Abstract
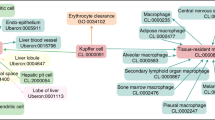
The biological cell, a natural self-contained unit of prime biological importance, is an enormously complex machine that can be understood at many levels. A higher-level perspective of the entire cell requires integration of various features into coherent, biologically meaningful descriptions. There are some efforts to model cells based on their genome, proteome or metabolome descriptions. However, there are no established methods as yet to describe cell morphologies, capture similarities and differences between different cells or between healthy and disease states. Here we report a framework to model various aspects of a cell and integrate knowledge encoded at different levels of abstraction, with cell morphologies at one end to atomic structures at the other. The different issues that have been addressed are ontologies, feature description and model building. The framework describes dotted representations and tree data structures to integrate diverse pieces of data and parametric models enabling size, shape and location descriptions. The framework serves as a first step in integrating different levels of data available for a biological cell and has the potential to lead to development of computational models in our pursuit to model cell structure and function, from which several applications can flow out.
Similar content being viewed by others
Abbreviations
- DAG:
-
Directed acyclic graph
- RER:
-
rough endoplasmic reticulum
- VRML:
-
virtual reality modelling language
References
Aleshin A E, Zeng C, Bourenkov G P, Bartunik H D, Fromm H J and Honzatko R B 1998 The mechanism of regulation of hexokinase: new insights from the crystal structure of recombinant human brain hexokinase complexed with glucose and glucose-6-phosphate; Structure 6 39–50
Ashburner M, Ball C A, Blake J A, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K et al 2000 Gene ontology: tool for the unification of biology. The Gene Ontology Consortium; Nat. Genet. 25 25–29 ( http://www.geneontology.org/)
Bard J, Rhee S Y and Ashburner M 2005 An Ontology for cell types; Genome Biol. 6 R21
Beuschlein F, Fassnacht M, Klink A, Allolio B and Reincke M. 2001 ACTH-receptor expression, regulation and role in adrenocortical tumor formation; Eur. J. Endocrinol. 144 199–206
Bourdeau I and Stratakis C A 2002 Cyclic AMP-dependent signaling aberrations in macronodular adrenal disease; Ann. N.Y. Acad. Sci. 968 240–255
Clark A J L, Cammas F M, Watt A, Kapas S and Weber A 1997 Familial glucocorticoid deficiency, one syndrome but more than one gene; J. Mol. Med. 75 394–399
Cormen T H, Leiserson C E, Rivest R L and Stein C 2001 Introduction to Algorithms Second edition (MIT Press)
Hunter P J and Borg T K 2003 Integration from protein to organs: The Physiome Project; Nat. Rev. Mol. Cell Biol. 4 237–243 ( http://www.physiome.org.nz/ )
Hunter P J 2004 The IUPS Physiome Project: a framework for computational physiology; Prog. Biophys. Mol. Biol. 85 551–569
Ideker T, Galitski T and Hood L 2001 A new approach to decoding life: systems biology; Annu. Rev. Genomics Hum. Genet. 2 343–372
Kiehl T R, Mattheyses R M and Simmons M K 2004 Hybrid simulation of cellular behavior; Bioinformatics, 20 316–322
Kremer J R, Mastronarde D N and McIntosh J R 1996 Computer visualization of three-dimensional image data using IMOD; J. Struct. Biol. 116 71–76
Lin C J, Jorge A A L, Latronico A C, Marui S, Fragoso M C V, Martin R M, Carvalho F M, Arnhold I J P, Mendonca B B 2000 Origin of ovarian steroid cell tumor causing isosexual pseudoprecocious puberty demonstrated by the expression of adrenal steroidogenic enzymes and adrenocorticotrophin receptor; J. Clin. Endocrinol. Metab. 85 1211–1214
Lloyd C M, Halstead M D and Nielsen P F 2004 CellML: its future, present and past; Prog. Biophys. Mol. Biol. 85 433–450 ( http://www.cellml.org/ )
Loew L M 2002 The virtual cell project.; Novartis Found. Symp. 247 151–160 (Virtual Cell. http://www.nrcam.uchc.edu )
Mesiano S, Fujimoto V Y, Nelson L R, Lee J Y, Voytek C C and Jaffe R B 1996 Localization and regulation of corticotrophin receptor expression in the midgestation human fetal adrenal cortex: implications for in utero homeostasis; J. Clin. Endocrinol. Metab. 81 340–345
Mountjoy K G, Robbins L S, Mortrud M T and Cone R D 1992 The cloning of a family of genes that encode the melanocortin receptors; Science 257 1248–1251
Rosse C and Mejino J L Jr 2003 A reference ontology for biomedical informatics: the Foundational Model of Anatomy; J. Biomed. Informat. 36 478–500 ( http://sig.biostr.washington.edu/projects/fm/AboutFM.html )
Selvais P, Donckier J, Buysscheart M and Maiter D 1998 Cushing’s disease: a comparison of pituitary corticotroph microadenomas and macroadenomas; Eur. J. Endocrinol. 138 153–159
Solminski A, Erma G and Mihm M 2004 ACTH receptor, CYP11A1, CYP17 and CYP21A2 genes are expressed in skin; J. Clin. Endocrinol. Metab. 81 2746–2749
Thomaseth K 2003 Multidisciplinary modelling of biomedical systems; Comput. Methods Prog. Biomed. 71 189–201
Tomita M, Hashimoto K, Takahashi K, Shimizu T S, Matsuzaki Y, Miyoshi F, Saito K, Tanida S, Yugi K, Venter J C and Hutchison C A 3rd 1999 E-CELL: software environment for whole-cell simulation; Bioinformatics, 15 72–84. ( http://www.e-cell.org/ ).
Zwermann O, Beuschlein F, Klink A, Stahl M and Reincke M 2004 The role of the ACTH receptor in adrenal tumors: Identification of a novel microsatellite marker; Horm. Metab. Res. 36 406–410
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Khodade, P., Malhotra, S., Kumar, N. et al. Cytoview: Development of a cell modelling framework. J Biosci 32 (Suppl 1), 965–977 (2007). https://doi.org/10.1007/s12038-007-0096-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12038-007-0096-y