Abstract
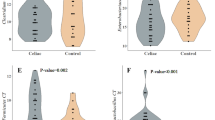
The fecal short-chain fatty acids concentration was higher (154 ± 46.9 mmol/L) in childhood patients than in healthy children (96.6 ± 19.2 mmol/L). On the other hand, pH values were nonsignificantly lower in patients stool (6.78 ± 0.75 vs. children 7.42 ± 0.74). Using denaturing gradient gel electrophoresis specific for total bacteria, lactobacilli and bifidobacteria the microbial population was characterized in fecal samples and in duodenal biopsies. Bacteria adhering to duodenal biopsies were not dominating in stool samples. More than 50 % of detected bacterial species belonged to as yet uncultured strains.
Similar content being viewed by others
References
Abdulkarim A. S., Murray J.A.: Review article: the diagnosis of celiac disease. Aliment.Pharmacol.Ther. 17, 987–995 (2003).
Flint H.J., Duncan S.H., Scott K.P., Louis P.: Interactions and competition within the microbial community of the human colon: links between diet and health. Environ.Microbiol. 9, 1101–1111 (2007).
Forsberg G., Fahlgren A., Horstedt P.: Presence of bacteria and innate immunity of intestinal epithelium in childhood celiac disease. Am.J.Gastroenterol. 99, 894–904 (2004).
Forster R.J., Gong J., Teather R.M.: Group-specific 16S rDNA hybridization probes for determinative and community structure studies of Butyrivibrio fibrisolvens in the rumen. Appl.Environ.Microbiol. 63, 1256–1260 (2007).
Kaufmann P., Pfefferkorn A., Teuber M., Meile L.: Identification and quantification of Bifidobacterium species isolated from food with genus-specific 16S rRNA-targeted probes by colony hybridization and PCR. Appl.Environ.Microbiol. 62, 3668–3672 (1997).
Kopečný J., Hajer J., Mrázek J.: Detection of cellulolytic bacteria from the human colon. Folia Microbiol. 49, 175–178 (2004).
Kopečný J., Mrázek J., Fliegerová K., Kott T.: The effect of gluten-free diet on microbes in the colon. Folia Microbiol. 51, 287–290 (2006).
Leffler D., Saha S., Farrell R.J.: Celiac disease. Am.J.Managed Care 9, 825–831 (2003).
Muyzer G., de Waal E.C., Uitterlinden A.G.: Profiłing of complex microbial population by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl.Environ.Microbiol. 59, 695–700 (1993).
Shan L., Molberg O., Parrot I., Hausch F., Filiz F., Gray G.M., Sollid F., Khosla C.: Structural basis for gluten intolerance in celiac sprue. Science 297, 2275–2279 (2002).
Sollid L.M., Gray G.M.: A role for bacteria in celiac disease? Am.J.Gastroenterol. 99, 905–906 (2004).
Temmerman R., Scherlinck I., Huys G., Swings J.: Culture-independent analysis of probiotic products by denaturing gradient gel electrophoresis. Appl.Environ.Microbiol. 69, 220–226 (2003).
Tursi A., Brandimarte G., Giorgetti G.M.: High prevalence of small intestinal bacterial overgrowth in celiac patients with persistence of gastrointestinal symptoms after gluten withdrawal. Am.J.Gastroenterol. 98, 839–843 (2003).
Wyeth J.W., Peters S.G., Fisher A., Yeong M.L., Stace N.H., Pomare E.W.: Gluten challenge in celiac-disease alters small-bowel and colonic pH. Gastroenterology 104 (Suppl.), a291–a291 (1993).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kopečný, J., Mrázek, J., Fliegerová, K. et al. The intestinal microflora of childhood patients with indicated celiac disease. Folia Microbiol 53, 214–216 (2008). https://doi.org/10.1007/s12223-008-0028-8
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-008-0028-8