New Engl J Med, 354(23), 2473–2483.
Stevens, L. A., & Levey, A. S. (2009). Measured GFR as a confirmatory test for estimated GFR. J Am Soc Nephrol, 20(11), 2305–2313.
Stevens, L. A., Schmid, C. H., Greene, T., Zhang, Y., Beck, G. J., Froissart, M., Hamm, L. L., Lewis, J. B., Mauer, M., Navis, G. J., Steffes, M. W., Eggers, P. W., Coresh, J., & Levey, A. S. (2010). Comparative performance of the CKD Epidemiology Collaboration (CKD-EPI) and the Modification of Diet in Renal Disease (MDRD) Study equations for estimating GFR levels above 60 mL/min/1.73 m2. Am J Kidney Dis, 56(3), 486–495.
Levey, A. S., Cattran, D., Friedman, A., Miller, W. G., Sedor, J., Tuttle, K., Kasiske, B., & Hostetter, T. (2009). Proteinuria as a surrogate outcome in CKD: report of a scientific workshop sponsored by the National Kidney Foundation and the US Food and Drug Administration. Am J Kidney Dis, 54(2), 205–226.
Julian, B. A., Suzuki, H., Suzuki, Y., Tomino, Y., Spasovski, G., & Novak, J. (2009). Sources of urinary proteins and their analysis by urinary proteomics for the detection of biomarkers of disease. Proteomics Clin Appl, 3(9), 1029–1043.
Caramori, M. L., Fioretto, P., & Mauer, M. (2000). The need for early predictors of diabetic nephropathy risk: is albumin excretion rate sufficient? Diabetes, 49(9), 1399–1408.
Perkins, B. A., Ficociello, L. H., Silva, K. H., Finkelstein, D. M., Warram, J. H., & Krolewski, A. S. (2003). Regression of microalbuminuria in type 1 diabetes. New Engl J Med, 348(23), 2285–2293.
Fioretto, P., & Mauer, M. (2007). Histopathology of diabetic nephropathy. Semin Nephrol, 27(2), 195–207.
American Diabetes Association. (2011). Standards of medical care in diabetes—2011. Diabetes Care, 34(Supplement 1), S11–S61.
Perkins, B. A., Ficociello, L. H., Ostrander, B. E., Silva, K. H., Weinberg, J., Warram, J. H., & Krolewski, A. S. (2007). Microalbuminuria and the risk for early progressive renal function decline in type 1 diabetes. J Am Soc Nephrol, 18(4), 1353–1361.
Perkins, B. A., Ficociello, L. H., Roshan, B., Warram, J. H., & Krolewski, A. S. (2010). In patients with type 1 diabetes and new-onset microalbuminuria the development of advanced chronic kidney disease may not require progression to proteinuria. Kidney Int, 77(1), 57–64.
MacIsaac, R., & Jerums, G. (2011). Diabetic kidney disease with and without albuminuria. Curr Opin Nephrol Hypertens, 20(3), 246–257.
Dwyer, J. P., Parving, H. H., Hunsicker, L. G., Ravid, M., Remuzzi, G., & Lewis, J. B. (2012). Renal dysfunction in the presence of normoalbuminuria in type 2 diabetes: results from the DEMAND study. Cardiorenal Medicine, 2(1), 1–10.
Pavkov, M. E., Knowler, W. C., Lemley, K. V., Mason, C. C., Myers, B. D., & Nelson, R. G. (2012). Early renal function decline in type 2 diabetes. Clin J Am Soc Nephrol, 7(1), 78–84.
Karalliedde, J., & Viberti, G. (2010). Proteinuria in diabetes: bystander or pathway to cardiorenal disease? J Am Soc Nephrol, 21(12), 2020–2027.
Tryggvason, K., & Pettersson, E. (2003). Causes and consequences of proteinuria: the kidney filtration barrier and progressive renal failure. J Intern Med, 254(3), 216–224.
Iyengar, S. K., Schelling, J. R., & Sedor, J. R. (2002). Approaches to understanding susceptibility to nephropathy: from genetics to genomics. Kidney Int, 61(Suppl. 1), S61–S67.
Iyengar, S. K., Abboud, H. E., Goddard, K. A. B., Saad, M. F., Adler, S. G., Arar, N. H., Bowden, D. W., Duggirala, R., Elston, R. C., Hanson, R. L., Ipp, E., Kao, W. H. L., Kimmel, P. L., Klag, M. J., Knowler, W. C., Meoni, L. A., Nelson, R. G., Nicholas, S. B., Pahl, M. V., Parekh, R. S., Quade, S. R. E., Rich, S. S., Rotter, J. I., Scavini, M., Schelling, J. R., Sedor, J. R., Sehgal, A. R., Shah, V. O., Smith, M. W., Taylor, K. D., Winkler, C. A., Zager, P. G., & Freedman, B. I. (2007). Genome-wide scans for diabetic nephropathy and albuminuria in multiethnic populations: The Family Investigation of Nephropathy and Diabetes (FIND). Diabetes, 56(6), 1577–1585.
Igo, J. R. P., Iyengar, S. K., Nicholas, S. B., Goddard, K. A. B., Langefeld, C. D., Hanson, R. L., Duggirala, R., Divers, J., Abboud, H., Adler, S. G., Arar, N. H., Horvath, A., Elston, R. C., Bowden, D. W., Guo, X., Ipp, E., Kao, W. H. L., Kimmel, P. L., Knowler, W. C., Meoni, L. A., Molineros, J., Nelson, R. G., Pahl, M. V., Parekh, R. S., Rasooly, R. S., Schelling, J. R., Shah, V. O., Smith, M. W., Winkler, C. A., Zager, P. G., Sedor, J. R., Freedman, B. I., & The Family Investigation of Nephropathy and Diabetes Research Group. (2011). Genomewide linkage scan for diabetic renal failure and albuminuria: the FIND study. Am J Nephrol., 33(5), 381–389.
Gohda, T., Tanimoto, M., Watanabe-Yamada, K., Matsumoto, M., Kaneko, S., Hagiwara, S., Shiina, K., Shike, T., Funabiki, K., & Tomino, Y. (2005). Genetic susceptibility to type 2 diabetic nephropathy in human and animal models. Nephrology, 10(Supplement s2), S22–S25.
Tanaka, N., & Babazono, T. (2005). Assessing genetic susceptibility to diabetic nephropathy. Nephrology, 10(Supplement s2), S17–S21.
Mueller, P. W., Rogus, J. J., Cleary, P. A., Zhao, Y., Smiles, A. M., Steffes, M. W., Bucksa, J., Gibson, T. B., Cordovado, S. K., Krolewski, A. S., Nierras, C. R., & Warram, J. H. (2006). Genetics of Kidneys in Diabetes (GoKinD) study: a genetics collection available for identifying genetic susceptibility factors for diabetic nephropathy in type 1 diabetes. J Am Soc Nephrol, 17(7), 1782–1790.
Freedman, B. I., Bostrom, M., Daeihagh, P., & Bowden, D. W. (2007). Genetic factors in diabetic nephropathy. Clin J Am Soc Nephrol, 2(6), 1306–1316.
Conway, B. R., & Maxwell, A. P. (2009). Genetics of diabetic nephropathy: are there clues to the understanding of common kidney diseases? Nephron Clin Pract, 112(4), c213–c221.
Pezzolesi, M. G., Poznik, G. D., Mychaleckyj, J. C., Paterson, A. D., Barati, M. T., Klein, J. B., Ng, D. P. K., Placha, G., Canani, L. H., Bochenski, J., Waggott, D., Merchant, M. L., Krolewski, B., Mirea, L., Wanic, K., Katavetin, P., Kure, M., Wolkow, P., Dunn, J. S., Smiles, A., Walker, W. H., Boright, A. P., Bull, S. B., the DCCT/EDIC Research Group, Doria, A., Rogus, J. J., Rich, S. S., Warram, J. H., & Krolewski, A. S. (2009). Genome-wide association scan for diabetic nephropathy susceptibility genes in type 1 diabetes. Diabetes, 58(6), 1403–1410.
Pezzolesi, M. G., Skupien, J., Mychaleckyj, J. C., Warram, J. H., & Krolewski, A. S. (2010). Insights to the genetics of diabetic nephropathy through a genome-wide association study of the GoKinD Collection. Semin Nephrol, 30(2), 126–140.
Schadt, E. E., Sachs, A., & Friend, S. (2005). Embracing complexity, inching closer to reality. Sci STKE, 2005(295), pe40.
Dobrin, R., Zhu, J., Molony, C., Argman, C., Parrish, M., Carlson, S., Allan, M., Pomp, D., & Schadt, E. (2009). Multi-tissue coexpression networks reveal unexpected subnetworks associated with disease. Genome Biol, 10(5), R55.
Sauer, U., Heinemann, M., & Zamboni, N. (2007). Getting closer to the whole picture. Science, 316(5824), 550–551.
Chuang, H.-Y., Hofree, M., & Ideker, T. (2010). A decade of systems biology. Annu Rev Cell Dev Biol, 26(1), 721–744.
Houle, D., Govindaraju, D. R., & Omholt, S. (2010). Phenomics: the next challenge. Nat Rev Genet, 11(12), 855–866.
Haring, R., and Wallaschofski, H. (2012) Diving through the “-Omics”: the case for deep phenotyping and systems epidemiology. OMICS: J Integrative Biol 16 (in press) (Epub ahead of print: February 9, 2012).
Tomlins, S. A., Rhodes, D. R., Perner, S., Dhanasekaran, S. M., Mehra, R., Sun, X.-W., Varambally, S., Cao, X., Tchinda, J., Kuefer, R., Lee, C., Montie, J. E., Shah, R. B., Pienta, K. J., Rubin, M. A., & Chinnaiyan, A. M. (2005). Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science, 310(5748), 644–648.
Hamburg, M. A., & Collins, F. S. (2010). The path to personalized medicine. New Engl J Med, 363(4), 301–304.
Allison, M. (2008). Is personalized medicine finally arriving? Nat Biotechnol, 26(5), 509–517.
Seaquist, E. R., Goetz, F. C., Rich, S., & Barbosa, J. (1989). Familial clustering of diabetic kidney disease: evidence for genetic susceptibility to diabetic nephropathy. New Engl J Med, 320(18), 1161–1165.
Borch-Johnsen, K., Norgaard, K., Hommel, E., Mathiesen, E. R., Jensen, J. S., Deckert, T., & Parving, H.-H. (1992). Is diabetic nephropathy an inherited complication? Kidney Int, 41(4), 719–722.
Imperatore, G., Knowler, W. C., Pettitt, D. J., Kobes, S., Bennett, P. H., & Hanson, R. L. (2000). Segregation analysis of diabetic nephropathy in Pima Indians. Diabetes, 49(6), 1049–1056.
Knowler, W. C., Coresh, J., Elston, R. C., Freedman, B. I., Iyengar, S. K., Kimmel, P. L., Olson, J. M., Plaetke, R., Sedor, J. R., & Seldin, M. F. (2005). The Family Investigation of Nephropathy and Diabetes (FIND): design and methods. J Diabetes Complications, 19(1), 1–9.
Pezzolesi, M. G., Poznik, G. D., Skupien, J., Smiles, A. M., Mychaleckyj, J. C., Rich, S. S., Warram, J. H., & Krolewski, A. S. (2011). An intergenic region on chromosome 13q33.3 is associated with the susceptibility to kidney disease in type 1 and 2 diabetes. Kidney Int, 80(1), 105–111.
Mooyaart, A., Valk, E., van Es, L., Bruijn, J., de Heer, E., Freedman, B., Dekkers, O., & Baelde, H. (2011). Genetic associations in diabetic nephropathy: a meta-analysis. Diabetologia, 54(3), 544–553.
Manolio, T. A., Collins, F. S., Cox, N. J., Goldstein, D. B., Hindorff, L. A., Hunter, D. J., McCarthy, M. I., Ramos, E. M., Cardon, L. R., Chakravarti, A., Cho, J. H., Guttmacher, A. E., Kong, A., Kruglyak, L., Mardis, E., Rotimi, C. N., Slatkin, M., Valle, D., Whittemore, A. S., Boehnke, M., Clark, A. G., Eichler, E. E., Gibson, G., Haines, J. L., Mackay, T. F. C., McCarroll, S. A., & Visscher, P. M. (2009). Finding the missing heritability of complex diseases. Nature, 461(7265), 747–753.
Zuk, O., Hechter, E., Sunyaev, S. R., & Lander, E. S. (2012). The mystery of missing heritability: genetic interactions create phantom heritability. Proc Natl Acad Sci USA, 109(4), 1193–1198.
McCarthy, M. I., Abecasis, G. R., Cardon, L. R., Goldstein, D. B., Little, J., Ioannidis, J. P. A., & Hirschhorn, J. N. (2008). Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat Rev Genet, 9(5), 356–369.
Genovese, G., Friedman, D. J., Ross, M. D., Lecordier, L., Uzureau, P., Freedman, B. I., Bowden, D. W., Langefeld, C. D., Oleksyk, T. K., Uscinski Knob, A. L., Bernhardy, A. J., Hicks, P. J., Nelson, G. W., Vanhollebeke, B., Winkler, C. A., Kopp, J. B., Pays, E., & Pollak, M. R. (2010). Association of trypanolytic ApoL1 variants with kidney disease in African Americans. Science, 329(5993), 841–845.
Lusis, A. J. (2012). Life after GWAS. Atertio Thromb Vasc Biol, 32(2), 169–169.
Pomerantz, M. M., Ahmadiyeh, N., Jia, L., Herman, P., Verzi, M. P., Doddapaneni, H., Beckwith, C. A., Chan, J. A., Hills, A., Davis, M., Yao, K., Kehoe, S. M., Lenz, H.-J., Haiman, C. A., Yan, C., Henderson, B. E., Frenkel, B., Barretina, J., Bass, A., Tabernero, J., Baselga, J., Regan, M. M., Manak, J. R., Shivdasani, R., Coetzee, G. A., & Freedman, M. L. (2009). The 8q24 cancer risk variant rs6983267 shows long-range interaction with MYC in colorectal cancer. Nat Genet, 41(8), 882–884.
Pearson, E. R. (2009). Pharmacogenetics in diabetes. Curr Diab Rep, 9(2), 172–181.
McCarthy, M. I. (2010). Genomics, type 2 diabetes, and obesity. New Engl J Med, 363(24), 2339–2350.
The Diabetes Control and Complications Trial Research Group. (1993). The effect of intensive treatment of diabetes on the development and progression of long-term complications in insulin-dependent diabetes mellitus. New Engl J Med, 329(14), 977–986.
UK Prospective Diabetes Study (UKPDS) Group. (1998). Intensive blood-glucose control with sulphonylureas or insulin compared with conventional treatment and risk of complications in patients with type 2 diabetes (UKPDS 33). The Lancet, 352(9131), 837–853.
Engerman, R. L., & Kern, T. S. (1987). Progression of incipient diabetic retinopathy during good glycemic control. Diabetes, 36(7), 808–812.
Ihnat, M. A., Thorpe, J. E., & Ceriello, A. (2007). Hypothesis: the ‘metabolic memory’, the new challenge of diabetes. Diabet Med, 24(6), 582–586.
El-Osta, A., Brasacchio, D., Yao, D., Pocai, A., Jones, P. L., Roeder, R. G., Cooper, M. E., & Brownlee, M. (2008). Transient high glucose causes persistent epigenetic changes and altered gene expression during subsequent normoglycemia. J Exp Med, 205(10), 2409–2417.
Feinberg, A. P. (2008). Epigenetics at the epicenter of modern medicine. JAMA, 299(11), 1345–1350.
Ceriello, A., Ihnat, M. A., & Thorpe, J. E. (2009). The “metabolic memory”: is more than just tight glucose control necessary to prevent diabetic complications? J Clin Endocrinol Metab, 94(2), 410–415.
Dwivedi, R. S., Herman, J. G., McCaffrey, T. A., & Raj, D. S. C. (2011). Beyond genetics: epigenetic code in chronic kidney disease. Kidney Int, 79(1), 23–32.
Reddy, M. A., & Natarajan, R. (2011). Epigenetics in diabetic kidney disease. J Am Soc Nephrol, 22(12), 2182–2185.
Villeneuve, L. M., & Natarajan, R. (2010). Epigenetics of diabetic complications. Expert Rev Endocrinol Metab, 5(1), 137–148.
Pirola, L., Balcerczyk, A., Okabe, J., & El-Osta, A. (2010). Epigenetic phenomena linked to diabetic complications. Nat Rev Endocrinol, 6(12), 665–675.
Mohtat, D., & Susztak, K. (2010). Fine tuning gene expression: the epigenome. Semin Nephrol, 30(5), 468–476.
Reddy, M. A., & Natarajan, R. (2011). Epigenetic mechanisms in diabetic vascular complications. Cardiovasc Res, 90(3), 421–429.
Thomas, M. C., Groop, P.-H., & Tryggvason, K. (2012). Towards understanding the inherited susceptibility for nephropathy in diabetes. Curr Opin Nephrol Hypertens, 21(2), 195–202.
Wetterstrand, K. (2012) DNA sequencing costs: data from the NHGRI Large-Scale Genome Sequencing Program. Available at: http://wwwgenomegov/sequencingcosts. Accessed March 20, 2012.
Schena, M., Shalon, D., Davis, R. W., & Brown, P. O. (1995). Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science, 270(5235), 467–470.
Kretzler, M., Cohen, C. D., Doran, P., Henger, A., Madden, S., Gröne, E. F., Nelson, P. J., Schlöndorff, D., & Gröne, H.-J. (2002). Repuncturing the renal biopsy: strategies for molecular diagnosis in nephrology. J Am Soc Nephrol, 13(7), 1961–1972.
Cohen, C. D., Frach, K., Schlondorff, D., & Kretzler, M. (2002). Quantitative gene expression analysis in renal biopsies: a novel protocol for a high-throughput multicenter application. Kidney Int, 61(1), 133–140.
Schmid, H., Henger, A., Cohen, C. D., Frach, K., Gröne, H.-J., Schlöndorff, D., & Kretzler, M. (2003). Gene expression profiles of podocyte-associated molecules as diagnostic markers in acquired proteinuric diseases. J Am Soc Nephrol, 14(11), 2958–2966.
Kieran, N. E., Doran, P. P., Connolly, S. B., Greenan, M.-C., Higgins, D. F., Leonard, M., Godson, C., Taylor, C. T., Henger, A., Kretzler, M., Burne, M. J., Rabb, H., & Brady, H. R. (2003). Modification of the transcriptomic response to renal ischemia/reperfusion injury by lipoxin analog. Kidney Int, 64(2), 480–492.
Henger, A., Kretzler, M., Doran, P., Bonrouhi, M., Schmid, H., Kiss, E., Cohen, C. D., Madden, S., Porubsky, S., Grone, E. F., Schlondorff, D., Nelson, P. J., & Grone, H.-J. (2004). Gene expression fingerprints in human tubulointerstitial inflammation and fibrosis as prognostic markers of disease progression. Kidney Int, 65(3), 904–917.
Baelde, H. J., Eikmans, M., Doran, P. P., Lappin, D. W. P., de Heer, E., & Bruijn, J. A. (2004). Gene expression profiling in glomeruli from human kidneys with diabetic nephropathy. Am J Kidney Dis, 43(4), 636–650.
Schmid, H., Boucherot, A., Yasuda, Y., Henger, A., Brunner, B., Eichinger, F., Nitsche, A., Kiss, E., Bleich, M., Gröne, H.-J., Nelson, P. J., Schlöndorff, D., Cohen, C. D., & Kretzler, M. (2006). Modular activation of nuclear factor-κB transcriptional programs in human diabetic nephropathy. Diabetes, 55(11), 2993–3003.
Lindenmeyer, M. T., Kretzler, M., Boucherot, A., Berra, S., Yasuda, Y., Henger, A., Eichinger, F., Gaiser, S., Schmid, H., Rastaldi, M. P., Schrier, R. W., Schlondorff, D., & Cohen, C. D. (2007). Interstitial vascular rarefaction and reduced VEGF-A expression in human diabetic nephropathy. J Am Soc Nephrol, 18(6), 1765–1776.
Woroniecka, K. I., Park, A. S. D., Mohtat, D., Thomas, D. B., Pullman, J. M., & Susztak, K. (2011). Transcriptome analysis of human diabetic kidney disease. Diabetes, 60(9), 2354–2369.
Lin, Y. L., Peng, S. J., Ferng, S. H., Tzen, C. Y., & Yang, C. S. (2009). Clinical indicators which necessitate renal biopsy in type 2 diabetes mellitus patients with renal disease. Int J Clin Pract, 63(8), 1167–1176.
Mauer, S. M., Steffes, M. W., Ellis, E. N., Sutherland, D. E., Brown, D. M., & Goetz, F. C. (1984). Structural-functional relationships in diabetic nephropathy. J Clin Invest, 74(4), 1143–1155.
Pagtalunan, M. E., Miller, P. L., Jumping-Eagle, S., Nelson, R. G., Myers, B. D., Rennke, H. G., Coplon, N. S., Sun, L., & Meyer, T. W. (1997). Podocyte loss and progressive glomerular injury in type II diabetes. J Clin Invest, 99(2), 342–348.
Moutzouris, D.-A., Herlitz, L., Appel, G. B., Markowitz, G. S., Freudenthal, B., Radhakrishnan, J., & D’Agati, V. D. (2009). Renal biopsy in the very elderly. Clin J Am Soc Nephrol, 4(6), 1073–1082.
Zhang, P.-P., Ge, Y.-C., Li, S.-J., Xie, H.-L., Li, L.-S., & Liu, Z.-H. (2011). Renal biopsy in type 2 diabetes: timing of complications and evaluating of safety in Chinese patients. Nephrology, 16(1), 100–105.
Cohen, C. D., Grone, H.-J., Grone, E. F., Nelson, P. J., Schlondorff, D., & Kretzler, M. (2002). Laser microdissection and gene expression analysis on formaldehyde-fixed archival tissue. Kidney Int, 61(1), 125–132.
Hodgin, J. B., Borczuk, A. C., Nasr, S. H., Markowitz, G. S., Nair, V., Martini, S., Eichinger, F., Vining, C., Berthier, C. C., Kretzler, M., & D’Agati, V. D. (2010). A molecular profile of focal segmental glomerulosclerosis from formalin-fixed, paraffin-embedded tissue. Am J Pathol, 177(4), 1674–1686.
Shen-Orr, S. S., Tibshirani, R., Khatri, P., Bodian, D. L., Staedtler, F., Perry, N. M., Hastie, T., Sarwal, M. M., Davis, M. M., & Butte, A. J. (2010). Cell type-specific gene expression differences in complex tissues. Nat Methods, 7(4), 287–289.
Kaiser, J. (2012). Biomarker tests need closer scrutiny, IOM concludes. Science, 335(6076), 1554–1554.
Consensus Report Institute of Medicine (2012) Evolution of translational OMICS: lessons learned and the path forward. At: http://wwwiomedu/Reports/2012/Evolution-of-Translational-Omics.aspx. Accessed March 23, 2012
Gygi, S. P., Rochon, Y., Franza, B. R., & Aebersold, R. (1999). Correlation between protein and mRNA abundance in yeast. Mol Cell Biol, 19(3), 1720–1730.
Cravatt, B. F., Simon, G. M., & Yates Iii, J. R. (2007). The biological impact of mass-spectrometry-based proteomics. Nature, 450(7172), 991–1000.
Vogel, C., & Marcotte, E. M. (2012). Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat Rev Genet, 13(4), 227–232.
Sébédio, J.-L., Pujos-Guillot, E., & Ferrara, M. (2009). Metabolomics in evaluation of glucose disorders. Curr Opin Clin Nutr Metab Care, 12(4), 412–418.
Wang, J. H., Byun, J., & Pennathur, S. (2010). Analytical approaches to metabolomics and applications to systems biology. Semin Nephrol, 30(5), 500–511.
Ben Ameur, R., Molina, L., Bolvin, C., Kifagi, C., Jarraya, F., Ayadi, H., Molina, F., & Granier, C. (2010). Proteomic approaches for discovering biomarkers of diabetic nephropathy. Nephrol Dial Transplant, 25(9), 2866–2875.
Thongboonkerd, V. (2011). Study of diabetic nephropathy in the proteomic era. Contrib Nephrol, 170, 172–183.
Mäkinen, V.-P., Tynkkynen, T., Soininen, P., Peltola, T., Kangas, A. J., Forsblom, C., Thorn, L. M., Kaski, K., Laatikainen, R., Ala-Korpela, M., & Groop, P.-H. (2012). Metabolic diversity of progressive kidney disease in 325 patients with type 1 diabetes (the FinnDiane Study). J Proteome Res, 11(3), 1782–1790.
Thongboonkerd, V., & Malasit, P. (2005). Renal and urinary proteomics: current applications and challenges. Proteomics, 5(4), 1033–1042.
Decramer, S., de Peredo, A. G., Breuil, B., Mischak, H., Monsarrat, B., Bascands, J.-L., & Schanstra, J. P. (2008). Urine in clinical proteomics. Mol Cell Proteomics, 7(10), 1850–1862.
Rossing, K., Mischak, H., Dakna, M., Zurbig, P., Novak, J., Julian, B. A., Good, D. M., Coon, J. J., Tarnow, L., Rossing, P., & on behalf of the PREDICTIONS Network. (2008). Urinary proteomics in diabetes and CKD. J Am Soc Nephrol, 19(7), 1283–1290.
Merchant, M. L., Perkins, B. A., Boratyn, G. M., Ficociello, L. H., Wilkey, D. W., Barati, M. T., Bertram, C. C., Page, G. P., Rovin, B. H., Warram, J. H., Krolewski, A. S., & Klein, J. B. (2009). Urinary peptidome may predict renal function decline in type 1 diabetes and microalbuminuria. J Am Soc Nephrol, 20(9), 2065–2074.
Otu, H. H., Can, H., Spentzos, D., Nelson, R. G., Hanson, R. L., Looker, H. C., Knowler, W. C., Monroy, M., Libermann, T. A., Karumanchi, S. A., & Thadhani, R. (2007). Prediction of diabetic nephropathy using urine proteomic profiling 10 years prior to development of nephropathy. Diabetes Care, 30(3), 638–643.
Overgaard, A., Hansen, H., Lajer, M., Pedersen, L., Tarnow, L., Rossing, P., McGuire, J., & Pociot, F. (2010). Plasma proteome analysis of patients with type 1 diabetes with diabetic nephropathy. Proteome Sci, 8(1), 4.
Overgaard, A. J., Thingholm, T. E., Larsen, M. R., Tarnow, L., Rossing, P., McGuire, J. N., & Pociot, F. (2010). Quantitative iTRAQ-based proteomic identification of candidate biomarkers for diabetic nephropathy in plasma of type 1 diabetic patients. Clin Proteomics, 6(4), 105–114.
Coon, J. J., Zürbig, P., Dakna, M., Dominiczak, A. F., Decramer, S., Fliser, D., Frommberger, M., Golovko, I., Good, D. M., Herget-Rosenthal, S., Jankowski, J., Julian, B. A., Kellmann, M., Kolch, W., Massy, Z., Novak, J., Rossing, K., Schanstra, J. P., Schiffer, E., Theodorescu, D., Vanholder, R., Weissinger, E. M., Mischak, H., & Schmitt-Kopplin, P. (2008). CE-MS analysis of the human urinary proteome for biomarker discovery and disease diagnostics. Proteomics Clin Appl, 2(7–8), 964–973.
Jantos-Siwy, J., Schiffer, E., Brand, K., Schumann, G., Rossing, K., Delles, C., Mischak, H., & Metzger, J. (2009). Quantitative urinary proteome analysis for biomarker evaluation in chronic kidney disease. J Proteome Res, 8(1), 268–281.
Maahs, D. M., Siwy, J., Argilés, À., Cerna, M., Delles, C., Dominiczak, A. F., Gayrard, N., Iphöfer, A., Jänsch, L., Jerums, G., Medek, K., Mischak, H., Navis, G. J., Roob, J. M., Rossing, K., Rossing, P., Rychlík, I., Schiffer, E., Schmieder, R. E., Wascher, T. C., Winklhofer-Roob, B. M., Zimmerli, L. U., Zürbig, P., & Snell-Bergeon, J. K. (2010). Urinary collagen fragments are significantly altered in diabetes: a link to pathophysiology. PLoS ONE, 5(9), e13051.
Alkhalaf, A., Zürbig, P., Bakker, S. J. L., Bilo, H. J. G., Cerna, M., Fischer, C., Fuchs, S., Janssen, B., Medek, K., Mischak, H., Roob, J. M., Rossing, K., Rossing, P., Rychlík, I., Sourij, H., Tiran, B., Winklhofer-Roob, B. M., Navis, G. J., & for the PREDICTIONS Group. (2010). Multicentric validation of proteomic biomarkers in urine specific for diabetic nephropathy. PLoS ONE, 5(10), e13421.
Good, D. M., Zurbig, P., Argiles, A., Bauer, H. W., Behrens, G., Coon, J. J., Dakna, M., Decramer, S., Delles, C., Dominiczak, A. F., Ehrich, J. H. H., Eitner, F., Fliser, D., Frommberger, M., Ganser, A., Girolami, M. A., Golovko, I., Gwinner, W., Haubitz, M., Herget-Rosenthal, S., Jankowski, J., Jahn, H., Jerums, G., Julian, B. A., Kellmann, M., Kliem, V., Kolch, W., Krolewski, A. S., Luppi, M., Massy, Z., Melter, M., Neususs, C., Novak, J., Peter, K., Rossing, K., Rupprecht, H., Schanstra, J. P., Schiffer, E., Stolzenburg, J.-U., Tarnow, L., Theodorescu, D., Thongboonkerd, V., Vanholder, R., Weissinger, E. M., Mischak, H., & Schmitt-Kopplin, P. (2010). Naturally occurring human urinary peptides for use in diagnosis of chronic kidney disease. Mol Cell Proteomics, 9(11), 2424–2437.
Jain, S., Rajput, A., Kumar, Y., Uppuluri, N., Arvind, A. S., & Tatu, U. (2005). Proteomic analysis of urinary protein markers for accurate prediction of diabetic kidney disorder. J Assoc Physicians India, 53(June), 513–520.
Fisher, W. G., Lucas, J. E., Mehdi, U. F., Qunibi, D. W., Garner, H. R., Rosenblatt, K. P., & Toto, R. D. (2011). A method for isolation and identification of urinary biomarkers in patients with diabetic nephropathy. Proteomics Clin Appl, 5(11–12), 603–612.
Lowe, J. B. (2001). Glycosylation, immunity, and autoimmunity. Cell, 104(6), 809–812.
Yang, N., Feng, S., Shedden, K., Xie, X., Liu, Y., Rosser, C. J., Lubman, D. M., & Goodison, S. (2011). Urinary glycoprotein biomarker discovery for bladder cancer detection using LC/MS-MS and label-free quantification. Clin Cancer Res, 17(10), 3349–3359.
Ahn, J.-M., Kim, B.-G., Yu, M.-H., Lee, I.-K., & Cho, J.-Y. (2010). Identification of diabetic nephropathy-selective proteins in human plasma by multi-lectin affinity chromatography and LC-MS/MS. Proteomics Clin Appl, 4(6–7), 644–653.
Vivekanandan-Giri, A., Slocum, J. L., Buller, C. L., Basrur, V., Ju, W., Pop-Busui, R., et al. (2011). Urine glycoprotein profile reveals novel markers for chronic kidney disease. Int J Proteomics, 2011, 18. Article ID 214715.
Pisitkun, T., Shen, R.-F., & Knepper, M. A. (2004). Identification and proteomic profiling of exosomes in human urine. Proc Natl Acad Sci USA, 101(36), 13368–13373.
Gonzales, P. A., Pisitkun, T., Hoffert, J. D., Tchapyjnikov, D., Star, R. A., Kleta, R., Wang, N. S., & Knepper, M. A. (2009). Large-scale proteomics and phosphoproteomics of urinary exosomes. J Am Soc Nephrol, 20(2), 363–379.
Konvalinka, A., Scholey, J. W., & Diamandis, E. P. (2012). Searching for new biomarkers of renal diseases through proteomics. Clin Chem, 58(2), 353–365.
Mischak, H., Apweiler, R., Banks, R. E., Conaway, M., Coon, J., Dominiczak, A., Ehrich, J. H. H., Fliser, D., Girolami, M., Hermjakob, H., Hochstrasser, D., Jankowski, J., Julian, B. A., Kolch, W., Massy, Z. A., Neusuess, C., Novak, J., Peter, K., Rossing, K., Schanstra, J., Semmes, O. J., Theodorescu, D., Thongboonkerd, V., Weissinger, E. M., Van Eyk, J. E., & Yamamoto, T. (2007). Clinical proteomics: a need to define the field and to begin to set adequate standards. Proteomics Clin Appl, 1(2), 148–156.
Good, D. M., Thongboonkerd, V., Novak, J., Bascands, J.-L., Schanstra, J. P., Coon, J. J., Dominiczak, A., & Mischak, H. (2007). Body fluid proteomics for biomarker discovery: lessons from the past hold the key to success in the future. J Proteome Res, 6(12), 4549–4555.
Kinsinger, C. R., Apffel, J., Baker, M., Bian, X., Borchers, C. H., Bradshaw, R., Brusniak, M.-Y., Chan, D. W., Deutsch, E. W., Domon, B., Gorman, J., Grimm, R., Hancock, W., Hermjakob, H., Horn, D., Hunter, C., Kolar, P., Kraus, H.-J., Langen, H., Linding, R., Moritz, R. L., Omenn, G. S., Orlando, R., Pandey, A., Ping, P., Rahbar, A., Rivers, R., Seymour, S. L., Simpson, R. J., Slotta, D., Smith, R. D., Stein, S. E., Tabb, D. L., Tagle, D., Yates, J. R., & Rodriguez, H. (2011). Recommendations for mass spectrometry data quality metrics for open access data (corollary to the Amsterdam principles). Proteomics Clin Appl, 5(11–12), 580–589.
Brazma, A., Hingamp, P., Quackenbush, J., Sherlock, G., Spellman, P., Stoeckert, C., Aach, J., Ansorge, W., Ball, C. A., Causton, H. C., Gaasterland, T., Glenisson, P., Holstege, F. C. P., Kim, I. F., Markowitz, V., Matese, J. C., Parkinson, H., Robinson, A., Sarkans, U., Schulze-Kremer, S., Stewart, J., Taylor, R., Vilo, J., & Vingron, M. (2001). Minimum information about a microarray experiment (MIAME)—toward standards for microarray data. Nat Genet, 29(4), 365–371.
Sreekumar, A., Poisson, L. M., Rajendiran, T. M., Khan, A. P., Cao, Q., Yu, J., Laxman, B., Mehra, R., Lonigro, R. J., Li, Y., Nyati, M. K., Ahsan, A., Kalyana-Sundaram, S., Han, B., Cao, X., Byun, J., Omenn, G. S., Ghosh, D., Pennathur, S., Alexander, D. C., Berger, A., Shuster, J. R., Wei, J. T., Varambally, S., Beecher, C., & Chinnaiyan, A. M. (2009). Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature, 457(7231), 910–914.
Kim, K., Aronov, P., Zakharkin, S. O., Anderson, D., Perroud, B., Thompson, I. M., & Weiss, R. H. (2009). Urine metabolomics analysis for kidney cancer detection and biomarker discovery. Mol Cell Proteomics, 8(3), 558–570.
Newgard, C. B., An, J., Bain, J. R., Muehlbauer, M. J., Stevens, R. D., Lien, L. F., Haqq, A. M., Shah, S. H., Arlotto, M., Slentz, C. A., Rochon, J., Gallup, D., Ilkayeva, O., Wenner, B. R., Yancy, W. S., Jr., Eisenson, H., Musante, G., Surwit, R. S., Millington, D. S., Butler, M. D., & Svetkey, L. P. (2009). A branched-chain amino acid-related metabolic signature that differentiates obese and lean humans and contributes to insulin resistance. Cell Metabolism, 9(4), 311–326.
Suhre, K., Meisinger, C., Döring, A., Altmaier, E., Belcredi, P., Gieger, C., Chang, D., Milburn, M. V., Gall, W. E., Weinberger, K. M., Mewes, H.-W., Hrabé de Angelis, M., Wichmann, H. E., Kronenberg, F., Adamski, J., & Illig, T. (2010). Metabolic footprint of diabetes: a multiplatform metabolomics study in an epidemiological setting. PLoS ONE, 5(11), e13953.
Wang, T. J., Larson, M. G., Vasan, R. S., Cheng, S., Rhee, E. P., McCabe, E., Lewis, G. D., Fox, C. S., Jacques, P. F., Fernandez, C., O’Donnell, C. J., Carr, S. A., Mootha, V. K., Florez, J. C., Souza, A., Melander, O., Clish, C. B., & Gerszten, R. E. (2011). Metabolite profiles and the risk of developing diabetes. Nat Med, 17(4), 448–453.
Rhee, E. P., Cheng, S., Larson, M. G., Walford, G. A., Lewis, G. D., McCabe, E., Yang, E., Farrell, L., Fox, C. S., O’Donnell, C. J., Carr, S. A., Vasan, R. S., Florez, J. C., Clish, C. B., Wang, T. J., & Gerszten, R. E. (2011). Lipid profiling identifies a triacylglycerol signature of insulin resistance and improves diabetes prediction in humans. J Clin Invest, 121(4), 1402–1411.
Ganti, S., & Weiss, R. H. (2011). Urine metabolomics for kidney cancer detection and biomarker discovery. Urol Oncol, 29(5), 551–557.
Rhee, E. P., & Gerszten, R. E. (2012). Metabolomics and cardiovascular biomarker discovery. Clin Chem, 58(1), 139–147.
Weiss, R. H., & Kim, K. (2012). Metabolomics in the study of kidney diseases. Nat Rev Nephrol, 8(1), 22–33.
Zhang, H., Saha, J., Byun, J., Schin, M., Lorenz, M., Kennedy, R. T., Kretzler, M., Feldman, E. L., Pennathur, S., & Brosius, F. C. (2008). Rosiglitazone reduces renal and plasma markers of oxidative injury and reverses urinary metabolite abnormalities in the amelioration of diabetic nephropathy. Am J Physiol Renal Physiol, 295(4), F1071–F1081.
Smith, C. A., Maille, G. O., Want, E. J., Qin, C., Trauger, S. A., Brandon, T. R., Custodio, D. E., Abagyan, R., & Siuzdak, G. (2005). METLIN: a metabolite mass spectral database. Ther Drug Monit, 27(6), 747–751.
Wishart, D. S., Tzur, D., Knox, C., Eisner, R., Guo, A. C., Young, N., Cheng, D., Jewell, K., Arndt, D., Sawhney, S., Fung, C., Nikolai, L., Lewis, M., Coutouly, M.-A., Forsythe, I., Tang, P., Shrivastava, S., Jeroncic, K., Stothard, P., Amegbey, G., Block, D., Hau, D. D., Wagner, J., Miniaci, J., Clements, M., Gebremedhin, M., Guo, N., Zhang, Y., Duggan, G. E., MacInnis, G. D., Weljie, A. M., Dowlatabadi, R., Bamforth, F., Clive, D., Greiner, R., Li, L., Marrie, T., Sykes, B. D., Vogel, H. J., & Querengesser, L. (2007). HMDB: the Human Metabolome Database. NAR, 35(suppl 1), D521–D526.
Barreto, F. C., Barreto, D. V., Liabeuf, S., Meert, N., Glorieux, G., Temmar, M., Choukroun, G., Vanholder, R., Massy, Z. A., & on behalf of the European Uremic Toxin Work Group. (2009). Serum indoxyl sulfate is associated with vascular disease and mortality in chronic kidney disease patients. Clin J Am Soc Nephrol, 4(10), 1551–1558.
Zhang, J., Yan, L., Chen, W., Lin, L., Song, X., Yan, X., Hang, W., & Huang, B. (2009). Metabonomics research of diabetic nephropathy and type 2 diabetes mellitus based on UPLC–oaTOF-MS system. Anal Chim Acta, 650(1), 16–22.
van der Kloet, F., Tempels, F., Ismail, N., van der Heijden, R., Kasper, P., Rojas-Cherto, M., van Doorn, R., Spijksma, G., Koek, M., van der Greef, J., Mäkinen, V., Forsblom, C., Holthöfer, H., Groop, P., Reijmers, T., & Hankemeier, T. (2012). Discovery of early-stage biomarkers for diabetic kidney disease using ms-based metabolomics (FinnDiane study). Metabolomics, 8(1), 109–119.
Xia, J.-F., Liang, Q.-L., Liang, X.-P., Wang, Y.-M., Hu, P., Li, P., & Luo, G.-A. (2009). Ultraviolet and tandem mass spectrometry for simultaneous quantification of 21 pivotal metabolites in plasma from patients with diabetic nephropathy. J Chromatogr B Analyt Technol Biomed Life Sci, 877(20–21), 1930–1936.
Sieberts, S., & Schadt, E. E. (2007). Moving toward a system genetics view of disease. Mamm Genome, 18(6), 389–401.
Emilsson, V., Thorleifsson, G., Zhang, B., Leonardson, A. S., Zink, F., Zhu, J., Carlson, S., Helgason, A., Walters, G. B., Gunnarsdottir, S., Mouy, M., Steinthorsdottir, V., Eiriksdottir, G. H., Bjornsdottir, G., Reynisdottir, I., Gudbjartsson, D., Helgadottir, A., Jonasdottir, A., Jonasdottir, A., Styrkarsdottir, U., Gretarsdottir, S., Magnusson, K. P., Stefansson, H., Fossdal, R., Kristjansson, K., Gislason, H. G., Stefansson, T., Leifsson, B. G., Thorsteinsdottir, U., Lamb, J. R., Gulcher, J. R., Reitman, M. L., Kong, A., Schadt, E. E., & Stefansson, K. (2008). Genetics of gene expression and its effect on disease. Nature, 452(7186), 423–428.
Ioannidis, J. P. A., Thomas, G., & Daly, M. J. (2009). Validating, augmenting and refining genome-wide association signals. Nat Rev Genet, 10(5), 318–329.
Keurentjes, J. J. B., Fu, J., de Vos, C. H. R., Lommen, A., Hall, R. D., Bino, R. J., van der Plas, L. H. W., Jansen, R. C., Vreugdenhil, D., & Koornneef, M. (2006). The genetics of plant metabolism. Nat Genet, 38(7), 842–849.
Gieger, C., Geistlinger, L., Altmaier, E., Hrabé de Angelis, M., Kronenberg, F., Meitinger, T., Mewes, H.-W., Wichmann, H. E., Weinberger, K. M., Adamski, J., Illig, T., & Suhre, K. (2008). Genetics meets metabolomics: a genome-wide association study of metabolite profiles in human serum. PLoS Genet, 4(11), e1000282.
Ferrara, C. T., Wang, P., Neto, E. C., Stevens, R. D., Bain, J. R., Wenner, B. R., Ilkayeva, O. R., Keller, M. P., Blasiole, D. A., Kendziorski, C., Yandell, B. S., Newgard, C. B., & Attie, A. D. (2008). Genetic networks of liver metabolism revealed by integration of metabolic and transcriptional profiling. PLoS Genet, 4(3), e1000034.
Shah, S. H., Hauser, E. R., Bain, J. R., Muehlbauer, M. J., Haynes, C., Stevens, R. D., et al. (2009). High heritability of metabolomic profiles in families burdened with premature cardiovascular disease. Mol Syst Biol, 5, 258.
Suhre, K., Wallaschofski, H., Raffler, J., Friedrich, N., Haring, R., Michael, K., Wasner, C., Krebs, A., Kronenberg, F., Chang, D., Meisinger, C., Wichmann, H. E., Hoffmann, W., Volzke, H., Volker, U., Teumer, A., Biffar, R., Kocher, T., Felix, S. B., Illig, T., Kroemer, H. K., Gieger, C., Romisch-Margl, W., & Nauck, M. (2011). A genome-wide association study of metabolic traits in human urine. Nat Genet, 43(6), 565–569.
Daly, A. K. (2010). Drug-induced liver injury: past, present and future. Pharmacogenomics, 11(5), 607–611.
Teslovich, T. M., Musunuru, K., Smith, A. V., Edmondson, A. C., Stylianou, I. M., Koseki, M., Pirruccello, J. P., Ripatti, S., Chasman, D. I., Willer, C. J., Johansen, C. T., Fouchier, S. W., Isaacs, A., Peloso, G. M., Barbalic, M., Ricketts, S. L., Bis, J. C., Aulchenko, Y. S., Thorleifsson, G., Feitosa, M. F., Chambers, J., Orho-Melander, M., Melander, O., Johnson, T., Li, X., Guo, X., Li, M., Shin Cho, Y., Jin Go, M., Jin Kim, Y., Lee, J.-Y., Park, T., Kim, K., Sim, X., Twee-Hee Ong, R., Croteau-Chonka, D. C., Lange, L. A., Smith, J. D., Song, K., Hua Zhao, J., Yuan, X., Luan, J. A., Lamina, C., Ziegler, A., Zhang, W., Zee, R. Y. L., Wright, A. F., Witteman, J. C. M., Wilson, J. F., Willemsen, G., Wichmann, H. E., Whitfield, J. B., Waterworth, D. M., Wareham, N. J., Waeber, G., Vollenweider, P., Voight, B. F., Vitart, V., Uitterlinden, A. G., Uda, M., Tuomilehto, J., Thompson, J. R., Tanaka, T., Surakka, I., Stringham, H. M., Spector, T. D., Soranzo, N., Smit, J. H., Sinisalo, J., Silander, K., Sijbrands, E. J. G., Scuteri, A., Scott, J., Schlessinger, D., Sanna, S., Salomaa, V., Saharinen, J., Sabatti, C., Ruokonen, A., Rudan, I., Rose, L. M., Roberts, R., Rieder, M., Psaty, B. M., Pramstaller, P. P., Pichler, I., Perola, M., Penninx, B. W. J. H., Pedersen, N. L., Pattaro, C., Parker, A. N., Pare, G., Oostra, B. A., O’Donnell, C. J., Nieminen, M. S., Nickerson, D. A., Montgomery, G. W., Meitinger, T., McPherson, R., McCarthy, M. I., McArdle, W., Masson, D., Martin, N. G., Marroni, F., Mangino, M., Magnusson, P. K. E., Lucas, G., Luben, R., Loos, R. J. F., Lokki, M.-L., Lettre, G., Langenberg, C., Launer, L. J., Lakatta, E. G., Laaksonen, R., Kyvik, K. O., Kronenberg, F., Konig, I. R., Khaw, K.-T., Kaprio, J., Kaplan, L. M., Johansson, A., Jarvelin, M.-R., Cecile, J. W., Janssens, A., Ingelsson, E., Igl, W., Kees Hovingh, G., Hottenga, J.-J., Hofman, A., Hicks, A. A., Hengstenberg, C., Heid, I. M., Hayward, C., Havulinna, A. S., Hastie, N. D., Harris, T. B., Haritunians, T., Hall, A. S., Gyllensten, U., Guiducci, C., Groop, L. C., Gonzalez, E., Gieger, C., Freimer, N. B., Ferrucci, L., Erdmann, J., Elliott, P., Ejebe, K. G., Doring, A., Dominiczak, A. F., Demissie, S., Deloukas, P., de Geus, E. J. C., de Faire, U., Crawford, G., Collins, F. S., Chen, Y.-D. I., Caulfield, M. J., Campbell, H., Burtt, N. P., Bonnycastle, L. L., Boomsma, D. I., Boekholdt, S. M., Bergman, R. N., Barroso, I., Bandinelli, S., Ballantyne, C. M., Assimes, T. L., Quertermous, T., Altshuler, D., Seielstad, M., Wong, T. Y., Tai, E. S., Feranil, A. B., Kuzawa, C. W., Adair, L. S., Taylor, H. A., Jr., Borecki, I. B., Gabriel, S. B., Wilson, J. G., Holm, H., Thorsteinsdottir, U., Gudnason, V., Krauss, R. M., Mohlke, K. L., Ordovas, J. M., Munroe, P. B., Kooner, J. S., Tall, A. R., Hegele, R. A., Kastelein, J. J. P., Schadt, E. E., Rotter, J. I., Boerwinkle, E., Strachan, D. P., Mooser, V., Stefansson, K., Reilly, M. P., Samani, N. J., Schunkert, H., Cupples, L. A., Sandhu, M. S., Ridker, P. M., Rader, D. J., van Duijn, C. M., Peltonen, L., Abecasis, G. R., Boehnke, M., & Kathiresan, S. (2010). Biological, clinical and population relevance of 95 loci for blood lipids. Nature, 466(7307), 707–713.
Köttgen, A., Pattaro, C., Boger, C. A., Fuchsberger, C., Olden, M., Glazer, N. L., Parsa, A., Gao, X., Yang, Q., Smith, A. V., O’Connell, J. R., Li, M., Schmidt, H., Tanaka, T., Isaacs, A., Ketkar, S., Hwang, S.-J., Johnson, A. D., Dehghan, A., Teumer, A., Pare, G., Atkinson, E. J., Zeller, T., Lohman, K., Cornelis, M. C., Probst-Hensch, N. M., Kronenberg, F., Tonjes, A., Hayward, C., Aspelund, T., Eiriksdottir, G., Launer, L. J., Harris, T. B., Rampersaud, E., Mitchell, B. D., Arking, D. E., Boerwinkle, E., Struchalin, M., Cavalieri, M., Singleton, A., Giallauria, F., Metter, J., de Boer, I. H., Haritunians, T., Lumley, T., Siscovick, D., Psaty, B. M., Zillikens, M. C., Oostra, B. A., Feitosa, M., Province, M., de Andrade, M., Turner, S. T., Schillert, A., Ziegler, A., Wild, P. S., Schnabel, R. B., Wilde, S., Munzel, T. F., Leak, T. S., Illig, T., Klopp, N., Meisinger, C., Wichmann, H. E., Koenig, W., Zgaga, L., Zemunik, T., Kolcic, I., Minelli, C., Hu, F. B., Johansson, A., Igl, W., Zaboli, G., Wild, S. H., Wright, A. F., Campbell, H., Ellinghaus, D., Schreiber, S., Aulchenko, Y. S., Felix, J. F., Rivadeneira, F., Uitterlinden, A. G., Hofman, A., Imboden, M., Nitsch, D., Brandstatter, A., Kollerits, B., Kedenko, L., Magi, R., Stumvoll, M., Kovacs, P., Boban, M., Campbell, S., Endlich, K., Volzke, H., Kroemer, H. K., Nauck, M., Volker, U., Polasek, O., Vitart, V., Badola, S., Parker, A. N., Ridker, P. M., Kardia, S. L. R., Blankenberg, S., Liu, Y., Curhan, G. C., Franke, A., Rochat, T., Paulweber, B., Prokopenko, I., Wang, W., Gudnason, V., Shuldiner, A. R., Coresh, J., Schmidt, R., Ferrucci, L., Shlipak, M. G., van Duijn, C. M., Borecki, I., Kramer, B. K., Rudan, I., Gyllensten, U., Wilson, J. F., Witteman, J. C., Pramstaller, P. P., Rettig, R., Hastie, N., Chasman, D. I., Kao, W. H., Heid, I. M., & Fox, C. S. (2010). New loci associated with kidney function and chronic kidney disease. Nat Genet, 42(5), 376–384.
Kettunen, J., Tukiainen, T., Sarin, A.-P., Ortega-Alonso, A., Tikkanen, E., Lyytikainen, L.-P., Kangas, A. J., Soininen, P., Wurtz, P., Silander, K., Dick, D. M., Rose, R. J., Savolainen, M. J., Viikari, J., Kahonen, M., Lehtimaki, T., Pietilainen, K. H., Inouye, M., McCarthy, M. I., Jula, A., Eriksson, J., Raitakari, O. T., Salomaa, V., Kaprio, J., Jarvelin, M.-R., Peltonen, L., Perola, M., Freimer, N. B., Ala-Korpela, M., Palotie, A., & Ripatti, S. (2012). Genome-wide association study identifies multiple loci influencing human serum metabolite levels. Nat Genet, 44(3), 269–276.
Suhre, K., Shin, S.-Y., Petersen, A.-K., Mohney, R. P., Meredith, D., Wagele, B., et al. (2011). Human metabolic individuality in biomedical and pharmaceutical research. Nature, 477(7362), 54–60.
Nicholson, G., Rantalainen, M., Li, J. V., Maher, A. D., Malmodin, D., Ahmadi, K. R., Faber, J. H., Barrett, A., Min, J. L., Rayner, N. W., Toft, H., Krestyaninova, M., Viksna, J., Neogi, S. G., Dumas, M.-E., Sarkans, U., Donnelly, P., Illig, T., Adamski, J., Suhre, K., Allen, M., Zondervan, K. T., Spector, T. D., Nicholson, J. K., Lindon, J. C., Baunsgaard, D., Holmes, E., McCarthy, M. I., Holmes, C. C., & The Mol, P. C. (2011). A genome-wide metabolic QTL analysis in Europeans implicates two loci shaped by recent positive selection. PLoS Genet, 7(9), e1002270.
Barabasi, A.-L., & Oltvai, Z. N. (2004). Network biology: understanding the cell’s functional organization. Nat Rev Genet, 5(2), 101–113.
Barabasi, A.-L., Gulbahce, N., & Loscalzo, J. (2011). Network medicine: a network-based approach to human disease. Nat Rev Genet, 12(1), 56–68.
Emmert-Streib, F., & Glazko, G. V. (2011). Network biology: a direct approach to study biological function. Wiley Interdiscip Rev Syst Biol Med, 3(4), 379–391.
Keller, B. J., Martini, S., Sedor, J. R., & Kretzler, M. (2012). A systems view of genetics in chronic kidney disease. Kidney Int, 81(1), 14–21.
He, J. C., Chuang, P. Y., Ma’Ayan, A., & Iyengar, R. (2012). Systems biology of kidney diseases. Kidney Int, 81(1), 22–39.
Schadt, E., Zhang, B., & Zhu, J. (2009). Advances in systems biology are enhancing our understanding of disease and moving us closer to novel disease treatments. Gen, 136(2), 259–269.
Ostrowski, J., & Wyrwicz, L. S. (2009). Integrating genomics, proteomics and bioinformatics in translational studies of molecular medicine. Expert Rev Mol Diagn, 9(6), 623–630.
Ideker, T., & Krogan, N. J. (2012). Differential network biology. Mol Syst Biol, 8, 565.
Schadt, E. E., Molony, C., Chudin, E., Hao, K., Yang, X., Lum, P. Y., Kasarskis, A., Zhang, B., Wang, S., Suver, C., Zhu, J., Millstein, J., Sieberts, S., Lamb, J., GuhaThakurta, D., Derry, J., Storey, J. D., Avila-Campillo, I., Kruger, M. J., Johnson, J. M., Rohl, C. A., van Nas, A., Mehrabian, M., Drake, T. A., Lusis, A. J., Smith, R. C., Guengerich, F. P., Strom, S. C., Schuetz, E., Rushmore, T. H., & Ulrich, R. (2008). Mapping the genetic architecture of gene expression in human liver. PLoS Biol, 6(5), e107.
Mirel, B., Eichinger, F., Keller, B. J., & Kretzler, M. (2011). A cognitive task analysis of a visual analytic workflow: exploring molecular interaction networks in systems biology. J Biomed Discov Collab, 6, 1–33.
Bergholdt, R., Brorsson, C., Palleja, A., Berchtold, L. A., Fløyel, T., Bang-Berthelsen, C. H., Frederiksen, K. S., Jensen, L. J., Størling, J., & Pociot, F. (2012). Identification of novel type 1 diabetes candidate genes by integrating genome-wide association data, protein-protein interactions, and human pancreatic islet gene expression. Diabetes, 61(4), 954–962.
Friend, S. H., & Ideker, T. (2011). POINT: are we prepared for the future doctor visit? Nat Biotech, 29(3), 215–218.
Kohane, I. S., & Margulies, D. M. (2011). COUNTERPOINT: do not opine before it’s time. Nat Biotech, 29(3), 218–219.
Shannon, P., Markiel, A., Ozier, O., Baliga, N. S., Wang, J. T., Ramage, D., Amin, N., Schwikowski, B., & Ideker, T. (2003). Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res, 13(11), 2498–2504.
Bhavnani, S., Ganesan, A., Hall, T., Maslowski, E., Eichinger, F., Martini, S., Saxman, P., Bellala, G., & Kretzler, M. (2010). Discovering hidden relationships between renal diseases and regulated genes through 3D network visualizations. BMC Res Notes, 3(1), 296.
Chen, R., Mias, G. I., Li-Pook-Than, J., Jiang, L., Lam, H. Y. K., Chen, R., Miriami, E., Karczewski, K. J., Hariharan, M., Dewey, F. E., Cheng, Y., Clark, M. J., Im, H., Habegger, L., Balasubramanian, S., O’Huallachain, M., Dudley, J. T., Hillenmeyer, S., Haraksingh, R., Sharon, D., Euskirchen, G., Lacroute, P., Bettinger, K., Boyle, A. P., Kasowski, M., Grubert, F., Seki, S., Garcia, M., Whirl-Carrillo, M., Gallardo, M., Blasco, M. A., Greenberg, P. L., Snyder, P., Klein, T. E., Altman, R. B., Butte, A. J., Ashley, E. A., Gerstein, M., Nadeau, K. C., Tang, H., & Snyder, M. (2012). Personal omics profiling reveals dynamic molecular and medical phenotypes. Cell, 148(6), 1293–1307.
Atkinson, A. J., Colburn, W. A., DeGruttola, V. G., DeMets, D. L., Downing, G. J., Hoth, D. F., Oates, J. A., Peck, C. C., Schooley, R. T., Spilker, B. A., Woodcock, J., Zeger, S. L., & Biomarkers Definitions Working Group. (2001). Biomarkers and surrogate endpoints: preferred definitions and conceptual framework. Clin Pharmacol Ther, 69(3), 89–95.
Mischak, H., Allmaier, G., Apweiler, R., Attwood, T., Baumann, M., Benigni, A., et al. (2010). Recommendations for biomarker identification and qualification in clinical proteomics. Sci Transl Med, 2(46), 46ps42.
Matheis, K., Laurie, D., Andriamandroso, C., Arber, N., Badimon, L., Benain, X., Bendjama, K., Clavier, I., Colman, P., Firat, H., Goepfert, J., Hall, S., Joos, T., Kraus, S., Kretschmer, A., Merz, M., Padro, T., Planatscher, H., Rossi, A., Schneiderhan-Marra, N., Schuppe-Koistinen, I., Thomann, P., Vidal, J.-M., & Molac, B. (2011). A generic operational strategy to qualify translational safety biomarkers. Drug Discov Today, 16(13–14), 600–608.
Hill, A. B. (1965). The environment and disease: association or causation? Proc R Soc Med, 58(5), 295–300.
Slocum, J. L., Heung, M., & Pennathur, S. (2012). Marking renal injury: can we move beyond serum creatinine? Transl Res, 159(4), 277–289.
Ju, W., Smith, S., & Kretzler, M. (2012). Genomic biomarkers for chronic kidney disease. Transl Res, 159(4), 290–302.
Sotiriou, C., & Piccart, M. J. (2007). Taking gene-expression profiling to the clinic: when will molecular signatures become relevant to patient care? Nat Rev Cancer, 7(7), 545–553.
Dabbs, D. J., Klein, M. E., Mohsin, S. K., Tubbs, R. R., Shuai, Y., & Bhargava, R. (2011). High false-negative rate of HER2 quantitative reverse transcription polymerase chain reaction of the oncotype DX test: an independent quality assurance study. J Clin Oncol, 29(32), 4279–4285.
Ignatiadis, M., & Sotiriou, C. (2012). Breast cancer: should we assess HER2 status by Oncotype DX®? Nat Rev Clin Oncol, 9(1), 12–14.
Coombes, K. R., Wang, J., & Baggerly, K. A. (2007). Microarrays: retracing steps. Nat Med, 13(11), 1276–1277.
Baggerly, K. A., & Coombes, K. R. (2009). Deriving chemosensitivity from cell lines: forensic bioinformatics and reproducible research in high-throughput biology. Ann Appl Stat, 3(4), 1309–1334.
No authors listed (2010). Retraction. Pharmacogenomic strategies provide a rational approach to the treatment of cisplatin-resistant patients with advanced cancer. J Clin Oncol 28(35):5229–5229
Potti, A., Mukherjee, S., Petersen, R., Dressman, H. K., Bild, A., Koontz, J., et al. (2011). Retraction: a genomic strategy to refine prognosis in early-stage non-small-cell lung cancer. N Engl J Med 2006;355:570–80. New Engl J Med, 364(12), 1176–1176.
Potti, A., Dressman, H. K., Bild, A., Riedel, R. F., Chan, G., Sayer, R., Cragun, J., Cottrill, H., Kelley, M. J., Petersen, R., Harpole, D., Marks, J., Berchuck, A., Ginsburg, G. S., Febbo, P., Lancaster, J., & Nevins, J. R. (2011). Retraction: genomic signatures to guide the use of chemotherapeutics. Nat Med, 17(1), 135–135.
Hayden, E.C. (2012) Lapses in oversight compromise omics results. US board calls for tighter control of test-based data. Nature [23 March 2012].
Curtis, C., Shah, S. P., Chin, S.-F., Turashvili, G., Rueda, O.M., Dunning, M.J., et al. (2012) The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature (in press) (Published online, April 18, 2012).
Monnier, V. M., Vishwanath, V., Frank, K. E., Elmets, C. A., Dauchot, P., & Kohn, R. R. (1986). Relation between complications of type i diabetes mellitus and collagen-linked fluorescence. New Engl J Med, 314(7), 403–408.
Sun, J. K., Keenan, H. A., Cavallerano, J. D., Asztalos, B. F., Schaefer, E. J., Sell, D. R., Strauch, C. M., Monnier, V. M., Doria, A., Aiello, L. P., & King, G. L. (2011). Protection from retinopathy and other complications in patients with type 1 diabetes of extreme duration. The Joslin 50-Year Medalist Study. Diabetes Care, 34(4), 968–974.
Mirel, B., Eichinger, F., Nair, V., & Kretzler, M. (2009). Integrating automated workflows, human intelligence and collaboration. Summit on Translat Bioinforma, 2009, 79–83.
Derry, J. M. J., Mangravite, L. M., Suver, C., Furia, M. D., Henderson, D., Schildwachter, X., Bot, B., Izant, J., Sieberts, S. K., Kellen, M. R., & Friend, S. H. (2012). Developing predictive molecular maps of human disease through community-based modeling. Nat Genet, 44(2), 127–130.
Acknowledgments
This work was supported by grants from the Juvenile Diabetes Research Foundation to M.K. and S.P., the German Research Foundation to C.V.K. (KO 4266/1-1), the National Institutes of Health, National Institute of Diabetes, and Digestive and Kidney Disease (P30-DK-081943 to F.C.B. and M.K; DK083912 to M.K.; DK089503, DK094292 to S.P.; DK082841 to M.K. and S.P.). The authors appreciate the assistance of Dr. Jeff B. Hodgin (Department of Pathology, University of Michigan) in providing the histological images. We further thank Chandra Hall for carefully proofreading the manuscript and the editor and referees for helpful comments and suggestions on an earlier version of the manuscript. The authors apologize to those colleagues whose work we were unable to cite in this review due to space limitation.
Conflict of Interest
The authors declare no competing financial interest.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Komorowsky, C.V., Brosius, F.C., Pennathur, S. et al. Perspectives on Systems Biology Applications in Diabetic Kidney Disease. J. of Cardiovasc. Trans. Res. 5, 491–508 (2012). https://doi.org/10.1007/s12265-012-9382-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12265-012-9382-7