Abstract
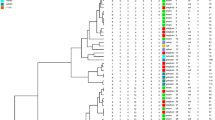
The aim of this study was to analyse genotypes, antimicrobial susceptibility patterns and serotypes in Pseudomonas aeruginosa clinical strains, including the clonal dissemination of particular strains throughout various intensive care units in one medical centre. Using random amplified polymorphic DNA (RAPD–PCR) and P. aeruginosa antisera, 22 different genotypes and 8 serotypes were defined among 103 isolates from 48 patients. No direct association between P. aeruginosa strain genotypes and serotypes was observed. RAPD typing in strains with the same serotype revealed different genotypes and, on the contrary, most strains with a different serotype displayed the same amplification pattern. The resulting banding patterns showed a high degree of genetic heterogeneity among all isolates from the patients examined, suggesting a non-clonal relationship between isolates from these patients. A higher degree of antibiotic resistance and stronger biofilm production in common genotypes compared to rare ones and genetic homogeneity of the most resistant strains indicated the role of antibiotic pressure in acquiring resistant and more virulent strains in our hospital. In conclusion, genetic characterisation of P. aeruginosa strains using RAPD method was shown to be more accurate in epidemiological analyses than phenotyping.
Similar content being viewed by others
References
Ahmadi H, Maleknia S, Tabaraie B et al (2010) Serotyping and cross-reactivity’s between different Pseudomonas aeruginosa isolates prevalent in Iran. Iran J Microbiol 2:85–88
Barnini S, Dodi C, Campa M (2004) Enhanced resolution of random amplified polymorphic DNA genotyping of Pseudomonas aeruginosa. Lett Appl Microbiol 39:274–277
Bouza E, Garcia-Garrote F, Cercenado E et al (1999) Pseudomonas aeruginosa: a survey of resistance in 136 hospitals in Spain. Antimicrob Agents Chemother 43:981–982
Dean CR, Franklund CV, Retief JD et al (1999) Characterization of the serogroup O11 O-antigen locus of Pseudomonas aeruginosa PA103. J Bacteriol 181:4275–4284
Delden CH, Iglewski BH (1998) Cell-to-cell signalling and Pseudomonas aeruginosa infections. Emerg Infect Dis 4:551–560
Deligianni E, Pattison S, Berrar D et al (2010) Pseudomonas aeruginosa cystic fibrosis isolates of similar RAPD genotype exhibit diversity in biofilm forming ability in vitro. BMC Microbiol 10:1471–2180
Extremina CI, Costa L, Aguiar AI, Peixe L, Fonseca AP (2011) Optimization of processing conditions for the quantification of enterococci biofilms using microtitre-plates. J Microbiol Methods 84:167–173
Fonseca AP, Correia P, Sousa JC et al (2007) Association patterns Pseudomonas aeruginosa clinical isolates as revealed by virulence traits, antibiotic resistance, serotype and genotype. FEMS Immunol Med Microbiol 51:505–516
Gasink LB, Fishman NO, Weiner MG et al (2006) Fluoroquinolone-resistant Pseudomonas aeruginosa: assessment of risk factors and clinical impact. Am J Med 119:526.e19–526.e25
Kalferstova L, Vilimovska Dedeckova K, Antuskova M, Melter O, Drevinek P (2016) How and why to monitor Pseudomonas aeruginosa infections in the long term at a cystic fibrosis centre. J Hosp Infect 92:54–60
Koutsogiannou M, Drougka E, Liakopoulos A et al (2013) Spread of multidrug-resistant Pseudomonas aeruginosa clones in a university hospital. J Clin Microbiol 51:665–668
Lambert PA (2002) Mechanism of antibiotic resistance in Pseudomonas aeruginosa. J R Soc Med 95:22–26
Mahenthiralingam E, Campbell M, Foster J et al (1996) Random amplified polymorphic DNA typing of Pseudomonas aeruginosa isolates recovered from patients with cystic fibrosis. J Clin Microbiol 34:1129–1135
Matar GM, Chaar MH, Araj GF et al (2005) Detection of highly prevalent and potentially virulent strain of Pseudomonas aeruginosa from nosocomial infections in a medical center. BMC Microbiol 5:1471–2180
Müller-Premru M, Gubina M (2000) Serotype, antimicrobial susceptibility and clone distribution of Pseudomonas aeruginosa in a university hospital. Zentralblatt für Bakteriologie: Int J Med Microbiol 289(8):57–67
Nanvazadeh F, Khosravi AD, Zolfaghari MR, Parhizgari N (2013) Genotyping of Pseudomonas aeruginosa strains isolated from burn patients by RAPD-PCR. Burns 39:1409–1413
Nazik H, Ongen B, Erturan Z et al (2007) Genotype and antibiotic susceptibility patterns of Pseudomonas aeruginosa and Stenotrophomonas maltophilia isolated from cystic fibrosis patients. Jpn J Infect Dis 60:82–86
Nemec A, Krizova L, Maixnerova M et al (2010) Multidrug-resistant epidemic clones among bloodstream isolates of Pseudomonas aeruginosa in the Czech Republic. Res Microbiol 161:234–242
Nguyen LH, Hsu DI, Ganapathy V et al (2008) Reducing empirical use of fluoroquinolones for Pseudomonas aeruginosa infections improves outcome. J Antimicrob Chemother 61:714–720
Nouér SA, Nucci M, de Oliveira MP et al (2005) Risk factors for acquisition of multidrug-resistant Pseudomonas aeruginosa producing SPM metallo-β-lactamase. Antimicrob Agents Chemother 49:3663–3667
Obritsch MD, Fish DN, MacLaren R et al (2005) Nosocomial infections due to multidrug-resistant Pseudomonas aeruginosa: epidemiology and treatment options. Pharmacotherapy 25:1353–1364
Ortiz-Herrera M, Gerónimo-Gallegos A, Cuevas-Schacht F et al (2004) RAPD-PCR characterization of Pseudomonas aeruginosa strains obtained from cystic fibrosis patients. Salud Publ Mex 46:149–157
Ouellet MM, Leduc A, Nadeau C, Barbeau J, Charette SJ (2015) Pseudomonas aeruginosa isolates from dental unit waterlines can be divided in two distinct groups, including one displaying phenotypes similar to isolates from cystic fibrosis patients. Front Microbiol 5:802. doi:10.3389/fmicb.2014.00802
Paramythiotou E, Lucet JC, Timsit JF et al (2004) Acquisition of multidrug-resistant Pseudomonas aeruginosa in patients in intensive care units: role of antibiotics with antipseudomonal activity. Clin Infect Dis 38:670–677
Patzer J, Dzierzanowska D (1991) The resistance patterns and serotypes of Pseudomonas aeruginosa strains isolated from children. J Antimicrob Chemother 28:869–875
Pier GB (2007) Pseudomonas aeruginosa lipopolysaccharide: a major virulence factor, initiator of inflammation and target for effective immunity. Int J Med Microbiol 297:277–295
Pirnay JP, De Vos D, Cochez C et al (2003) Molecular epidemiology of Pseudomonas aeruginosa colonization in a burn unit: persistence of a multidrug-resistant clone and a silver sulfadiazine-resistant clone. J Clin Microbiol 41:1192–1202
Renders N, Römling U, Verbrugh H et al (1996) Comparative typing of Pseudomonas aeruginosa by random amplification of polymorphic DNA or pulsed-field gel electrophoresis of DNA macrorestriction fragments. J Clin Microbiol 34:3190–3195
Stepanović S, Vuković D, Hola V et al (2007) Quantification of biofilm in microtiter plates: overview of testing conditions and practical recommendations for assessment of biofilm production by staphylococci. APMIS 115:891–899
Tam VH, Rogers CA, Chang KT et al (2010) Impact of multidrug-resistant Pseudomonas aeruginosa bacteremia on patient outcomes. Antimicrob Agents Chemother 54:3717–3722
Trautmann M, Bauer C, Schumann C et al (2006) Common RAPD pattern of Pseudomonas aeruginosa from patients and tap water in a medical intensive care unit. Int J Hyg Environ Health 209:325–331
Wolska K, Kot B, Jakubczak A (2011) Phenotypic and genotypic diversity of Pseudomonas aeruginosa strains isolated from hospitals in Siedlce (Poland). BJM 43:274–282
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work was supported by the grant of Ministry of Health, CZ, [16-31593A], and the internal grant of the Centre for Cardiovascular Surgery and Transplantation Brno, CZ [201404].
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Vaněrková, M., Mališová, B., Kotásková, I. et al. Biofilm formation, antibiotic susceptibility and RAPD genotypes in Pseudomonas aeruginosa clinical strains isolated from single centre intensive care unit patients. Folia Microbiol 62, 531–538 (2017). https://doi.org/10.1007/s12223-017-0526-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-017-0526-7