Abstract
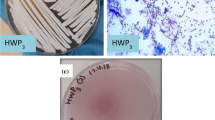
Ligninolytic enzyme complexes are involved in lignin degradation. Among them laccases are outstanding because they use molecular oxygen as a co-substrate instead of hydrogen peroxide as used by peroxidases. Bacterial laccase of Bacillus genus was first reported in Claus and Filip (Microbiol Res 152:209–216, 1997), since then more bacterial laccases have been found. In this research, laccase-producing bacteria were screened from pulp and paper industry wastewater, bagass and sugarcane rhizosphere. Nutrient agar medium containing 0.5 mM of guaiacol was used. It was observed that the laccase-producing strains developed brown colour from which 16 strains of Bacillus were identified. One of the isolated strains was identified as Bacillus subtilis WPI based on the results of biochemical tests and 16S rDNA sequence analysis. This strain showed laccase-like activity towards the oxidizing substrates ABTS and guaiacol. In this study guaiacol was used as the substrate of laccase activity assay. For determination of laccase activity of this isolate guaiacol was used as a substrate of assay for the first time in this study. SDS-PAGE and Native-PAGE confirmed the presence of laccase.





Similar content being viewed by others
References
Claus H, Filip Z (1997) The evidence of a laccase-like enzyme activity in a Bacillus sphaericus strain. Microbiol Res 152:209–216
Thurston CF (1994) The structure and function of fungal laccases. Microbiology 140:19–26
Desai SS, Nityanand C (2011) Microbial laccases and their application. Asian J Biotechnol 3:98–124
Pazarlıoglu NK, Sariisik M, Telefoncu A (2005) Laccase: production by Trametes versicolor and application to denim washing. Process Biochem 40(1):673–1678
Xu F (2005) Applications of oxidoreductases: recent progress. Ind Biotechol 1:38–50
Gardiol AE, Hernandez RJ, Harte BR (1998) Device for detecting oxygen with oxidase, US Pat. 5, 804, 401
Faure D, Bouillant ML, Bally R (1994) Isolation of Azospirillum lipoferum 4T mutants affected in melanization and laccase activity. Appl Environ Microbiol 60:3413–3415
Endo K, Hosono K, Beppu T, Ueda K (2002) A novel extra cytoplasmatic phenol oxidase of Streptomyces: its possible involvement in the onset of morphogenesis. Microbiology 148:1767–1776
Hullo MF, Moszer I, Danchin A, Martin-Verstraete I (2001) CotA of Bacillus subtilisis a copper-dependent laccase. J Bacteriol 183:5426–5430
Augustine AJ, Kragh ME, Sarangi R, Fujii S, Liboiron BD, Stoj CS, Kosman DJ, Hodgson KO, Hedman B, Solomon EI (2008) Spectroscopic studies of perturbed T1 Cu sites in the multicopper oxidases Saccharmycescerevisiae Fet3p and Rhusvernicifera laccase: allosteric coupling between the T1 and trinuclear Cu sites. Biochemistry 47:2036–2045
Rodgers CJ, Blanford CF, Giddens SR, Skamnioti P, Armstrong FA, Gurr SJ (2010) Designer laccases: a vogue for high-potential fungal enzymes? Trends Biotechnol 28:63–72
Koschorreck K, Schmid RD, Urlacher VB (2009) Improving the functional expression of a Bacillus licheniformis laccase by random and site-directed mutagenesis. BMC Biotechnol 9:12–21
BadoeiDalfard A, Khajeh K, Soudi MR, Naderi-Manesh H, Ranjbar B, Hassan Sajed R (2006) Isolation and biochemical characterization of laccase and tyrosinase activities in a novel melanogenic soil bacterium. Enzyme Microb Technol 39:1409–1416
Bains J, Capalash N, Sharma P (2003) Laccase from a non-melanogenic, alkalotolerantγ-proteobacterium JB isolated from industrial wastewater drained soil. Biotechnol Lett 25(1):155–1159
Machado KMG, Matheus DR (2006) “Potential of a ligninolytic enzymatic complex produced by Pleurotusostreatus during growth on solid substrate for the biodegradation of organic pollutants”Brazilian. J Microbiol 37:468–473
Niku-Paavola ML, Raaska L, Itvaara M (1990) Detection of white-rot fungi by a nontoxic stain. Mycol Res 94:27–31
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacterio phage T4. Nature 227:680–685
Papizadeh M, Roayaei Ardakani M, Ebrahimipour Gh, Motamedi H (2010) Utilization of dibenzothiophene as sulfur source by Microbacterium sp. NISOC-06. World J Microbiol Biotechnol 26:1195–1200
Duarte GF, Rosado AS, Seldin L, De Araujo W, Van Elsas JD (2001) Analysis of bacterial community structure in sulfurous-oil containing soils and detection of species carrying dibenzothiophene desulfurization (dsz) genes. Appl Environ Microbiol 67:1052–1062
Liers C, Ullrich R, Pecyna M, Schlosser D, Hofrichter M (2007) Production, purification and partial enzymatic and molecular characterization of a laccase from the wood-rotting ascomycete Xylariapolymorpha. Enzyme Microbial Technol 41:785–793
Li A, Zhu Y, Xu L, Zhu W, Tian X (2008) Comparative study on the determination of assay for laccase of Trametes sp. Af J Biochem Res 2:181–183
Robles A, Lucas R, Cienfuegos GA, Galvez A (2000) Phenol-oxidase (laccase) activity in strains of the hyphomycete Chalaraparadoxa isolated from olive mill wastewater disposal ponds. Enzyme Microb Technol 26:484–490
Ruijssenaars HJ, Hartmans S (2004) A cloned Bacillus halodurans multicopper oxidase exhibiting alkaline laccase activity. Appl Microbiol Biotechnol 65:177–182
Niladevi KN, Jacob N, Prema P (2008) Evidence for a halotolerant-alkaline laccase in Streptomyces psammoticus Purification and characterization. Process Biochem 43:654–660
Mohammadian M, Fathi-Roudsari M, Mollania N, Badoei-Dalfard A, Khajeh K (2010) Enhanced expression of a recombinant bacterial laccase at low temperature and microaerobic conditions: purification and biochemical characterization. J Ind Microbiol Biotechnol 37:863–869
Reiss R, Ihssen J, Thony-Meyer L (2011) Bacillus pumilus laccase: a heat stable enzyme with a wide substrate spectrum. BMC Biotechnol 11:1–11
Acknowledgments
This research was supported by Biotechnology and Biological Research Center of Shahid Chamran University of Ahvaz (Iran).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sheikhi, F., Roayaei Ardakani, M., Enayatizamir, N. et al. The Determination of Assay for Laccase of Bacillus subtilis WPI with Two Classes of Chemical Compounds as Substrates. Indian J Microbiol 52, 701–707 (2012). https://doi.org/10.1007/s12088-012-0298-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12088-012-0298-3