Summary
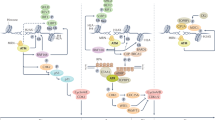
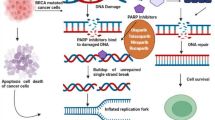
The efficacy of cancer chemotherapy and radiotherapy relies on generation of DNA damage. Since intrinsic DNA repair pathways enable cancer cells to survive by repairing these damaged lesions, inactivation of DNA repair coupled with chemotherapy and radiotherapy has a potential to enhance the effect of these therapies. Small molecule compounds that inhibit specific DNA repair proteins have been developed, and early clinical trials are ongoing. While DNA repair inhibitors have been tried mostly as a part of combination therapies with cytotoxic agents, recent reports highlighted a new concept in cancer therapy where DNA repair inhibitors could be used as single agents to selectively kill cancer cells. This concept is based on the findings that cancer cells are frequently defective in particular DNA repair pathway(s) and the presumption that inhibition of the compensatory repair pathway(s) in such cells might be useful to kill them. For example, poly(ADP-ribose) polymerase (PARP) plays a critical role in DNA base-excision repair, and inactivation of this protein increases the number of single-strand breaks, leading to double-strand breaks that require to be repaired by homologous recombination (HR) mediated by BRCA1 and BRCA2. Recently, BRCA1- or BRCA2-defective tumour cells were shown to be sensitive to PARP inhibitors alone. This treatment may be tumour-specific because only the BRCA1−/− or BRCA2−/− tumours in the BRCA1+/− or BRCA2+/− patients are completely defective in HR repair. The following short review aims at summarizing the basic mechanisms of DNA repair and the therapeutic options using DNA repair inhibitors in cancer therapy.
Similar content being viewed by others
References
Khanna KK, Jackson SP. DNA double-strand breaks: signaling, repair and the cancer connection. Nat Genet, 27: 247–254, 2001
van Gent DC, Hoeijmakers JH, Kanaar R. Chromosomal stability and the DNA double-stranded break connection. Nat Rev Genet, 2: 196–206, 2001
Helleday T, Petermann E, Lundin C, et al. DNA repair pathways as targets for cancer therapy. Nat Rev Cancer, 8: 193–204, 2008
Martin SA, Lord CJ, Ashworth A. DNA repair deficiency as a therapeutic target in cancer. Curr Opin Genet Dev, 18: 80–86, 2008
Helleday T, Lo J, van Gent DC, Engelward BP. DNA double-strand break repair: from mechanistic understanding to cancer treatment. DNA Repair, 6: 923–935, 2007
Sung P, Klein H. Mechanism of homologous recombination: mediators and helicases take on regulatory functions. Nat Rev Mol Cell Biol, 7: 739–750, 2006
Sharma RA, Dianov GL. Targeting base excision repair to improve cancer therapies. Mol Aspects Med, 28: 345–374, 2007
Sugasawa K, Okamoto T, Shimizu Y, et al. A multistep damage recognition mechanism for global genomic nucleotide excision repair. Genes Dev, 15: 507–521, 2001
Lindahl T, Demple B, Robins P. Suicide inactivation of the E. coli O6-methylguanine-DNA methyltransferase. EMBO J, 1: 1359–1363, 1982
Dronkert MLG, Kanaar R. Repair of DNA interstrand cross-links. Mutat Res, 486: 217–247, 2001
Pommier Y. Topoisomerase I inhibitors: camptothecins and beyond. Nature Rev Cancer, 6: 789–802, 2006
Arnaudeau C, Lundin C, Helleday T. DNA double-strand breaks associated with replication forks are predominantly repaired by homologous recombination involving an exchange mechanism in mammalian cells. J Mol Biol, 307: 1235–1245, 2001
Patel KJ, Joenje H. Fanconi anemia and DNA replication repair. DNA Repair (Amst), 6: 885–890, 2007
Hanada K, Budzowska M, Davies SL, et al. The structure-specific endonuclease Mus81 contributes to replication restart by generating double-strand DNA breaks. Nat Struct Mol Biol, 14: 1096–1104, 2007
Wu L, Hickson ID. DNA helicases required for homologous recombination and repair of damaged replication forks. Annu Rev Genet, 40: 279–306, 2006
Middleton MR, Margison GP. Improvement of chemotherapy efficacy by inactivation of a DNA-repair pathway. Lancet Oncol, 4: 37–44, 2003
Durkacz BW, Omidiji O, Gray DA, et al. (ADP-ribose)n participates in DNA excision repair. Nature, 283: 593–596, 1980
de Murcia JM, Niedergang C, Trucco C, et al. Requirement of poly(ADP-ribose) polymerase in recovery from DNA damage in mice and in cells. Proc Natl Acad Sci USA, 94: 7303–7307, 1997
Schultz N, Lopez E, Saleh-Gohari N, et al. Poly(ADP-ribose) polymerase (PARP-1) has a controlling role in homologous recombination. Nucleic Acids Res, 31: 4959–4964, 2003
Ratnam K, Low JA. Current development of clinical inhibitors of poly(ADP-ribose) polymerase in oncology. Clin Cancer Res, 13: 1383–1388, 2007
Donawho CK, Luo Y, Luo Y. ABT-888, an orally active poly(ADP-ribose) polymerase inhibitor that potentiates DNA-damaging agents in preclinical tumor models. Clin Cancer Res, 13: 2728–2737, 2007
Adimoolam S, Sirisawad M, Chen J, et al. HDAC inhibitor PCI-24781 decreases RAD51 expression and inhibits homologous recombination. Proc Natl Acad Sci USA, 104: 19482–19487, 2007
Ismail IH, Martensson S, Moshinsky D, et al. SU11752 inhibits the DNA-dependent protein kinase and DNA double-strand break repair resulting in ionizing radiation sensitization. Oncogene, 23: 873–882, 2004
Zhao Y, Thomas HD, Batey MA, et al. Preclinical evaluation of a potent novel DNA-dependent protein kinase inhibitor NU7441. Cancer Res, 66: 5354–5362, 2006
Marone R, Cmiljanovic V, Giese B, et al. Targeting phosphoinositide 3-kinase: moving towards therapy. Biochim Biophys Acta, 1784: 159–185, 2007
Ranson M, Middleton MR, Bridgewater J, et al. Lomeguatrib, a potent inhibitor of O6-alkylguanine-DNA-alkyltransferase: phase I safety, pharmacodynamic, and pharmacokinetic trial and evaluation in combination with temozolomide in patients with advanced solid tumors. Clin Cancer Res, 12: 1577–1584, 2006
Ranson M, Hersey P, Thompson D, et al. Randomized trial of the combination of lomeguatrib and temozolomide compared with temozolomide alone in chemotherapy naive patients with metastatic cutaneous melanoma. J Clin Oncol, 25: 2540–2545, 2007
Khan OA, Ranson M, Michael M, et al. A phase II trial of lomeguatrib and temozolomide in metastatic colorectal cancer. Br J Cancer, 98: 1614–1618, 2008
Lara PN Jr, Mack PC, Synold T, et al. The cyclin-dependent kinase inhibitor UCN-01 plus cisplatin in advanced solid tumors: a California cancer consortium phase I pharmacokinetic and molecular correlative trial. Clin Cancer Res, 11: 4444–4450, 2005
Hotte SJ, Oza A, Winquist EW, et al. Phase I trial of UCN-01 in combination with topotecan in patients with advanced solid cancers: a Princess Margaret Hospital Phase II Consortium study. Ann Oncol, 17: 334–340, 2006
Hickson I, Zhao Y, Richardson CJ, et al. Identification and characterization of a novel and specific inhibitor of the ataxia-telangiectasia mutated kinase ATM. Cancer Res, 64: 9152–9159, 2004
Farmer H, McCabe N, Lord CJ, et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature, 434: 917–921, 2005
Bryant HE, Schultz N, Thomas HD, et al. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature, 434: 913–917, 2005
Sakai W, Swisher EM, Karlan BY, et al. Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers. Nature, 28: 1116–1120, 2008
Edwards SL, Brough R, Lord CJ, et al. Resistance to therapy caused by intragenic deletion in BRCA2. Nature, 451: 1111–1115, 2008
Swisher EM, Sakai W, Karlan BY, et al. Secondary BRCA1 mutations in BRCA1-mutated ovarian carcinomas with platinum resistance. Cancer Res, 68: 2581–2586, 2008
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Hosoya, N., Miyagawa, K. Clinical importance of DNA repair inhibitors in cancer therapy. memo 2, 9–14 (2009). https://doi.org/10.1007/s12254-008-0081-7
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s12254-008-0081-7