Abstract
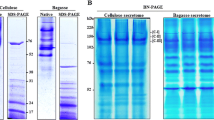
The fungus Trichoderma reesei is a major producer of plant cell wall-degrading enzymes, such as cellulases and hemicellulases. These enzymes have been used to hydrolyze agro-industrial wastes in processes of second-generation biofuel production. The disaccharide lactose, the main by-product of dairy industry, is considered the most attractive substrate for the commercial production of cellulases by T. reesei. In the present work, it was shown that T. reesei, cultivated in media supplemented with either lactose or galactose, secrete proteins which behave as multi-enzymatic complexes. Blue Native electrophoresis (BN-PAGE) displayed two major bands which were specific or present in higher amounts in each secretome sample, complex I in galactose medium and complex II in lactose medium. The protein components of complexes I and II were identified by means of LC-MS/MS. The complexes were composed of different glycosyl hydrolases and predicted proteins related to biopolymer metabolism. Zymography coupled to BN-PAGE showed that T. reesei secretome presents higher cellulolytic and xylanolytic activities in lactose than in galactose-induced medium. The results presented here corroborate the existence of hydrolysis based on multi-enzymatic complexes in T. reesei secretome, as observed previously in other fungi, such as Trichoderma harzianum and Penicillium purpurogenum.


Similar content being viewed by others
References
de Vries RP, Kester HC, Poulsen CH, Benen JA, Visser J (2000) Synergy between enzymes from Aspergillus involved in the degradation of plant cell wall polysaccharides. Carbohydr Res 327(4):401–410
Kuhad RC, Gupta R, Singh A (2011) Microbial cellulases and their industrial applications. Enzyme Res 2011:280696. doi:10.4061/2011/280696
Ivanova C, Bååth JA, Seiboth B, Kubicek CP (2013) Systems analysis of lactose metabolism in Trichoderma reesei identifies a lactose permease that is essential for cellulase induction. PLoS One 8(5), e62631. doi:10.1371/journal.pone.0062631
Xu J, Zhao G, Kou Y, Zhang W, Zhou Q, Chen G, Liu W (2014) Intracellular β-glucosidases CEL1a and CEL1b are essential for cellulase induction on lactose in Trichoderma reesei. Eukaryot Cell 13(8):1001–1013. doi:10.1128/EC.00100-14
Stricker A, Grosstessner-Hain K, Würleitner E, Mach R (2006) Xyr1 (xylanase regulator 1) regulates both the hydrolytic enzyme system and D-xylose metabolism in Hypocrea jecorina. Eukaryot Cell 5(12):2128–2137. doi:10.1128/EC.00211-06
Stricker AR, Steiger MG, Mach RL (2007) Xyr1 receives the lactose induction signal and regulates lactose metabolism in Hypocrea jecorina. FEBS Lett 581(21):3915–3920. doi:10.1016/j.febslet.2007.07.025
da Silva AJ, Gómez-Mendoza DP, Junqueira M, Domont GB, Ximenes Ferreira Filho E, de Sousa MV, Ricart CA (2012) Blue native-PAGE analysis of Trichoderma harzianum secretome reveals cellulases and hemicellulases working as multienzymatic complexes. Proteomics 12(17):2729–2738. doi:10.1002/pmic.201200048
Gonzalez-Vogel A, Eyzaguirre J, Oleas G, Callegari E, Navarrete M (2011) Proteomic analysis in non-denaturing condition of the secretome reveals the presence of multienzyme complexes in Penicillium purpurogenum. Appl Microbiol Biotechnol 89(1):145–155. doi:10.1007/s00253-010-2953-0
Chávez R, Bull P, Eyzaguirre J (2006) The xylanolytic enzyme system from the genus Penicillium. J Biotechnol 123(4):413–433. doi:10.1016/j.jbiotec.2005.12.036
Karaffa L, Fekete E, Gamauf C, Szentirmai A, Kubicek C, Seiboth B (2006) D-Galactose induces cellulase gene expression in Hypocrea jecorina at low growth rates. Microbiology 152(Pt 5):1507–1514. doi:10.1099/mic.0.28719-0
Schägger H, Cramer W, von Jagow G (1994) Analysis of molecular masses and oligomeric states of protein complexes by blue native electrophoresis and isolation of membrane protein complexes by two-dimensional native electrophoresis. Anal Biochem 217(2):220–230
Sunna A, Antranikian G (1997) Xylanolytic enzymes from fungi and bacteria. Crit Rev Biotechnol 17(1):39–67. doi:10.3109/07388559709146606
Matsui H, Ban-Tokuda T (2008) Studies on carboxymethyl cellulase and xylanase activities of anaerobic fungal isolate CR4 from the bovine rumen. Curr Microbiol 57(6):615–619. doi:10.1007/s00284-008-9252-3
Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18):3674–3676. doi:10.1093/bioinformatics/bti610
Gamauf C, Marchetti M, Kallio J, Puranen T, Vehmaanperä J, Allmaier G, Kubicek CP, Seiboth B (2007) Characterization of the bga1-encoded glycoside hydrolase family 35 beta-galactosidase of Hypocrea jecorina with galacto-beta-D-galactanase activity. FEBS J 274(7):1691–1700. doi:10.1111/j.1742-4658.2007.05714.x
Seiboth B, Hartl L, Salovuori N, Lanthaler K, Robson GD, Vehmaanperä J, Penttilä ME, Kubicek CP (2005) Role of the bga1-encoded extracellular {beta}-galactosidase of Hypocrea jecorina in cellulase induction by lactose. Appl Environ Microbiol 71(2):851–857. doi:10.1128/AEM.71.2.851-857.2005
Mouyna I, Sarfati J, Recco P, Fontaine T, Henrissatz B, Latge JP (2002) Molecular characterization of a cell wall-associated beta(1-3)endoglucanase of Aspergillus fumigatus. Med Mycol 40(5):455–464
Mouyna I, Morelle W, Vai M, Monod M, Léchenne B, Fontaine T, Beauvais A, Sarfati J, Prévost MC, Henry C, Latgé JP (2005) Deletion of GEL2 encoding for a beta(1-3)glucanosyltransferase affects morphogenesis and virulence in Aspergillus fumigatus. Mol Microbiol 56(6):1675–1688. doi:10.1111/j.1365-2958.2005.04654.x
Gastebois A, Mouyna I, Simenel C, Clavaud C, Coddeville B, Delepierre M, Latgé JP, Fontaine T (2010) Characterization of a new beta(1-3)-glucan branching activity of Aspergillus fumigatus. J Biol Chem 285(4):2386–2396. doi:10.1074/jbc.M109.077545
Hartl L, Gastebois A, Aimanianda V, Latgé JP (2011) Characterization of the GPI-anchored endo β-1,3-glucanase Eng2 of Aspergillus fumigatus. Fungal Genet Biol 48(2):185–191. doi:10.1016/j.fgb.2010.06.011
Chelikani P, Fita I, Loewen PC (2004) Diversity of structures and properties among catalases. Cell Mol Life Sci 61(2):192–208. doi:10.1007/s00018-003-3206-5
Zámocký M, Droghetti E, Bellei M, Gasselhuber B, Pabst M, Furtmüller PG, Battistuzzi G, Smulevich G, Obinger C (2012) Eukaryotic extracellular catalase-peroxidase from Magnaporthe grisea—biophysical/chemical characterization of the first representative from a novel phytopathogenic KatG group. Biochimie 94(3):673–683. doi:10.1016/j.biochi.2011.09.020
Tanabe S, Nishizawa Y, Minami E (2009) Effects of catalase on the accumulation of H(2)O(2) in rice cells inoculated with rice blast fungus, Magnaporthe oryzae. Physiol Plant 137(2):148–154. doi:10.1111/j.1399-3054.2009.01272.x
Tanabe S, Ishii-Minami N, Saitoh K, Otake Y, Kaku H, Shibuya N, Nishizawa Y, Minami E (2011) The role of catalase-peroxidase secreted by Magnaporthe oryzae during early infection of rice cells. Mol Plant Microbe Interact 24(2):163–171. doi:10.1094/MPMI-07-10-0175
Wang ZJ, Xu XP, Fan KQ, Jia CJ, Yang KQ (2007) Sample preparation for two-dimensional blue native/SDS polyacrylamide gel electrophoresis in the identification of Streptomyces coelicolor cytoplasmic protein complexes. J Biochem Biophys Methods 70(4):565–572. doi:10.1016/j.jbbm.2007.01.001
Heuberger EH, Veenhoff LM, Duurkens RH, Friesen RH, Poolman B (2002) Oligomeric state of membrane transport proteins analyzed with blue native electrophoresis and analytical ultracentrifugation. J Mol Biol 317(4):591–600. doi:10.1006/jmbi.2002.5416
Wittig I, Schägger H (2008) Features and applications of blue-native and clear-native electrophoresis. Proteomics 8(19):3974–3990. doi:10.1002/pmic.200800017
Geiger A, Hirtz C, Bécue T, Bellard E, Centeno D, Gargani D, Rossignol M, Cuny G, Peltier JB (2010) Exocytosis and protein secretion in Trypanosoma. BMC Microbiol 10:20. doi:10.1186/1471-2180-10-20
Acknowledgments
This project was supported by the FAPDF (Federal District Foundation for Science and Research Supporting—grant nos. 193.000.584/2009 and 193.000.470/2011). FINEP (Design and Project Financing Agency), Petrobras (Brazilian Oil Corporation, CENPES 11007), CNPq (National Council for Scientific and Technological Development grant nos. 478066/2010-4 MCT/CNPq–Universal, 470265/2011-6, 563260/2010-6, and 563823/2010-0), University of Brasilia (UnB), Universidade Federal do Rio de Janeiro (UFRJ), and the National Institute of Science and Technology of Bioethanol. CNPq provided scholarship for DPGM.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 214 kb)
Rights and permissions
About this article
Cite this article
da Silva, A.J., Gómez-Mendoza, D.P., Junqueira, M. et al. Secretomic Analysis Reveals Multi-Enzymatic Complexes in Trichoderma reesei Grown in Media Containing Lactose or Galactose. Bioenerg. Res. 8, 1906–1911 (2015). https://doi.org/10.1007/s12155-015-9648-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12155-015-9648-4