Abstract
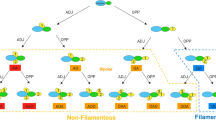
Many yeasts can produce filamentous elongated cells identifiable as hyphae, pseudohyphae or invasive filaments. Filament formation has been understood as a foraging response that occurs in nutrient-poor conditions. However, fusel alcohols were observed to induce filament formation in rich nutrient conditions in every yeast species examined. Fusel alcohols, e.g., 3-methyl-1-butanol (3Me-BuOH; ‘isoamyl alcohol’), 2-methyl-1-propanol (isobutyl alcohol), (−)-2-methyl-1-butanol (‘active amyl alcohol’), 2-phenylethanol and 3-(2-hydroxyethyl)indole (tryptophol) (the end products of leucine, valine, isoleucine, phenylalanine and tryptophan catabolism, respectively) are the end products of amino acid catabolism that accumulate when nutrients become limiting. Thus, yeast responds to its own metabolic by-products. Considerable effort was made to define the cell biological and biochemical changes that take place during 3Me-BuOH-induced filam entation. In Saccharomyces cerevisiae filaments contain significantly greater mitochondrial mass and increased chitin content in comparison with yeast-form cells. The global transcriptional response of S. cerevisiae during the early stages of 3Me-BuOH-induced filament formation has been described. Four ORFs displayed very significant (more than 10-fold) increases in their RNA species, and 12 ORFs displayed increases in transcription of more than 5-fold. The transcription of five genes (all of which encode transporters) decreased by similar amounts. Where examined, the activity of the proteins encoded reflected the transcriptional pattern of their respective mRNAs. To understand this regulation, studies were performed to see whether deletion or overexpression of key genes affects the ability to filament and invade solid YEPD medium. This has led to identification of those proteins that are essential for filament formation, repressors and those which are simply not required. It also leads to the conclusion that 3Me-BuOH-induced filament formation is not a foraging response but a response to reduced growth rate.
Similar content being viewed by others
Abbreviations
- 3ME-BuOH:
-
3-methyl-1-butanol (‘isoamyl alcohol’)
- PDR:
-
pleiotropic drug resistance
- YEPD:
-
medium (%, W/V): bacteriological peptone 2, dextrose (glucose) 2, yeast extract 1; also contains 0.01% of both adenine and uracil
References
Aguilera J., Van Dijken J.P., Dewinde J.H., Pronk J.T.: Carbonic anhydrase (Nce103): an essential biosynthetic enzyme for growth of Saccharomyces cerevisiae at atmospheric carbon dioxide pressure. Biochem.J. 391, 311–316 (2005).
Amoah-Buahin E., Bone N., Armstrong J.: Hyphal growth in the fission yeast Schizosaccharomyces pombe. Euk.Cell 4, 1287–1297 (2005).
Ashe M.P., Slaven J.W., Delong S.K., Ibrahimo S., Sachs A.B.: A novel eIF2B-dependent mechanism of translational control in yeast as a response to fusel alcohols. EMBO J. 20, 6464–6474 (2001).
Buziol S., Becker J., Baumeister A, Jung S., Mauch K., Reuss M., Boles E.: Determination of in vivo kinetics of the starvation-induced Hxt5 glucose transporter of Saccharomyces cerevisiae. FEMS Yeast Res. 2, 283–291 (2002).
Chelstowska A., Liu Z., Jia Y., Amberg D., Butow R.A.: Signaling between the mitochondria and the nucleus regulates the expression of a new d-lactate dehydrogenase activity in yeast. Yeast 15, 1377–1391 (1999).
Chen H., Fink G.R.: Feedback control of morphogenesis in fungi by aromatic alcohols. Genes Dev. 20, 1150–1161 (2006).
Chen S., Brockenbrough J.S., Dove J.E., Aris J.P.: Homocitrate synthase is located in the nucleus in the yeast Saccharomyces cerevisiae. J.Biol.Chem. 272, 10839–10846 (1997).
Chen C.-N., Porubleva L., Shearer G., Svrakic M., Holden L. G., Dover J.L., Johnston M., Chitnis P.R., Kohl D.H.: Associating protein activities with their genes: rapid identification of a gene encoding a methylglyoxal reductase in the yeast Saccharomyces cerevisiae. Yeast 20, 545–554 (2003).
Chen H., Fujita M., Feng Q., Clardy J., Fink G.R.: Tyrosol is a quorum-sensing molecule in Candida albicans. Proc.Nat.Acad.Sci.USA 101, 5048–5052 (2004).
Destruelle M., Kolzer H., Klionsky D.J.: Identification and characterization of a novel yeast gene: the YGP1 gene product is a highly glycosylated secreted protein that is synthesized in response to nutrient limitation. Mol.Cell.Biol. 14, 2740–2754 (1994).
Dickinson J.R.: Irreversible formation of pseudohyphase by haploid Saccharomyces cerevisiae. FEMS Microbiol.Lett. 119, 99–104 (1994).
Dickinson J.R.: ’Fusel’ alcohols induce hyphal-like extensions and pseudohyphal formation in yeasts. Microbiology 142, 1391–1397 (1996).
Dickinson J.R.: Life cycle and morphogenesis, pp. 1–19 in J.R. Dickinson, M. Schweizer (Eds): The Metabolism and Molecular Physiology of Saccharomyces cerevisiae, 2nd ed. CRC Press, Boca Raton 2004.
Dickinson J.R., Lanterman M.M., Danner D.J., Pearson B.M., Sanz P., Harrison S.J., Hewlins M.J.E.: A 13C nuclear magnetic resonance investigation of the metabolism of leucine to isoamyl alcohol in Saccharomyces cerevisiae. J.Biol.Chem. 272, 26871–26878 (1977).
Dickinson J.R., Harrison S.J., Hewlins M.J.E.: An investigation into the metabolism of valine to isobutyl alcohol in Saccharomyces cerevisiae. J.Biol.Chem. 273, 25751–25756 (1998).
Dickinson J.R., Harrison S.J., Dickinson J.A., Hewlins M.J.E.: An investigation of the metabolism of isoleucine to active amyl alcohol in Saccharomyces cerevisiae. J.Biol.Chem. 275, 10937–10942 (2000).
Dickinson J.R., Salgado L.E.J., Hewlins M.J.E.: The catabolism of amino acids to long-chain and complex alcohols in Saccharomyces cerevisiae. J.Biol.Chem. 278, 8028–8034 (2003).
Diderich J.A., Schuurmans J.M., Van Gaalen M.C., Kruckeberg A.L., Van Dam K.: A functional analysis if the hexose transporter homologue HXT5 in Saccharomyces cerevisiae. Yeast 18, 1515–1524 (2001).
Esposito R.E., Klapholz S.: Meiosis and ascospore development, pp. 211–287 in J.N. Strathern, E.W. Jones, J.R. Broach (Eds): The Molecular Biology of the Yeast Saccharomyces: Life Cycle and Inheritance. Cold Spring Harbor Laboratory, Cold Spring Harbor 1981.
Feller A., Ramos F., Piérard A., Dubois E.: In Saccharomyces cerevisiae, feedback inhibition of homocitrate synthase isoenzymes by lysine modulates the activation of LYS gene expression by Lys14p. Eur.J.Biochem. 261, 163–170 (1999).
Fuge E.K., Braun E.L., Werner-Washbourne M.: Protein synthesis in long-term stationary-phase cultures of Saccharomyces cerevisiae. J.Bacteriol. 176, 5802–5813 (1994).
Gimeno C.J., Ljungdahl P.O., Styles C.A., Fink G.R.: Unipolar cell divisions in the yeast Saccharomyces cerevisiae lead to filamentous growth: regulation by starvation and RAS. Cell 68, 1077–1090 (1992).
Hancock L.C., Behta R.P., Lopes J.M.: Genomic analysis of the Opi− phenotype. Genetics 173, 621–634 (2006).
Hauser M., Horn P., Tournu H., Hauser N.C., Hoheisel J.D., Brown A.J.P., Dickinson J.R.: A transcriptome analysis of isoamyl alcohol-induced filamentation in yeast reveals a novel role for Gre2p as isovaleraldehyde reductase. FEMS Yeast Res. 7, 84–92 (2007).
Hayashi M., Ohkuni K., Yamashita I.: An extracellular meiosis-promoting factor in Saccharomyces cerevisiae. Yeast 14, 617–622 (1998).
Jiang Y.W., Kang C.M.: Induction of S. cerevisiae filamentous differentiation by slowed DNA synthesis involves Mec1, Rad53 and Swe1 checkpoint proteins. Mol.Biol.Cell 14, 5116–5124 (2002).
Kern K., Nunn C.D., Pichova A., Dickinson J.R.: Isoamyl alcohol-induced morphological change in Saccharomyces cerevisiae involves increases in mitochondria and cell wall chitin content. FEMS Yeast Res. 5, 43–49 (2004).
Kron S.J., Gow N.A.R.: Budding yeast morphogenesis: signaling, cytoskeleton and cell cycle. Curr.Opin.Cell Biol. 7, 845–855 (1995).
La Valle R., Wittenberg C.: A role for the Swe1 checkpoint kinase during filamentous growth of Saccharomyces cerevisiae. Genetics 158, 549–562 (2001).
Liger D., Quevillon-Cheruel S., Sorel I., Bremang M., Blondeau K., Aboulfath I., Janin J., Van Tilbeurgh H., Leulliot N.: Crystal structure of YHI9, the yeast member of the phenazine biosynthesis PhzF enzyme superfamily. Proteins 60, 778–786 (2005).
Lorenz M.C., Heitman J.: The MEP2 ammonium permease regulates pseudohyphal differentiation in Saccharomyces cerevisiae. EMBO J. 17, 1236–1247 (1998).
Lorenz M.C., Cutler N.S., Heitman J.: Characterization of alcohol-induced filamentous growth in Saccharomyces cerevisiae. Mol.Biol.Cell 11, 183–199 (2000).
Martinez-Anaya C., Dickinson J.R., Sudbery P.E.: In yeast, the pseudohyphal phenotype induced by isoamyl alcohol results from the operation of the morphogenesis checkpoint. J.Cell Sci. 116, 3423–3431 (2003).
Nyunoya H., Lusty C.J.: Sequence of the small subunit of yeast carbamyl phosphate synthetase and identification of its catalytic domain. J.Biol.Chem. 259, 9790–9798 (1984).
Ohkuni K., Hayashi M., Yamashita I.: Bicarbonate-mediated social communication stimulates meiosis and sporulation of Saccharomyces cerevisiae. Yeast 14, 623–631 (1998).
Påhlman A.-K., Granath K., Ansell R., Hohmann S., Adler L.: The yeast glycerol-3-phosphatases Gpp1 and Gpp2 are required for glycerol biosynthesis and differentially involved in the cellular responses to osmotic, anaerobic and oxidative stress. J.Biol.Chem. 276, 3555–3563 (2001).
Paulsen I.T., Sliwinski M.K., Nelissen B., Goffeau A., Saier M.H. Jr.: Unified inventory of established and putative transporters encoded within the complete genome of Saccharomyces cerevisiae. FEBS Lett. 430, 116–125 (1998).
Perry J.R., Basrai M.A., Steiner H.-Y., Naider F., Becker J.M.: Isolation and characterization of a Saccharomyces cerevisiae peptide transporter gene. Mol.Cell.Biol. 14, 104–115 (1994).
Regenberg B., During-Olsen L., Kieland-Brandt M.C., Holmberg S.: Substrate specificity and gene expression of the amino acid permeases in Saccharomyces cerevisiae. Curr.Genet. 36, 317–328 (1999).
Rua D., Tobe B.T., Kron S.J.: Cell cycle control of yeast filamentous growth. Curr.Opin.Microbiol. 4, 720–727 (2001).
Schmidt A., Hall M.N., Koller A.: Two FK506 resistance-conferring genes in Saccharomyces cerevisiae, TAT1 and TAT2, encode amino acid permeases mediating tyrosine and tryptophan uptake. Mol.Cell.Biol. 14, 6597–6606 (1994).
Schreve J.L., Sin J.K., Garrett J.M.: The Saccharomyces cerevisiae YCC5 (YCL025c) gene encodes an amino acid permease, Agp1p, which transports asparagine and glutamine. J.Bacteriol. 180, 2556–2559 (1998).
Sipiczki M., Takeo K., Yamaguchi M., Yoshida S., Miklos I.: Environmentally controlled dimorphic cycle in a fission yeast. Microbiology 144, 1319–1330 (1998a).
Sipiczki M., Takeo K., Grallert A.: Growth polarity transitions in a dimorphic fission yeast. Microbiology 144, 3475–3485 (1998b).
Stark M.J.R.: Protein phosphorylation and dephosphorylation, pp. 284–375 in J.R. Dickinson, M. Schweizer (Eds): The Metabolism and Molecular Physiology of Saccharomyces cerevisiae, 2nd ed. CRC Press, Boca Raton 2004.
Vewaal R., Paalman J.W.G., Hogenkamp A., Verkleij A.J., Verrips C.T., Boonstra J.: HXT5 expression is determined by growth rates in Saccharomyces cerevisiae. Yeast 19, 1029–1038 (2002).
Warringer J., Blomberg A.: Involvement of yeast YOL151/GRE2 in ergosterol metabolism. Yeast 23, 389–398 (2006).
Winge D.R., Nielson K.B., Gray W.R., Hamer D.H., Karin M., Najarian R., Haslinger A., Valenzuela P., Welch J., Fogel S.: Yeast metallothionein. Sequence and metal-binding properties. J.Biol.Chem. 260, 14464–14470 (1985).
Wu X., Jiang Y.W.: Possible integration of upstream signals at Cdc42 in filamentous differentiation of S. cerevisiae. Yeast 22, 1069–1077 (2005).
Author information
Authors and Affiliations
Corresponding author
Additional information
Dedicated to the memory of Anna Kocková-Kratochvílová
Rights and permissions
About this article
Cite this article
Dickinson, J.R. Filament formation in Saccharomyces cerevisiae — a review. Folia Microbiol 53, 3–14 (2008). https://doi.org/10.1007/s12223-008-0001-6
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-008-0001-6