Abstract
Contactin-associated protein-like 2 (CNTNAP2) gene, located on chromosome 7q35, is one of the largest genes in the human genome. CNTNAP2 protein is a type-I transmembrane protein specifically expressed in the nervous system, with versatile roles in the axonal organization, synaptic functions, neuronal migration, and functional connectivity. CNTNAP2 has been widely investigated as a risk gene for autism spectrum disorder (ASD), and recent studies also implicated CNTNAP2 in Alzheimer’s disease (AD). Knowledge of the regulations on CNTNAP2’s life cycle is necessary for understanding the related physiological functions and pathological conditions. However, the mechanisms underlying CNTNAP2 protein degradation remain elusive. Therefore, we systematically investigated the half-life and degradation pathway of the human CNTNAP2 protein. We discovered that CNTNAP2 has C-terminal fragments (CTF), which may have essential physiological functions. Our results demonstrated that CNTNAP2 full-length protein and CTF have a short half-life of about 3–4 h. CNTNAP2 proteins are degraded by the ubiquitin–proteasome system and the macroautophagy-lysosome pathway, while the lysosome pathway is more common for CNTNAP2 degradation. This study will provide novel insights and valuable tools for CNTNAP2 functional research in physiological and pathological scenarios.







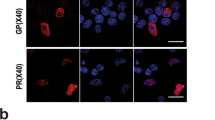
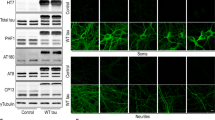
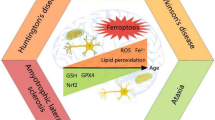
Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article.
References
Liu B, Ruan J, Chen M, Li Z, Manjengwa G, Schluter D, Song W, Wang X (2021) Deubiquitinating enzymes (DUBs): decipher underlying basis of neurodegenerative diseases. MolPsychiatr 27:259–268
Liu H, Wang P, Song W, Sun X (2009) Degradation of regulator of calcineurin 1 (RCAN1) is mediated by both chaperone-mediated autophagy and ubiquitin proteasome pathways. FASEB J 23(10):3383–3392. https://doi.org/10.1096/fj.09-134296
Glenner GG, Wong CW (1984) Alzheimer’s disease: initial report of the purification and characterization of a novel cerebrovascular amyloid protein. Biochem Biophys Res Commun 120(3):885–890
Braak H, Braak E (1991) Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol 82(4):239–259
Zhang Y, Song W (2017) Islet amyloid polypeptide: another key molecule in Alzheimer's pathogenesis? Prog Neurobiol 153:100-120. https://doi.org/10.1111/dom.137331016/j.pneurobio.2017.03.001.
Mawuenyega KG, Sigurdson W, Ovod V, Munsell L, Kasten T, Morris JC, Yarasheski KE, Bateman RJ (2010) Decreased clearance of CNS β-amyloid in Alzheimer’s disease. Science 330(6012):1774–1774
Gowrishankar S, Yuan P, Wu Y, Schrag M, Paradise S, Grutzendler J, De Camilli P, Ferguson SM (2015) Massive accumulation of luminal protease-deficient axonal lysosomes at Alzheimer’s disease amyloid plaques. Proc Natl Acad Sci U S A 112(28):E3699–3708. https://doi.org/10.1073/pnas.1510329112
He G, Qing H, KwoK C, Xu H, Yu G, Bernstein A, Song W (2006) Ubiquitin-proteasome pathway mediates degradation of APH-1. J Neurochem 99(5):1403–1412
He G, Qing H, Tong Y, Cai F, Ishiura S, Song W (2007) Degradation of nicastrin involves both proteasome and lysosome. J Neurochem 101(4):982–992. https://doi.org/10.1111/j.1471-4159.2007.04449.x
Liu S, Bromley-Brits K, Xia K, Mittelholtz J, Wang R, Song W (2008) TMP21 degradation is mediated by the ubiquitin-proteasome pathway. Eur J Neurosci 28(10):1980–1988. https://doi.org/10.1111/j.1460-9568.2008.06497.x
Liu X, Wang Z, Wu Y, Wang J, Song W (2013) BACE2 degradation mediated by the macroautophagy-lysosome pathway. Eur J Neurosci 37(12):1970–1977. https://doi.org/10.1111/ejn.12204
Qing H, Zhou W, Christensen MA, Sun X, Tong Y, Song W (2004) Degradation of BACE by the ubiquitin-proteasome pathway. FASEB J 18(13):1571–1573. https://doi.org/10.1096/fj.04-1994fje
Hetz C (2021) Adapting the proteostasis capacity to sustain brain healthspan. Cell 184(6):1545–1560. https://doi.org/10.1016/j.cell.2021.02.007
Winckler B, Faundez V, Maday S, Cai Q, Guimas Almeida C, Zhang H (2018) The endolysosomal system and proteostasis: from development to degeneration. J Neurosci 38(44):9364–9374. https://doi.org/10.1523/jneurosci.1665-18.2018
Louros SR, Osterweil EK (2016) Perturbed proteostasis in autism spectrum disorders. J Neurochem 139(6):1081–1092. https://doi.org/10.1111/jnc.13723
Levy SE, Mandell DS, Schultz RT (2009) Autism. Lancet 374(9701):1627–1638. https://doi.org/10.1016/s0140-6736(09)61376-3
Jiang YH, Armstrong D, Albrecht U, Atkins CM, Noebels JL, Eichele G, Sweatt JD, Beaudet AL (1998) Mutation of the Angelman ubiquitin ligase in mice causes increased cytoplasmic p53 and deficits of contextual learning and long-term potentiation. Neuron 21(4):799–811. https://doi.org/10.1016/s0896-6273(00)80596-6
Pignatelli M, Piccinin S, Molinaro G, Di Menna L, Riozzi B, Cannella M, Motolese M, Vetere G et al (2014) Changes in mGlu5 receptor-dependent synaptic plasticity and coupling to homer proteins in the hippocampus of Ube3A hemizygous mice modeling angelman syndrome. J Neurosci 34(13):4558–4566. https://doi.org/10.1523/jneurosci.1846-13.2014
Basel-Vanagaite L, Dallapiccola B, Ramirez-Solis R, Segref A, Thiele H, Edwards A, Arends MJ, Miró X et al (2012) Deficiency for the ubiquitin ligase UBE3B in a blepharophimosis-ptosis-intellectual-disability syndrome. Am J Hum Genet 91(6):998–1010. https://doi.org/10.1016/j.ajhg.2012.10.011
Chahrour MH, Yu TW, Lim ET, Ataman B, Coulter ME, Hill RS, Stevens CR, Schubert CR, Greenberg ME, Gabriel SB, Walsh CA (2012) Whole-exome sequencing and homozygosity analysis implicate depolarization-regulated neuronal genes in autism. PLoS Genet 8(4):e1002635. https://doi.org/10.1371/journal.pgen.1002635
O’Roak BJ, Vives L, Girirajan S, Karakoc E, Krumm N, Coe BP, Levy R, Ko A et al (2012) Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 485(7397):246–250. https://doi.org/10.1038/nature10989
Rapanelli M, Tan T, Wang W, Wang X, Wang ZJ, Zhong P, Frick L, Qin L et al (2021) Behavioral, circuitry, and molecular aberrations by region-specific deficiency of the high-risk autism gene Cul3. Mol Psychiatry 26(5):1491–1504. https://doi.org/10.1038/s41380-019-0498-x
Poliak S, Gollan L, Martinez R, Custer A, Einheber S, Salzer JL, Trimmer JS, Shrager P et al (1999) Caspr2, a new member of the neurexin superfamily, is localized at the juxtaparanodes of myelinated axons and associates with K+ channels. Neuron 24(4):1037–1047. https://doi.org/10.1016/s0896-6273(00)81049-1
Spiegel I, Salomon D, Erne B, Schaeren-Wiemers N, Peles E (2002) Caspr3 and caspr4, two novel members of the caspr family are expressed in the nervous system and interact with PDZ domains. Mol Cell Neurosci 20(2):283–297
Poliak S, Salomon D, Elhanany H, Sabanay H, Kiernan B, Pevny L, Stewart CL, Xu X et al (2003) Juxtaparanodal clustering of Shaker-like K+ channels in myelinated axons depends on Caspr2 and TAG-1. J Cell Biol 162(6):1149–1160. https://doi.org/10.1083/jcb.200305018
Traka M, Goutebroze L, Denisenko N, Bessa M, Nifli A, Havaki S, Iwakura Y, Fukamauchi F et al (2003) Association of TAG-1 with Caspr2 is essential for the molecular organization of juxtaparanodal regions of myelinated fibers. J Cell Biol 162(6):1161–1172. https://doi.org/10.1083/jcb.200305078
Denisenko-Nehrbass N, Oguievetskaia K, Goutebroze L, Galvez T, Yamakawa H, Ohara O, Carnaud M, Girault JA (2003) Protein 4.1B associates with both Caspr/paranodin and Caspr2 at paranodes and juxtaparanodes of myelinated fibres. Eur J Neurosci 17(2):411–416. https://doi.org/10.1046/j.1460-9568.2003.02441.x
Fernandes D, Santos SD, Coutinho E, Whitt JL, Beltrao N, Rondao T, Leite MI et al. (2019) Disrupted AMPA receptor function upon genetic- or antibody-mediated loss of autism-associated CASPR2. Cereb Cortex 29:4919–4931 https://doi.org/10.1002/dvdy.3210.1093/cercor/bhz032
Varea O, Martin-de-Saavedra MD, Kopeikina KJ, Schurmann B, Fleming HJ, Fawcett-Patel JM, Bach A, Jang S et al (2015) Synaptic abnormalities and cytoplasmic glutamate receptor aggregates in contactin associated protein-like 2/Caspr2 knockout neurons. Proc Natl Acad Sci U S A 112(19):6176–6181. https://doi.org/10.1073/pnas.1423205112
Canali G, Garcia M, Hivert B, Pinatel D, Goullancourt A, Oguievetskaia K, Saint-Martin M, Girault JA et al (2018) Genetic variants in autism-related CNTNAP2 impair axonal growth of cortical neurons. Hum Mol Genet 27(11):1941–1954. https://doi.org/10.1093/hmg/ddy102
Anderson GR, Galfin T, Xu W, Aoto J, Malenka RC, Sudhof TC (2012) Candidate autism gene screen identifies critical role for cell-adhesion molecule CASPR2 in dendritic arborization and spine development. Proc Natl Acad Sci USA 109(44):18120–18125. https://doi.org/10.1073/pnas.1216398109
Gdalyahu A, Lazaro M, Penagarikano O, Golshani P, Trachtenberg JT, Geschwind DH (2015) The autism related protein contactin-associated protein-like 2 (CNTNAP2) stabilizes new spines: an in vivo mouse study. PloS one 10(5):e0125633. https://doi.org/10.1371/journal.pone.0125633
Penagarikano O, Abrahams BS, Herman EI, Winden KD, Gdalyahu A, Dong H, Sonnenblick LI, Gruver R et al (2011) Absence of CNTNAP2 leads to epilepsy, neuronal migration abnormalities, and core autism-related deficits. Cell 147(1):235–246. https://doi.org/10.1016/j.cell.2011.08.040
Cope EC, Briones BA, Brockett AT, Martinez S, Vigneron PA, Opendak M, Wang SS, Gould E (2016) Immature neurons and radial glia, but not astrocytes or microglia, are altered in adult Cntnap2 and Shank3 mice, models of autism. Eneuro 3 (5):ENEURO.0196-16.2016.
Zerbi V, Ielacqua GD, Markicevic M, Haberl MG, Ellisman MH, A-Bhaskaran A, Frick A, Rudin M et al (2018) Dysfunctional autism risk genes cause circuit-specific connectivity deficits with distinct developmental trajectories. Cereb Cortex 28(7):2495–2506. https://doi.org/10.1093/cercor/bhy046
Liska A, Bertero A, Gomolka R, Sabbioni M, Galbusera A, Barsotti N, Panzeri S, Scattoni ML, Pasqualetti M, Gozzi A (2018) Homozygous loss of autism-risk gene CNTNAP2 results in reduced local and long-range prefrontal functional connectivity. Cereb Cortex 28(4):1141–1153. https://doi.org/10.1093/cercor/bhx022
Choe KY, Bethlehem RAI, Safrin M, Dong H, Salman E, Li Y, Grinevich V, Golshani P et al (2022) Oxytocin normalizes altered circuit connectivity for social rescue of the Cntnap2 knockout mouse. Neuron 110(5):795-808.e796. https://doi.org/10.1016/j.neuron.2021.11.031
Penagarikano O, Geschwind DH (2012) What does CNTNAP2 reveal about autism spectrum disorder? Trends Mol Med 18(3):156–163. https://doi.org/10.1016/j.molmed.2012.01.003
Bakkaloglu B, O’Roak BJ, Louvi A, Gupta AR, Abelson JF, Morgan TM, Chawarska K, Klin A et al (2008) Molecular cytogenetic analysis and resequencing of contactin associated protein-like 2 in autism spectrum disorders. Am J Hum Genet 82(1):165–173. https://doi.org/10.1016/j.ajhg.2007.09.017
Maekawa M, Yamada K, Toyoshima M, Ohnishi T, Iwayama Y, Shimamoto C, Toyota T, Nozaki Y et al (2015) Utility of scalp hair follicles as a novel source of biomarker genes for psychiatric illnesses. Biol Psychiat 78(2):116–125. https://doi.org/10.1016/j.biopsych.2014.07.025
Sampath S, Bhat S, Gupta S, O’Donnor A, West AB, Arking DE, Chakravarti A (2013) Defining the contribution of CNTNAP2 to autism susceptibility. PloS one 8(10):e77906. https://doi.org/10.1371/journal.pone.0077906
Scott KE, Kazazian K, Mann RS, Möhrle D, Schormans AL, Schmid S, Allman BL (2020) Loss of Cntnap2 in the rat causes autism-related alterations in social interactions, stereotypic behavior, and sensory processing. Autism Res 13(10):1698–1717
Hirano A, Ohara T, Takahashi A, Aoki M, Fuyuno Y, Ashikawa K, Morihara T, Takeda M et al (2015) A genome-wide association study of late-onset Alzheimer’s disease in a Japanese population. Psychiatr Genet 25(4):139–146. https://doi.org/10.1016/j.mcn.2015.06.00510.1097/ypg.0000000000000090
Iakoubov L, Mossakowska M, Szwed M, Puzianowska-Kuznicka M (2015) A common copy number variation polymorphism in the CNTNAP2 gene: sexual dimorphism in association with healthy aging and disease. Gerontology 61(1):24–31. https://doi.org/10.1159/000363320
Bi XA, Cai R, Wang Y, Liu Y (2019) Effective diagnosis of Alzheimer’s disease via multimodal fusion analysis framework. Front Genet 10:976. https://doi.org/10.3389/fgene.2019.00976
van Abel D, Michel O, Veerhuis R, Jacobs M, van Dijk M, Oudejans CB (2012) Direct downregulation of CNTNAP2 by STOX1A is associated with Alzheimer’s disease. Journal of Alzheimer’s disease : JAD 31(4):793–800. https://doi.org/10.3233/jad-2012-120472
Falivelli G, De Jaco A, Favaloro FL, Kim H, Wilson J, Dubi N, Ellisman MH, Abrahams BS et al (2012) Inherited genetic variants in autism-related CNTNAP2 show perturbed trafficking and ATF6 activation. Hum Mol Genet 21(21):4761–4773. https://doi.org/10.1093/hmg/dds320
Wang Z, Xu Q, Cai F, Liu X, Wu Y, Song W (2019) BACE2, a conditional β-secretase, contributes to Alzheimer's disease pathogenesis. JCI Insight 4 (1):e123431. https://doi.org/10.1038/s41392-019-0091-410.1172/jci.insight.123431
Mizushima N, Levine B, Cuervo AM, Klionsky DJ (2008) Autophagy fights disease through cellular self-digestion. Nature 451(7182):1069–1075. https://doi.org/10.1038/nature06639
Seglen PO, Gordon PB (1982) 3-Methyladenine: specific inhibitor of autophagic/lysosomal protein degradation in isolated rat hepatocytes. Proc Natl Acad Sci 79(6):1889–1892
Struhl G, Adachi A (1998) Nuclear access and action of notch in vivo. Cell 93(4):649–660. https://doi.org/10.1016/s0092-8674(00)81193-9
Zhang ZH, Nadeau P, Song WH, Donoviel D, Yuan ML, Bernstein A, Yankner BA (2000) Presenilins are required for gamma-secretase cleavage of beta-APP and transmembrane cleavage of Notch-1. Nat Cell Biol 2(7):463–465
López Salon M, Morelli L, Castaño EM, Soto EF, Pasquini JM (2000) Defective ubiquitination of cerebral proteins in Alzheimer’s disease. J Neurosci Res 62(2):302–310
Nunan J, Shearman MS, Checler F, Cappai R, Evin G, Beyreuther K, Masters CL, Small DH (2001) The C-terminal fragment of the Alzheimer’s disease amyloid protein precursor is degraded by a proteasome-dependent mechanism distinct from γ-secretase. Eur J Biochem 268(20):5329–5336
Cole GM, Huynh TV, Saitoh T (1989) Evidence for lysosomal processing of amyloid β-protein precursor in cultured cells. Neurochem Res 14(10):933–939
Saido T, Leissring MA (2012) Proteolytic degradation of amyloid β-protein. Cold Spring Harb Perspect Med 2(6):a006379
Bel C, Oguievetskaia K, Pitaval C, Goutebroze L, Faivre-Sarrailh C (2009) Axonal targeting of Caspr2 in hippocampal neurons via selective somatodendritic endocytosis. J Cell Sci 122(Pt 18):3403–3413. https://doi.org/10.1242/jcs.050526
Funding
This study was partly supported by Jack Brown and Family Alzheimer's Research Foundation. QZ and IBL are supported by the Four Year Fellowship for PhD and the Faculty of Medicine Graduate Award from the University of British Columbia. QZ received DMCBH Innovation Fund Graduate Trainee Award. IBL is also supported by Jock and Irene Graham Brain Endowment Award.
Author information
Authors and Affiliations
Contributions
QZ and WS conceived and designed the experiments. QZ, KS, FC, LX, and SY performed the experiments. QZ, KS, FC, LX, XM, IBL, and WS analyzed and contributed reagents/materials/analytical tools. QZ and WS wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Ethics approval
Animal experiments were approved by the University of British Columbia Animal Care and Use Committee (Protocol ID: A21-0303, Neuronal Culture) and conducted in accordance with guidelines.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Q., Sterling, K., Xu, L. et al. CNTNAP2 Protein Is Degraded by the Ubiquitin–Proteasome System and the Macroautophagy-Lysosome Pathway. Mol Neurobiol 60, 2455–2469 (2023). https://doi.org/10.1007/s12035-023-03227-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-023-03227-9