Abstract
Epilepsy is a severe neurological condition affecting 50–65 million individuals worldwide that can lead to brain damage. Nevertheless, the etiology of epilepsy remains poorly understood. Meta-analyses of genome-wide association studies involving 15,212 epilepsy cases and 29,677 controls of the ILAE Consortium cohort were used to conduct transcriptome-wide association studies (TWAS) and protein-wide association studies (PWAS). Furthermore, a protein-protein interaction (PPI) network was generated using the STRING database, and significant epilepsy-susceptible genes were verified using chip data. Chemical-related gene set enrichment analysis (CGSEA) was performed to determine novel drug targets for epilepsy. TWAS analysis identified 21,170 genes, of which 58 were significant (TWASfdr < 0.05) in ten brain regions, and 16 differentially expressed genes were verified based on mRNA expression profiles. The PWAS identified 2249 genes, of which 2 were significant (PWASfdr < 0.05). Through chemical-gene set enrichment analysis, 287 environmental chemicals associated with epilepsy were identified. We identified five significant genes (WIPF1, IQSEC1, JAM2, ICAM3, and ZNF143) that had causal relationships with epilepsy. CGSEA identified 159 chemicals that were significantly correlated with epilepsy (Pcgsea < 0.05), such as pentobarbital, ketone bodies, and polychlorinated biphenyl. In summary, we performed TWAS, PWAS (for genetic factors), and CGSEA (for environmental factors) analyses and identified several epilepsy-associated genes and chemicals. The results of this study will contribute to our understanding of genetic and environmental factors for epilepsy and may predict novel drug targets.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Epilepsy is a severe neurological condition characterized by repeated seizures that can lead to brain damage, disability, and even death [1]. Seizures are abnormal, excessive, or synchronous increases in neuronal activity, which can lead to behavioral manifestations [2]. Studies have shown that the lifetime prevalence of epilepsy is 7.60%, affecting 50–65 million individuals worldwide [3]. The etiology of epilepsy is unclear [4] and includes genetic, structural, infectious, metabolic, and immune factors. Moreover, the definitive cause cannot be determined in > 60% of patients [5]. Furthermore, current studies lack reliable human data from brain tissue or biomarker studies of common epilepsies [6] due to samples of human brain tissue being difficult to obtain [7]. Therefore, the etiology and pathogenesis of epilepsy must be further explored.
Genetic factors are one of the most important etiologies of epilepsy [8]. Genome-wide association studies (GWAS) for common epilepsy have identified genetic risk variants for generalized and focal epilepsy and febrile seizures [9]. Several candidate genes for epilepsy include calmodulin-regulated spectrin-associated protein 1-like protein 1 (CAMSAP1L1) [10], glutamate metabotropic receptor 3 (GRM3) [11], CD3 gamma subunit of T-cell receptor complex (CD3G), and solute carrier organic anion transporter family member 3A1 (SLCO3A1) [12]. However, it is difficult to identify genes that influence traits through GWAS alone [13]. Transcriptome-wide association study (TWAS) is a gene-based association approach that integrates GWAS data [14]. The International League Against Epilepsy Consortium on Complex Epilepsies (ILAE) has performed a TWAS with dorsolateral prefrontal cortex tissue, and identified 21 most likely biological epilepsy genes [15]. A TWAS has suggested three significant risk genes for epilepsy: tetratricopeptide repeat domain 21 B (TTC21B), RP11-375N15.2, and tankyrase (TNKS) [11]. These studies contributed to the understanding of the etiology and pathogenesis of epilepsy; however, they rarely explored the correlation between the brain proteome and epilepsy.
Previous studies have established an association between genotype and quantitative trait loci (QTL) on protein (pQTL) abundance and gene expression (eQTL) [16]. Proteome-wide association studies (PWAS) are based on the premise that causal variants in coding regions affect phenotypes by altering the biochemical functions of the gene’s protein products. In PWAS, variant-level associations are decomposed into individual gene values (determined by the association between variants and protein levels), and the combined association of these variants with outcomes is evaluated [17]. PWAS have been widely used to identify candidate genes in brain disorders such as Alzheimer’s disease [18], depression [19], stroke [20], and post-traumatic stress disorder [21]. Therefore, the possible pathogenesis of epilepsy can be explored from gene expression levels of proteins and transcriptomes using GWAS summary data.
In this study, we aimed to determine the influence of genetic and environmental factors on epilepsy by performing TWAS and PWAS analyses based on a GWAS dataset. We investigated gene expression levels in the amygdala, caudate, cingulate, cortex, frontal cortex, hippocampus, hypothalamus, nucleus accumbens, putamen, and substantia nigra. Furthermore, we evaluated the relationship between genes identified by TWAS and PWAS and identified epilepsy-associated chemicals (Fig. 1).
Methods and Materials
Epilepsy GWAS Summary Data
The published GWAS summary data [15] for epilepsy used in our study was based on the International League Against Epilepsy (ILAE) launched by the Consortium on Complex Epilepsies. The analyzed data included 15, 212 cases (childhood absence epilepsy, juvenile absence epilepsy, juvenile myoclonic epilepsy, and generalized tonic-clonic seizures alone) and 29, 677 controls. This study is a large trans-ancestral GWAS meta-analysis that included not only Caucasian populations but also 531 and 147 cases from Asian and African populations, respectively. Single-nucleotide polymorphisms (SNPs) and genetic predictors of relative risk for all epilepsies were performed by using the PLINK-indep-pairwise command and PGA. Further information was detailed in a previously published study [15]. Ethical approval was not applicable to our study as publicly available data were used for all analyses.
TWAS of Epilepsy
TWAS is a powerful method that integrates gene expression with GWAS to identify the genes associated with certain traits. Therefore, it is appropriate to study the genetic etiology of multiple phenotypes [22]. All genome-wide testing burdens were corrected to ensure the TWAS false positive rate was well-controlled in order to measure significant SNP-trait associations [23]. The software program, Functional Summary-based Imputation (FUSION, http://gusevlab.org/projects/fusion/), was used for TWAS and joint analyses of regions containing multiple significant associations [24]. Briefly, we calculated the reference weights of RnaSeq and human brain gene expression panels through the multiple prediction models running in FUSION, selected the best reference weights, unified the GWAS and reference SNPs, and removed/flipped alleles as appropriate. Each feature that expanded to 100,000 bp was defined as contiguous. The minimum p-value for including features in the model was 0.05. Features with r2 > 0.9 were considered identical, while features with r2 < 0.008 were considered independent. Then, we imputed GWAS Z-scores for any reference SNPs that were missing using the “ImpG” algorithm (https://github.com/huwenboshi/ImpG). Output columns included CHR (chromosome), P0 (gene start), P1, HSQ (heritability of the gene), BEST.GWAS.ID (rsID of the most significant GWAS SNP in locus), EQTL.R2 (cross-validation R2 of the best eQTL in the locus), MODELCV.R2 (cross-validation R2 of the best performing model), MODELCV.PV (cross-validation P-value of the best performing model), TWAS.Z (TWAS Z-score), and TWAS.P (TWAS P-value). After estimating the functional-GWAS association statistic, all features tested results were reported. All P values were then subjected to multiple testing corrections using the Benjamini–Hochberg procedure to gather Q values, which represent the minimum false discovery rate (FDR) threshold at which the contact is deemed significant [25]. FDR is less conservative than the Bonferroni approach and has greater ability (i.e., power) to find truly significant results [26]; thus, all P values are subjected to multiple testing correction using the FDR [27]. TWASfdr <0.05 was used as the significance threshold [28, 29].
High-frequency oscillations (HFOs, 80–500 Hz) recorded in intracranial electroencephalography have been regarded as neurobiomarkers for epileptogenic tissues [30]. HFOs can be identified not only in epileptic cerebral tissue but also in non-epileptic sites [31]. Thus, gene expression weights from many brain regions, including the amygdala, caudate, cingulate, cortex, frontal cortex, hippocampus, hypothalamus, nucleus accumbens, putamen, and substantia nigra, were used as references.
PWAS of Epilepsy
Human brain proteome reference weight data were obtained using the Religious Orders Study and Rush Memory and Aging Project (ROS/MAP) and Banner Sun Health Research Institute (Banner) study data [32]. Individuals from the ROS/MAP and Banner were genotyped using the Affymetrix Precision Medicine Array following the manufacturer’s protocol, using the Qiagen GenePure kit to extract and purify DNA from brain cells. Genotyping was imputed to the 1000 Genome Project Phase 356 using the Michigan Imputation Server 57, and SNPs with imputation R2 > 0.3 were retained [19]. Genotyping was filtered to include only sites on the linkage disequilibrium reference panel provided by the FUSION pipeline. The epilepsy genetic effect (PWAS z-score) was calculated to evaluate the effects of significant SNPs in the GWAS on protein abundance. Finally, FUSION identified candidate genes that were associated with epilepsy that regulate protein abundance in the brain. The default settings and parameters recommended by FUSION were used for the analysis. All P values are subjected to multiple testing corrections using the FDR. PWASfdr <0.05 was used as the significance threshold.
Gene Expression Validation of Epilepsy
Gene profiles (GSE88992) were downloaded from the Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/). This study aimed to investigate the underlying molecular mechanisms in epilepsy. The epilepsy mouse model obtained by intrahippocampal microinjection of kainate (KA; 1 nmol/50 nL) was used in parallel with saline-injected animals as controls [33]. GSE88992 was downloaded from the GEO database using the GEO query package. Statistical analysis and visualization were performed using the R packages, “GEOquery” [34], “limma” [35], “ComplexHeatmap” [36], and “ggplot2.” Differentially expressed genes (DEGs) were identified based on adjusted P values. Veen plot was performed using the R packages “ggplot2.”
Protein-Protein Interaction (PPI) Network and Transcription Factor (TF) Prediction
PPI networks and TFs were analyzed by using the STRING v11.5 database (STRING, https://string-db.org), which required a confidence score of 0.15 and “active interaction sources,” based on a previous study [28]. Cytoscape was used to visualize all interaction networks [29]. AnimalTFDB 4.0 database (http://bioinfo.life.hust.edu.cn) [37] and UCSC Genome Browser database (http://genome.ucsc.edu) [38] were used to predict TFs.
Chemical-Related Gene Set Enrichment Analysis (CGSEA)
The chemical-gene expression annotation dataset used in this study was downloaded from the Comparative Toxicology Genomics Database (http://ctdbase.org/downloads/), which is a publicly available database that provides manually curated information about chemical–gene/protein interactions, chemical–disease, and gene–disease relationships [39]. CGSEA is a flexible tool for assessing the associations between chemicals and complex diseases. A detailed analysis method is provided in the original article [40]. In the present study, 5000 permutations were performed to obtain the empirical distribution of GSEA statistical data [41] for each chemical. Subsequently, the p-value of each chemical was calculated based on the empirical distribution of the CGSEA data. Based on previous studies [42], gene sets containing < 10 or > 200 genes were excluded to control for the influence of gene set size on the results. Chemicals with a p-value < 0.05 and absolute normalized enrichment score (NES) values of ≥ 1 were considered significantly enriched according to GSEA [43].
Results
TWAS Analysis of Epilepsy
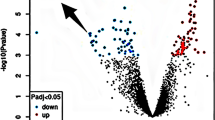
TWAS analysis identified 21,170 genes from the epilepsy GWAS summary data (Fig. 2). After FDR correction, 58 unduplicated genes showed a significant association with epilepsy (TWASfdr < 0.05; Supplementary Information 1).
Manhattan plots from the epilepsy transcriptome-wide association study. The horizon line represents Ptwas = 0.001. Each dot represents the genetically predicted expression of one specific gene for the amygdala, caudate, cingulate, cortex, frontal cortex, hippocampus, hypothalamus,nucleus accumbens, putamen, and substantia nigra. The x axis represents the genomic position of the corresponding gene, and the y axis represents the negative logarithm of the association PTWAS
A comprehensive TWAS was performed to predict the relationship between epilepsy and significant genes found in the amygdala, caudate, cingulate, cortex, frontal cortex, hippocampus, hypothalamus, nucleus accumbens, putamen, and substantia nigra. These results are consistent with previous studies that have identified some risk loci for epilepsy [11, 15] (see Table 1).
Common Genes Identified by TWAS and mRNA Expression Profiling
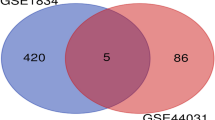
To verify the reliability of the 58 TWAS-identified significant genes, we selected and analyzed the GEO dataset (GSE88992, Supplementary Information 2). GSE88992 was normalized and corrected (Fig. 3A). GSE88992 (Fig. 3B) contained 8520 genes and shared 16 common genes with TWAS-identified significant genes (Table 2, Fig. 3C). The expression of 16 overlapping genes are shown in Fig. 3D.
PWAS Analysis of Epilepsy
PWAS analysis identified 2249 protein-coding genes from the epilepsy GWAS summary data (Fig. 4), wherein two protein-coding genes showed a significant association with epilepsy (PWASfdr < 0.05; Supplementary Information 3). These two epilepsy-susceptibility genes identified by PWAS were IQSEC1 (IQ motif and Sec7 Domain ArfGEF 1, chromosome 3), and JAM2 (junctional adhesion molecule 2, chromosome 21).
Manhattan plots from the epilepsy protein-wide association study. The horizon line represents Ppwas = 0.001. Each dot represents the genetically predicted expression of one specific protein-coding gene from the human brain. The x axis represents the genomic position of the corresponding gene, and the y axis represents the negative logarithm of the association PPWAS
Molecular Interaction of the Genes Identified by TWAS and PWAS
To verify the relationship between genes identified by TWAS and PWAS genes, we searched PWAS-identified gene-related proteins and TFs (Supplementary Information 4). Interestingly, ICAM3 (intercellular adhesion molecule 3, chromosome 19) was observed to interact with JAM2. ZNF143 (zinc finger protein 143, chromosome 13) is predicted to be a TF common to both IQSEC1 and JAM2 (Fig. 5).
CGSEA of the TWAS-Identified Genes
CGSEA was performed to investigate the environmental factors that influence epilepsy. A total of 14,918 chemicals were identified, including 159 chemicals that correlated significantly with epilepsy (PCGSEA < 0.05, |NES| ≥ 1, Supplementary Information 5). Figure 6 illustrates the constructed network of chemicals based on TWAS-identified and PWAS-identified genes.
Discussion
Epilepsy has a higher incidence in children aged younger than 5 years than in those aged >5 years; moreover, children who present with epilepsy experience a high burden of cognitive and behavioral comorbidities [3]. Most seizures in patients with epilepsy arise from discrete pathological regions of abnormal brain tissue [44], including the frontal [45], temporal [46], occipital [47], and parietal lobes [48]. Thus, exploring as many brain regions as possible will provide new information for identifying biomarkers and understanding the etiology of epilepsy.
In this study, we identified several new candidate genes for epilepsy, such as WIPF1, IQSEC1, JAM2, ICAM3, and ZNF143. WIPF1 (WAS/WASL interacting protein family member 1, related to endocytosis) has been identified by TWAS and mRNA expression profiles. An animal study demonstrated the potential significance of DNA methylation-dependent endocytosis regulation in the pathophysiology of temporal lobe epilepsy [49]. IQSEC1 has been predicted to be involved in the regulation of postsynaptic neurotransmitter receptor internalization. Mutations of IQSEC1 were found to be associated with intellectual disability [50], Alzheimer’s disease [51], and attention-deficit/hyperactivity disorder [52]. However, the potential role of IQSEC1 in epilepsy has not been identified. Primary familial brain calcification (PFBC) is a neurodegenerative disorder characterized by calcium deposition in bilateral and symmetric brain [53]. Some PFBC patients would exhibit symptoms of neuropsychiatric symptoms, movement disorders, and epilepsy [54]. A case report showed Bi-allelic JAM2 variants lead to learning difficulties and seizures [55]. Inflammatory cells, molecules, and associated pathways found in the nervous system and systemic tissues have been regarded as key factors in the development of epilepsy [56]. The intercellular adhesion molecule (ICAM) family is a subfamily of the immunoglobulin superfamily associated with immune responses, inflammation, and intracellular signaling [57]. ICAM3 is a member of the ICAM family and can be expressed in gitter cells [58]. The neuronal protein Kidins220/ARMS, a protein that is mainly expressed in brain and neural cells, has been verified to coimmunoprecipitate with ICAM3 to regulate T-cell motility [59]. Therefore, we speculate that ICAM3 may have a potential role in inflammation-related epilepsy, though further research is needed. Primary mitochondrial disease is a systemic disease, and seizures may be its presenting feature [60].
Prior studies have noted that mitochondrial oxidative stress and dysfunction have played a key role not only in epileptogenesis, but also result from seizures [61]. ZNF143 may protect cells from oxidative damage during mitochondrial dysfunction [62]. Cobalamin deficiency [63] and cobalamin-related remethylation disorder [64] may contribute to epilepsy. Whole-exome sequencing suggested mutations in ZNF143 are associated with inborn error of cobalamin metabolism [65]. These results implied the association between ZNF143 and epilepsy.
Antiseizure medications are the mainstay of treatment for patients with newly diagnosed epilepsy [66]. Although many antiepileptic drugs are available, approximately 30% of patients with epilepsy have uncontrolled seizures; thus, new therapeutic approaches must still be developed [67]. In this study, we extended the classic GSEA approach to detect the association between chemicals and epilepsy from our TWAS and PWAS results and identified 159 significantly enriched chemicals. Antiepileptic drugs [68] such as ketamine, corticosterone, pentobarbital, and ketone bodies were identified in this study. A ketogenic diet, particularly polyunsaturated fatty acids, is an effective treatment for refractory epilepsy [69] and may be beneficial for these patients [70]. This study confirms that polyunsaturated fatty acids (palm and fish oils) are associated with epilepsy, providing genetic evidence for the effects of a ketogenic diet on epilepsy. Vitamin E as a neuroprotective agent has been proven to be a useful therapeutic approach in epilepsy treatment [71]. Seizures can be manifestations of intracranial tumors [72]; thus, our results also identified some antineoplastic drugs, such as buthionine sulfoximine, methotrexate, vincristine, temozolomide, etoposide, and cyclophosphamide. Selenium has been suggested to be associated with epilepsy, which further supports the idea that selenium deficiency increases the risk of seizures, while supplementation may help alleviate seizure frequency and duration [73]. Genetic and environmental factors in early life are also important for neurodevelopment [74]. The etiology of neurodevelopmental disorders involves complex interactions between genes and the environment, which has been confirmed in schizophrenia [75] and autism [76]. Environmental chemical exposures in early life, including endocrine-disrupting chemicals (EDCs), influence health and disease susceptibility across the life course [77]. Multiple EDCs were identified in our study, including zearalenone, paraquat, trichloroethylene, polychlorinated biphenyls, and triclosan. Animal research has shown that polychlorinated biphenyl exposure increases the susceptibility to seizures in adulthood [78, 79]. EDCs have been widely elucidated as important risk factors for metabolic diseases such as diabetes and obesity [80]. Although EDCs are ubiquitous in the environment and affect human health [81], there are few published studies on the relationship between EDCs and epilepsy. Thus, more research is urgently needed to understand the mechanisms underlying the genetic predispositions and EDCs related to epilepsy.
Nevertheless, this study has some limitations. First, some genes related to epilepsy susceptibility identified here have not been verified by molecular biology experiments, which should be performed in future studies. Second, the pooled GWAS data are predominantly from European population and it is common to find type-I errors that are not under control for various reasons in GWAS analyses. Therefore, our results should be used with caution when studying other populations. Furthermore, some chemicals that were identified in this study have been previously demonstrated to play a role in epilepsy, whereas others have not yet been validated, which will require more clinical observations and cohort studies. However, to the best of our knowledge, this is the first large-scale study that used PWAS and CGSEA to identify candidate genes and chemicals related to epilepsy.
In this study, we aimed to determine the effects of genetic and environmental factors on epilepsy. Therefore, we performed TWAS and PWAS on epilepsy and identified multiple epilepsy-associated genes. We further performed CGSEA to identify multiple epilepsy-associated chemicals, including EDCs. Our results provide genetic evidence of the effects of EDCs on epilepsy. The results of this study expand our understanding of the genetic and environmental factors that affect epilepsy as well as potential novel drug targets.
Data Availability
The datasets analyzed during the current study are available from the Database of Genomic Variants (http://projects.tcag.ca/variation/); the URLs for Consortia and Groups (https://www.preventcd.com); the BioGPS (http://biogps.gnf.org); the Gene Expression Omnibus database (https://www.ncbi.nlm.nih.gov/gds) (accession numbers GSE88992); and the UK biobank (http://geneatlas.roslin.ed.ac.uk/) (fields: 20002).
References
Fernandez-Albert J et al (2019) Immediate and deferred epigenomic signatures of in vivo neuronal activation in mouse hippocampus. Nat Neurosci 22(10):1718–1730
Dabrowska N et al (2019) Parallel pathways of seizure generalization. Brain 142(8):2336–2351
Symonds JD et al (2019) Incidence and phenotypes of childhood-onset genetic epilepsies: a prospective population-based national cohort. Brain 142(8):2303–2318
Oyrer J et al (2018) Ion channels in genetic epilepsy: from genes and mechanisms to disease-targeted therapies. Pharmacol Rev 70(1):142–173
Razaz N et al (2017) Maternal body mass index in early pregnancy and risk of epilepsy in offspring. JAMA Neurol 74(6):668–676
Whelan CD et al (2018) Structural brain abnormalities in the common epilepsies assessed in a worldwide ENIGMA study. Brain 141(2):391–408
Cieśluk M et al (2020) Nanomechanics and histopathology as diagnostic tools to characterize freshly removed human brain tumors. Int J Nanomedicine 15:7509–7521
de Kovel CGF et al (2017) Neurodevelopmental disorders caused by de novo variants in KCNB1 genotypes and phenotypes. JAMA Neurol 74(10):1228–1236
Leu C et al (2019) Polygenic burden in focal and generalized epilepsies. Brain 142(11):3473–3481
Guo Y et al (2012) Two-stage genome-wide association study identifies variants in CAMSAP1L1 as susceptibility loci for epilepsy in Chinese. Hum Mol Genet 21(5):1184–1189
Song M et al (2021) Genome-wide meta-analysis identifies two novel risk loci for epilepsy. Front Neurosci 15:722592
Lin H et al (2022) Potential pleiotropic genes and shared biological pathways in epilepsy and depression based on GWAS summary statistics. Comput Intell Neurosci 2022:6799285
Wainberg M et al (2019) Opportunities and challenges for transcriptome-wide association studies. Nat Genet 51(4):592–599
Cao C et al (2021) Power analysis of transcriptome-wide association study: implications for practical protocol choice. PLoS Genet 17(2):e1009405
The International League Against Epilepsy Consortium on Complex Epilepsies (2018) Genome-wide mega-analysis identifies 16 loci and highlights diverse biological mechanisms in the common epilepsies. Nat Commun 9(1):5269. https://doi.org/10.1038/s41467-018-07524-z
Wang M et al (2018) The Mount Sinai cohort of large-scale genomic, transcriptomic and proteomic data in Alzheimer’s disease. Sci Data 5:180185
Brandes N, Linial N, Linial M (2020) PWAS: proteome-wide association study-linking genes and phenotypes by functional variation in proteins. Genome Biol 21(1):173
Wingo AP et al (2021) Integrating human brain proteomes with genome-wide association data implicates new proteins in Alzheimer’s disease pathogenesis. Nat Genet 53(2):143–146
Wingo TS et al (2021) Brain proteome-wide association study implicates novel proteins in depression pathogenesis. Nat Neurosci 24(6):810–817
Wu BS et al (2022) Identifying causal genes for stroke via integrating the proteome and transcriptome from brain and blood. J Transl Med 20(1):181
Zhang Z et al (2022) Brain proteome-wide association study identifies candidate genes that regulate protein abundance associated with post-traumatic stress disorder. Genes (Basel) 13(8)
Gusev A et al (2016) Integrative approaches for large-scale transcriptome-wide association studies. Nat Genet 48(3):245–252
Yao DW et al (2020) Quantifying genetic effects on disease mediated by assayed gene expression levels. Nat Genet 52(6):626–633
Pain O et al (2019) Novel insight into the etiology of autism spectrum disorder gained by integrating expression data with genome-wide association statistics. Biol Psychiatry 86(4):265–273
Noble WS (2009) How does multiple testing correction work? Nat Biotechnol 27(12):1135–1137
Perneger TV (1998) What’s wrong with Bonferroni adjustments. Bmj 316(7139):1236–1238
Dall’Aglio L, Lewis CM, Pain O (2021) Delineating the genetic component of gene expression in major depression. Biol Psychiatry 89(6):627–636
Konuma T, Ogawa K, Okada Y (2021) Integration of genetically regulated gene expression and pharmacological library provides therapeutic drug candidates. Hum Mol Genet 30(3-4):294–304
Liu D et al (2022) A transcriptome-wide association study identifies novel candidate susceptibility genes for prostate cancer risk. Int J Cancer 150(1):80–90
Zijlmans M et al (2011) Ictal and interictal high frequency oscillations in patients with focal epilepsy. Clin Neurophysiol 122(4):664–671
Liu S et al (2018) Stereotyped high-frequency oscillations discriminate seizure onset zones and critical functional cortex in focal epilepsy. Brain 141(3):713–730
Beach TG et al (2015) Arizona study of aging and neurodegenerative disorders and brain and body donation program. Neuropathology 35(4):354–389
Kalozoumi G et al (2018) Glial responses during epileptogenesis in Mus musculus point to potential therapeutic targets. PLoS One 13(8):e0201742
Davis S, Meltzer PS (2007) GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 23(14):1846–1847
Ritchie ME et al (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43(7):e47
Gu Z, Eils R, Schlesner M (2016) Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32(18):2847–2849
Shen WK et al (2023) AnimalTFDB 4.0: a comprehensive animal transcription factor database updated with variation and expression annotations. Nucleic Acids Res 51(D1):D39–d45
Kent WJ et al (2002) The human genome browser at UCSC. Genome Res 12(6):996–1006
Mattingly CJ et al (2004) Promoting comparative molecular studies in environmental health research: an overview of the comparative toxicogenomics database (CTD). Pharmacogenomics J 4(1):5–8
Cheng S et al (2020) CGSEA: A flexible tool for evaluating the associations of chemicals with complex diseases. G3 (Bethesda) 10(3):945–949
Subramanian A et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102(43):15545–15550
Cheng S et al (2020) Identifying 5 common psychiatric disorders associated chemicals through integrative analysis of genome-wide association study and chemical-gene interaction datasets. Schizophr Bull 46(5):1182–1190
Mooney MA, Wilmot B (2015) Gene set analysis: a step-by-step guide. Am J Med Genet B Neuropsychiatr Genet 168(7):517–527
Li P et al (2017) Loss of CLOCK results in dysfunction of brain circuits underlying focal epilepsy. Neuron 96(2):387–401.e6
Vignatelli L et al (2015) Prevalence of nocturnal frontal lobe epilepsy in the adult population of Bologna and Modena, Emilia-Romagna region Italy. Sleep 38(3):479–485
Gourmaud S et al (2020) Alzheimer-like amyloid and tau alterations associated with cognitive deficit in temporal lobe epilepsy. Brain 143(1):191–209
Vaudano AE et al (2017) Photosensitive epilepsy is associated with reduced inhibition of alpha rhythm generating networks. Brain 140(4):981–997
Lin JJ, Mula M, Hermann BP (2012) Uncovering the neurobehavioural comorbidities of epilepsy over the lifespan. Lancet 380(9848):1180–1192
Pensold D et al (2020) DNA methylation-mediated modulation of endocytosis as potential mechanism for synaptic function regulation in murine inhibitory cortical interneurons. Cereb Cortex 30(7):3921–3937
Ansar M et al (2019) Bi-allelic variants in IQSEC1 cause intellectual disability, developmental delay, and short stature. Am J Hum Genet 105(5):907–920
Brabec JL et al (2021) System-level analysis of Alzheimer’s disease prioritizes candidate genes for neurodegeneration. Front Genet 12:625246
Sun Y et al (2023) Associations of DNA methylation with behavioral problems, gray matter volumes, and negative life events across adolescence: evidence from the longitudinal IMAGEN study. Biol Psychiatry 93(4):342–351
Zhan FX et al (2020) Primary familial brain calcification presenting as paroxysmal kinesigenic dyskinesia: genetic and functional analyses. Neurosci Lett 714:134543
Knowles JK et al (2018) Refractory focal epilepsy in a paediatric patient with primary familial brain calcification. Seizure 56:50–52
Schottlaender LV et al (2020) Bi-allelic JAM2 variants lead to early-onset recessive primary familial brain calcification. Am J Hum Genet 106(3):412–421
Rana A, Musto AE (2018) The role of inflammation in the development of epilepsy. J Neuroinflammation 15(1):144
Shen W et al (2018) ICAM3 mediates inflammatory signaling to promote cancer cell stemness. Cancer Lett 422:29–43
Serafini B et al (2006) Dendritic cells in multiple sclerosis lesions: maturation stage, myelin uptake, and interaction with proliferating T cells. J Neuropathol Exp Neurol 65(2):124–141
Jean-Mairet RM et al (2011) The neuronal protein Kidins220/ARMS associates with ICAM-3 and other uropod components and regulates T-cell motility. Eur J Immunol 41(4):1035–1046
Lopriore P et al (2022) Mitochondrial epilepsy, a challenge for neurologists. Int J Mol Sci 23(21)
Wu Y, Chen M, Jiang J (2019) Mitochondrial dysfunction in neurodegenerative diseases and drug targets via apoptotic signaling. Mitochondrion 49:35–45
Lu W et al (2012) ZNF143 transcription factor mediates cell survival through upregulation of the GPX1 activity in the mitochondrial respiratory dysfunction. Cell Death Dis 3(11):e422
Anastogiannis H et al (2014) Cobalamin deficiency triggering de novo status epilepticus. Epileptic Disord 16(1):138–139
Gowda VK et al (2021) A treatable cause of global developmental delay with epileptic spasm and delayed myelination due to cobalamin-related remethylation disorder. Indian J Pediatr 88(11):1156–1157
Pupavac M et al (2016) Inborn error of cobalamin metabolism associated with the intracellular accumulation of transcobalamin-bound cobalamin and mutations in ZNF143, which codes for a transcriptional activator. Hum Mutat 37(9):976–982
Alsfouk BAA et al (2020) Tolerability of antiseizure medications in individuals with newly diagnosed epilepsy. JAMA Neurol 77(5):574–581
Delahaye-Duriez A et al (2016) Rare and common epilepsies converge on a shared gene regulatory network providing opportunities for novel antiepileptic drug discovery. Genome Biol 17(1):245
Vossler DG et al (2020) Treatment of refractory convulsive status epilepticus: a comprehensive review by the American Epilepsy Society Treatments Committee. Epilepsy Curr 20(5):245–264
Olson CA et al (2018) The gut microbiota mediates the anti-seizure effects of the ketogenic diet. Cell 173(7):1728–1741.e13
Nathan J et al (2019) A switch to polyunsaturated fatty acid based ketogenic diet improves seizure control in patients with drug-resistant epilepsy on the mixed fat ketogenic diet: a retrospective open label trial. Cureus 11(12):e6399
Upaganlawar AB et al (2021) Interweaving epilepsy and neurodegeneration: vitamin E as a treatment approach. Biomed Pharmacother 143:112146
Tewari BP et al (2018) Perineuronal nets decrease membrane capacitance of peritumoral fast spiking interneurons in a model of epilepsy. Nat Commun 9(1):4724
Pillai R, Uyehara-Lock JH, Bellinger FP (2014) Selenium and selenoprotein function in brain disorders. IUBMB Life 66(4):229–239
Oberg AS et al (2016) Association of labor induction with offspring risk of autism spectrum disorders. JAMA Pediatr 170(9):e160965
Reay WR, Cairns MJ (2020) The role of the retinoids in schizophrenia: genomic and clinical perspectives. Mol Psychiatry 25(4):706–718
Courchet V et al (2018) Haploinsufficiency of autism spectrum disorder candidate gene NUAK1 impairs cortical development and behavior in mice. Nat Commun 9(1):4289
Treviño LS et al (2020) Epigenome environment interactions accelerate epigenomic aging and unlock metabolically restricted epigenetic reprogramming in adulthood. Nat Commun 11(1):2316
Poon E et al (2015) Developmental PCB exposure increases susceptibility to audiogenic seizures in adulthood. Neurotoxicology 46:117–124
Bandara SB et al (2016) Developmental PCB exposure increases audiogenic seizures and decreases glutamic acid decarboxylase in the inferior colliculus. Toxicol Sci 149(2):335–345
Sun J et al (2022) A review of environmental metabolism disrupting chemicals and effect biomarkers associating disease risks: where exposomics meets metabolomics. Environ Int 158:106941
Kumar M et al (2020) Environmental endocrine-disrupting chemical exposure: role in non-communicable diseases. Front Public Health 8:553850
Acknowledgements
We are indebted to all the individuals who participated in and helped with our research.
Author information
Authors and Affiliations
Contributions
Author Mengnan Lu and Ruoyang Feng collected and processed the data and wrote the article. Chenglin Zhang provided language help and writing assistance. Chunyan Yin and Yanfeng Xiao designed the study. All authors contributed to the revisions, approved the final manuscript, and made the decision to submit the manuscript for publication.
Corresponding authors
Ethics declarations
Ethics Approval
Ethical approval was not applicable to our study as publicly available data were used for all analyses.
Consent to Participate
We are indebted to all the individuals who participated in and helped with our research.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lu, M., Feng, R., Zhang, C. et al. Identifying Novel Drug Targets for Epilepsy Through a Brain Transcriptome-Wide Association Study and Protein-Wide Association Study with Chemical-Gene-Interaction Analysis. Mol Neurobiol 60, 5055–5066 (2023). https://doi.org/10.1007/s12035-023-03382-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-023-03382-z