Abstract
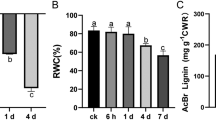
Lignocellulosic biomass has the potential to be a significant source of renewable energy, provided that current challenges regarding sufficient biomass production and efficient biomass conversion are met. Floral transition is a major developmental switch in the life of flowering plants which dictates whether photosynthetic products should be invested in vegetative growth or reproductive development. To directly evaluate the contribution of floral transition to biomass yield, we manipulated the onset of flowering in the model legume Medicago truncatula using a mutagenesis approach. Through forward genetics screening of Tnt1-tagged mutants, we identified three M. truncatula lines altered in flowering time and fertility: a delayed flowering mutant named vernalization-insensitive delayed flowering in long days (vdf), a nonflowering stemless mutant named headless (hdl), and a male sterile mutant named medicago male sterile 1 (mms1). Biomass yield analyses at different stages of development in these mutants revealed that the vdf mutant had the highest aboveground biomass and the hdl mutant had the lowest biomass at 70 days after germination. The difference in biomass yield between the vdf mutant and wild-type R108 became apparent after floral initiation and peaked at 90 days after germination, the late blooming stage for R108, where the vdf plants produced approximately twofold more biomass than R108. Interestingly, vdf, hdl, and mms1 mutants produced significantly less lignin than R108. Our results suggest that delaying floral initiation could be employed as a convenient tool to simultaneously improve biomass quantity and quality, provided that this is achieved without pleiotropic developmental defects.





Similar content being viewed by others
References
Somerville C (2007) Biofuels. Curr Biol 17(4):R115–R119
Stamm P, Verma V, Ramamoorthy R, Kumar PP (2012) Manipulation of plant architecture to enhance lignocellulosic biomass. AoB PLANTS pls026
Carroll A, Somerville, C (2009) Cellulosic biofuels. Annu Rev Plant Biol 60:165–182
Perlack RD, Wright LL, Turhollow AF, Graham RL, Stokes BJ, Erbach DC (2005) Biomass as feedstock for a bioenergy and bioproducts industry: the technical feasibility of a billion-ton annual supply. Oakridge National Laboratory, p. 75
Demura T, Ye Z-H (2010) Regulation of plant biomass production. Curr Opin Plant Biol 13:299–304
Chen F, Dixon RA (2007) Lignin modification improves fermentable sugar yields for biofuel production. Nat Biotechnol 25(7):759–761
Pauly M, Keegstra K (2010) Plant cell wall polymers as precursors for biofuels. Curr Opin Plant Biol 13:305–312
Fu C, Mielenz J, Xiao X, Ge Y, Hamilton CY, Rodriguez M, Chen F, Foston M, Ragauskas A, Bouton J, Dixon RA, Wang ZY (2011) Genetic manipulation of lignin reduces recalcitrance and improves ethanol production from switchgrass. Proc Natl Acad Sci U S A 108:3803–3808
Chundawat SPS, Venkatesh B, Dale BE (2007) Effect of particle size based separation of milled corn stover on AFEX pretreatment and enzymatic digestibility. Biotechnol Bioeng 96:219–231
Weng J-K, Akiyama T, Ralph J, Chapple C (2011) Independent recruitment of an O-methyltransferase for syringyl lignin biosynthesis in Selaginella moellendorffii. Plant Cell 23(7):2708–2724
Vanholme R, Ralph J, Akiyama T, Lu F, Pazo JR, Kim H, Christensen JH, Van Reusel B, Storme V, De Rycke R, Rohde A, Morreel K, Boerjan W (2010) Engineering traditional monolignols out of lignin by concomitant up-regulation of F5H1 and down-regulation of COMT in Arabidopsis. Plant J 64(6):885–897
Vanholme R, Demedts B, Morreel K, Ralph J, Boerjan W (2010) Lignin biosynthesis and structure. Plant Physiol 153(3):895–905
Boerjan W, Ralph J, Baucher M (2003) Lignin biosynthesis. Annu Rev Plant Biol 54:519–546
Tanaka K, Murata K, Yamazaki M, Onosato K, Miyao A, Hirochika H (2003) Three distinct rice cellulose synthase catalytic subunit genes required for cellulose synthesis in the secondary wall. Plant Physiol 133(1):73–83
Liepman AH, Wilkerson, Curtis G, Keegstra K (2005) Expression of cellulose synthase-like (Csl) genes in insect cells reveals that CslA family members encode mannan synthases. Proc Natl Acad Sci U S A 102(6):2221–2226
Chen F, Srinivasa R, Marry S, Temple S, Jackson L, Shadle G, Dixon RA (2006) Multi-site genetic modulation of monolignol biosynthesis suggests new routes for formation of syringyl lignin and wall-bound ferulic acid in alfalfa (Medicago sativa L.). Plant J 48(1):113–124
Somerville C (2006) Cellulose synthesis in higher plants. Annu Rev Cell Dev Biol 22:53–78
Petersen PD, Lau J, Ebert B, Yang F, Verhertbruggen Y, Kim JS, Varanasi P, Suttangkakul A, Auer M, Loque D, Scheller HV (2012) Engineering of plants with improved properties as biofuels feedstocks by vessel-specific complementation of xylan biosynthesis mutants. Biotechnol Biofuels 5(1):84
Holland N, Holland D, Helentjaris T, Dhugga KS, Xoconostle-Cazares B, Delmer DP (2000) A comparative analysis of the plant cellulose synthase (CesA) gene family. Plant Physiol 123(4):1313–1324
Yang C, Xu Z, Song J, Conner K, Vizcay Barrena G, Wilson ZA (2007) Arabidopsis MYB26/MALE STERILE35 regulates secondary thickening in the endothecium and is essential for anther dehiscence. Plant Cell 19(2):534–548
Mitsuda N, Seki M, Shinozaki K, Ohme-Takagi M (2005) The NAC transcription factors NST1 and NST2 of Arabidopsis regulate secondary wall thickenings and are required for anther dehiscence. Plant Cell 17(11):2993–3006
Yamaguchi M, Mitsuda N, Ohtani M, Ohme-Takagi M, Kato K, Demura T (2011) Vascular-related NAC-domain 7 directly regulates the expression of a broad range of genes for xylem vessel formation. Plant J 66(4):579–590
Mitsuda N, Ohme-Takagi M, Iwase A, Yamamoto H, Yoshida M, Seki M, Shinozaki K, Ohme-Takagi M (2007) NAC transcription factors, NST1 and NST3, are key regulators of the formation of secondary walls in woody tissues of Arabidopsis. Plant Cell 19(1):270–280
Zhong R, Ye Z-H (2012) MYB46 and MYB83 bind to the SMRE sites and directly activate a suite of transcription factors and secondary wall biosynthetic genes. Plant Cell Physiol 53(2):368–380
Zhong R, Lee C, Zhou J, McCarthy RL, Ye Z-H, Richardson EA, Ye Z-H (2007) Two NAC domain transcription factors, SND1 and NST1, function redundantly in regulation of secondary wall synthesis in fibers of Arabidopsis. Planta 225(6):1603–1611
Mitsuda N, Ohme-Takagi M (2008) NAC transcription factors NST1 and NST3 regulate pod shattering in a partially redundant manner by promoting secondary wall formation after the establishment of tissue identity. Plant J 56(5):768–778
Zhong R, Lee C, Zhou J, McCarthy RL, Ye Z-H (2008) A battery of transcription factors involved in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell 20(10):2763–2782
Zhou J, Lee C, Zhong R, Ye Z-H (2009) MYB58 and MYB63 are transcriptional activators of the lignin biosynthetic pathway during secondary cell wall formation in Arabidopsis. Plant Cell 21(1):248–266
Wang H, Avci U, Nakashima J, Hahn MG, Chen F, Dixon RA (2010) Mutation of WRKY transcription factors initiates pith secondary wall formation and increases stem biomass in dicotyledonous plants. Proc Natl Acad Sci U S A 107(51):22338–22343
Mele G, Ori N, Sato Y, Hake S (2003) The knotted1-like homeobox gene BREVIPEDICELLUS regulates cell differentiation by modulating metabolic pathways. Genes Dev 17(17):2088–2093
Tadege M, Lin H, Bedair M, Berbel A, Wen J, Rojas CM, Niu L, Tang Y, Sumner L, Ratet P, McHale NA, Madueño F, Mysore KS (2011) STENOFOLIA regulates blade outgrowth and leaf vascular patterning in Medicago truncatula and Nicotiana sylvestris. Plant Cell 23(6):2125–2142
Vandenbussche M, Horstman A, Zethof J, Koes R, Rijpkema AS, Gerats T (2009) Differential recruitment of WOX transcription factors for lateral development and organ fusion in Petunia and Arabidopsis. Plant Cell 21(8):2269–2283
Nardmann J, Ji J, Werr W, Scanlon MJ (2004) The maize duplicate genes narrow sheath1 and narrow sheath2 encode a conserved homeobox gene function in a lateral domain of shoot apical meristems. Development 131(12):2827–2839
Ishiwata A, Ozawa M, Nagasaki H, Kato M, Noda Y, Yamaguchi T, Nosaka M, Shimizu-Sato S, Nagasaki A, Maekawa M, Hirano HY, Sato Y (2013) Two WUSCHEL-related homeobox genes, narrow leaf2 and narrow leaf3, control leaf width in rice. Plant Cell Physiol 54(5):779–792
Cho SH, Yoo SC, Zhang H, Pandeya D, Koh HJ, Hwang JY, Kim GT, Paek NC (2013) The rice narrow leaf2 and narrow leaf3 loci encode WUSCHEL-related homeobox 3A (OsWOX3A) and function in leaf, spikelet, tiller and lateral root development. New Phytol 198(4):1071–1084
Sheldon CC, Burn JE, Perez PP, Metzger J, Edwards JA, Peacock WJ, Dennis ES (1999) The FLF MADS box gene: a repressor of flowering in Arabidopsis regulated by vernalization and methylation. Plant Cell 11(3):445–458
Michaels SD, Amasino RM (1999) FLOWERING LOCUS C encodes a novel MADS domain protein that acts as a repressor of flowering. Plant Cell 11(5):949–956
Yoshida N, Yanai Y, Chen L, Kato Y, Hiratsuka J, Miwa T, Sung ZR, Takahashi S (2001) EMBRYONIC FLOWER2, a novel polycomb group protein homolog, mediates shoot development and flowering in Arabidopsis. Plant Cell 13:2471–2481
Sung ZR, Belachew A, Shunong B, Bertrand-Garcia R (1992) EMF, an Arabidopsis gene required for vegetative shoot development. Science 258:1645–1647
Aubert D, Chen L, Moon Y-H, Martin D, Castle LA, Yang C-H, Sung ZR (2001) EMF1, a novel protein involved in the control of shoot architecture and flowering in Arabidopsis. Plant Cell 13:1865–1875
Kardailsky I, Shukla VK, Ahn JH, Dagenais N, Christensen SK, Nguyen JT, Chory J, Harrison MJ, Weigel D (1999) Activation tagging of the floral inducer FT. Science 286:1962–1965
Gu Q, Ferrándiz C, Yanofsky MF, Martienssen R (1998) The FRUITFULL MADS-box gene mediates cell differentiation during Arabidopsis fruit development. Development 125(8):1509–1517
Ferrándiz C, Gu Q, Martienssen R, Yanofsky MF (2000) Redundant regulation of meristem identity and plant architecture by FRUITFULL, APETALA1 and CAULIFLOWER. Development 127(4):725–734
Samach A, Onouchi H, Gold SE, Ditta GS, Schwarz-Sommer Z, Yanofsky MF, Coupland G (2000) Distinct roles of CONSTANS target genes in reproductive development of Arabidopsis. Science 288(5471):1613–1616
Melzer S, Lens F, Gennen J, Vanneste S, Rohde A, Beeckman T (2008) Flowering-time genes modulate meristem determinacy and growth form in Arabidopsis thaliana. Nat Genet 40(12):1489–1492
Jensen E et al (2013) Flowering induction in the bioenergy grass Miscanthus sacchariflorus is a quantitative short-day response, whilst delayed flowering under long days increases biomass accumulation. J Exp Bot 64(2):541–552
Jensen E et al (2011) Characterization of flowering time diversity in Miscanthus species. Global Chang Biol Bioenergy 3(5):387–400
Alexopoulou E et al (2008) Biomass yields for upland and lowland switchgrass varieties grown in the Mediterranean region. Biomass Bioenergy 32(10):926–933
Casler MD et al (2007) Genetic diversity, plant adaptation regions, and gene pools for switchgrass. Crop Sci 47(6):2261–2273
Stroup JA et al (2003) Comparison of growth and performance in upland and lowland switchgrass types to water and nitrogen stress. Bioresour Technol 86(1):65–72
Mullet J et al (2014) Energy sorghum—a genetic model for the design of C-4 grass bioenergy crops. J Exp Bot 65(13):3479–3489
Olson SN et al (2012) High biomass yield energy sorghum: developing a genetic model for C4 grass bioenergy crops. Biofuels Bioprod Bioref-Biofpr 6(6):640–655
Rooney WL et al (2007) Designing sorghum as a dedicated bioenergy feedstock. Biofuels Bioprod Bioref-Biofpr 1(2):147–157
Yang SS et al. (2014) Sorghum phytochrome B inhibits flowering in long days by activating expression of SbPRR37 and SbGHD7, repressors of SbEHD1, SbCN8 and SbCN12. PLoS One 9(8)
Murphy RL et al. (2014) Ghd7 (Ma(6)) represses sorghum flowering in long days: Ghd7 alleles enhance biomass accumulation and grain production. Plant Genome 7(2)
Murphy RL et al (2011) Coincident light and clock regulation of pseudoresponse regulator protein 37 (PRR37) controls photoperiodic flowering in sorghum. Proc Natl Acad Sci U S A 108(39):16469–16474
Yarce JC, Lee HK, Tadege M, Ratet P, Mysore KS (2013) Forward genetics screening of Medicago truncatula Tnt1 insertion lines. Methods Mol Biol 1069:93–100
Tadege M, Wang TL, Wen J, Ratet P, Mysore KS (2009) Mutagenesis and Beyond! Tools for understanding legume biology. Plant Physiol 151(3):978–984
Tadege M, Wen J, He J, Tu H, Kwak Y, Eschstruth A, Cayrel A, Endre G, Zhao PX, Chabaud M, Ratet P, Mysore KS (2008) Large-scale insertional mutagenesis using the Tnt1 retrotransposon in the model legume Medicago truncatula. Plant J 54(2):335–347
Tadege M, Ratet P, Mysore KS (2005) Insertional mutagenesis: a Swiss Army knife for functional genomics of Medicago truncatula. Trends Plant Sci 10(5):229–235
Vogel KP, Pedersen JF, Materson SD, Toy JJ (1999) Evaluation of a filter bag system for NDF, ADF, and IVDMD forage analysis. Crop Sci 39:276–279
Casler MD, Vogel KP (2014) Selection for biomass yield in upland, lowland, and hybrid switchgrass. Crop Sci 54(2):626–636
Arnoult S, Quillet MC, Brancourt-Hulmel M (2014) Miscanthus clones display large variation in floral biology and different environmental sensitivities useful for breeding. Bioenergy Res 7(1):430–441
Casler MD et al (2007) Latitudinal and longitudinal adaptation of switchgrass populations. Crop Sci 47(6):2249–2260
Vermerris W, Thompson KJ, McIntyre LM (2002) The maize Brown midrib1 locus affects cell wall composition and plant development in a dose-dependent manner. Heredity 88(6):450–457
Xu B, Sathitsuksanoh N, Tang Y, Udvardi MK, Zhang JY, Shen Z, Balota M, Harich K, Zhang PY, Zhao B (2012) Overexpression of AtLOV1 in switchgrass alters plant architecture, lignin content, and flowering time. PLoS One 7(12):e47399
Yoo SY, Kim Y, Kim SY, Lee JS, Ahn JH (2007) Control of flowering time and cold response by a NAC-domain protein in Arabidopsis. PLoS One 2(7):e642
Shi Y, Xu Z, Xu ZY, Li L, Zhang C, Schläppi M, Xu ZQ (2011) Influence of EARLI1-like genes on flowering time and lignin synthesis of Arabidopsis thaliana. Plant Biol 13(5):731–739
Wang YW, Wang WC, Jin SH, Wang J, Wang B, Hou BK (2012) Over-expression of a putative poplar glycosyltransferase gene, PtGT1, in tobacco increases lignin content and causes early flowering. J Exp Bot 63(7):2799–2808
Chuck GS, Tobias C, Sun L, Kraemer F, Li C, Dibble D, Arora R, Bragg JN, Vogel JP, Singh S, Simmons BA, Pauly M, Hake S (2011) Overexpression of the maize Corngrass1 microRNA prevents flowering, improves digestibility, and increases starch content of switchgrass. Proc Natl Acad Sci U S A 108(42):17550–17555
Fu C, Sunkar R, Zhou C, Shen H, Zhang JY, Matts J, Wolf J, Mann DG, Stewart CN Jr, Tang Y, Wang ZY (2012) Overexpression of miR156 in switchgrass (Panicum virgatum L.) results in various morphological alterations and leads to improved biomass production. Plant Biotechnol J 10(4):443–452
Fukushima RS, Hatfield RD (2004) Comparison of the acetyl bromide spectrophotometric method with other analytical lignin methods for determining lignin concentration in forage samples. J Agri Food Chem 52:3713–3720
Zhao Q, Gallego-Giraldo L, Wang H, Zeng Y, Ding S-Y, Chen F, Dixon RA (2010) An NAC transcription factor orchestrates multiple features of cell wall development in Medicago truncatula. Plant J 63(1):100–114
Acknowledgments
This work was supported by the Samuel Roberts Noble Foundation and the National Science Foundation (EPSCoR Grant EPS-0814361). The insertion mutant lines of M. truncatula were created through research support, in part, from the National Science Foundation (DBI-0703285).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Tadege, M., Chen, F., Murray, J. et al. Control of Vegetative to Reproductive Phase Transition Improves Biomass Yield and Simultaneously Reduces Lignin Content in Medicago truncatula . Bioenerg. Res. 8, 857–867 (2015). https://doi.org/10.1007/s12155-014-9565-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12155-014-9565-y