Abstract
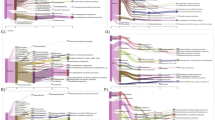
Pyrosequence data was used to analyze the composition and metabolic potential of a metagenome from a hydrocarbon-contaminated site. Unamplified and whole genome amplified (WGA) sequence data was compared from this source. According to MG-RAST, an additional 2,742,252 bp of DNA was obtained with the WGA, indicating that WGA has the ability to generate a large amount of DNA from a small amount of starting sample. However, it was observed that WGA introduced a bias with respect to the distribution of the amplified DNA and the types of microbial populations that were accessed from the metagenome. The dominant order in the WGA metagenome was Flavobacteriales, whereas the unamplified metagenome was dominated by Actinomycetales as determined by RDPII and CARMA databases. According to the SEED database, the subsystems shown to be present for the individual metagenomes were associated with the metabolic potential that was expected to be present in the contaminated groundwater, such as the metabolism of aromatic compounds. A higher percentage (4.4) of genes associated with the metabolism of aromatic compounds was identified in the unamplified metagenome when compared to the WGA metagenome (0.66%). This could be attributed to the increased number of hydrocarbon degrading bacteria that had been accessed from this metagenome (Mycobacteria, Nocardia, Brevibacteria, Clavibacter, Rubrobacter, and Rhodoccocus). Therefore, it was possible to relate the taxonomic groups accessed to the contamination profile of the metagenome. By collating the sequencing data obtained pre- and post-amplification, this study provided insight regarding the survival strategies of microbial communities inhabiting contaminated environments.






Similar content being viewed by others
Abbreviations
- CARMA:
-
CARMA is a software pipeline for the characterization of species composition and the genetic potential of microbial samples using short reads. In contrast to the traditional 16S-rRNA approach for taxonomical classification, CARMA uses reads that encode for known proteins. By assigning the taxonomic origins to each read, a profile is constructed which characterizes the taxonomic composition of the corresponding community
- EGT (Expressed Gene Tag):
-
A unique stretch of DNA within a coding region of a gene that is useful for identifying full-length genes and serves as a landmark for mapping. An EGT is a sequence tagged site (STS) derived from cDNA
- Metagenomics:
-
The genomic analysis of microorganisms by direct extraction and/or cloning of DNA from an assemblage of microorganisms (also refereed to as environmental and community genomics)
- MGRAST:
-
MetaGenome Rapid Annotation Subsystems Technology MGRAST: is a fully automated service for annotating metagenome samples that provides annotation of sequence fragments, their phylogenetic classification, metabolic reconstructions and comparison tools
- Pyrosequencing:
-
A method of DNA sequencing (determining the order of nucleotides in DNA) based on the “sequencing by synthesis” principle. It differs from Sanger sequencing, in that it relies on the detection of pyrophosphate release on nucleotide incorporation, rather than chain termination with dideoxynucleotides
- RDPII:
-
The Ribosomal Database Project (RDP) provides ribosome related data and services to the scientific community, including online data analysis and aligned and annotated Bacterial and Archaeal small-subunit 16S rRNA sequences
- Whole genome amplification:
-
An increasingly common technique through which minute amounts of DNA can be multiplied to generate quantities suitable for genetic testing and analysis
References
Rodríguez-Martínez, E. M., Pérez, E. X., Schadt, C. W., Zhou, J., & Massol-Deyá, A. A. (2006). Microbial diversity and bioremediation of a hydrocarbon-contaminated aquifer (Vega Baja, Puerto Rico). International Journal of Environmental Research and Public Health, 3, 292–300.
Rees, H. C., Oswald, S. E., Banwart, S. A., Pickup, R. W., & Lerner, D. N. (2007). Biodegradation processes in a laboratory-scale groundwater contaminant plume assessed by flourescence imaging and microbial analysis. Applied and Environmental Microbiology, 73, 3865–3876.
Lovely, D. R. (2003). Cleaning up with genomics: Applying molecular biology to bioremediation. Nature Reviews, 1, 35–44.
Grostern, A., & Edwards, E. A. (2006). Growth of Dehalobacter and Dehalococcoides spp. during degradation of chlorinated ethanes. Applied and Environmental Microbiology, 72, 428–432.
Simon, C., Wiezer, A., Strittmatter, A. W., & Daniel, R. (2009). Phylogenetic diversity and metabolic potential revealed in a glacier ice metagenome. Applied and Environmental Microbiology, 75, 7519–7526.
Clarke, J., Wu, H. C., Jayasinghe, L., Patel, A., Reid, S., & Bayley, H. (2009). Continuous base identification for single-molecule nanopore DNA sequencing. Nature Nanotechnology, 4, 265–270.
Zheng, Z., Melefors, O., Glavas, S., Nordström, H., Ye, W., Engstrand, L., et al. (2010). Titration-free massively parallel pyrosequencing using trace amounts of starting material. Nucleic Acids Research, 38, e137. doi:10.1093.
Pinard, E., de Winter, A., Sarkis, G. J., Gerstein, M. B., Tartaro, K. R., Plant, R. N., et al. (2006). Assessment of whole genome amplification-induced bias through high-throughput, massively parallel whole genome sequencing. BMC Genomics, 7, 216. doi:10.1186/1471-2164-7-216.
Hawkins, T. L., Detter, J. C., & Richardson, P. M. (2002). Whole genome amplification—applications and advances. Current Opinion in Biotechnology, 13, 65–67.
Marguiles, M., Egholm, M., Altman, W. E., Attiya, S., Bader, J. S., Bemben, L. A., et al. (2005). Genome sequencing in microfabricated high-density picolitre reactors. Nature, 437, 376–380.
Meyer, F., Paarmann, D., D’Souza, M., Olson, R., Glass, E. M., Kubal, M., et al. (2008). The metagenomics RAST server—a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinformatics, 9, 386. doi:10.1186/1471-2105-9-386.
Altschul, S. F., Madden, T. L., Schaffer, A. A., Zhang, J., Zhang, Z., Miller, W., et al. (1997). Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Research, 25, 3389–3402.
Krause, L., Diaz, N. N., Goesmann, A., Kelley, S., Nattkemper, T. W., Rohwer, F., et al. (2008). Phylogenetic classification of short environmental DNA fragments. Nucleic Acids Research, 36, 2230–2239.
Ventura, M., Canchaya, C., Tauch, A., Chandra, G., Fitzgerald, G. F., Chater, K. F., et al. (2007). Genomics of Actinobacteria: Tracing the evolutionary history of an ancient phylum. Molecular Microbiology Reviews, 71, 495–548.
Woyke, T., Xie, G., Copeland, A., González, J. M., Han, C., Kiss, H., Saw, J. H., Senin, P., Yanga, C., Chatterji, S., Cheng, J.-F., Eisen, J. A., Sierackis, M. E., & Stepanauskas, R. (2009). Assembling the marine metagenome, one cell at a time. 4, e5299. doi:10.1371/journal.pone.0005299. http://www.plosone.org.
Tringe, S. G., von Mering, C., Kobayashi, A., Salamov, A. A., Chen, K., Chang, H. W., et al. (2005). Comparative metagenomics of microbial communities. Science, 308, 554–557.
Edwards, R. A., Rodriquez-Brito, B., Wegley, L., Haynes, Breitbart, M., Peterson, M. D. M., et al. (2006). Using pyrosequencing to shed light on deep mine microbiol ecology. BMC Genomics, 7, 57. doi:10.1186/1471-2164-7-57.
Sanapareddy, N., Hamp, T. J., Gonzalez, L. C., Hilger, H. A., Fodor, A. A., & Clinton, S. M. (2009). Molecular diversity of a North Carolina wastewater treatment plant as revealed by pyrosequencing. Applied and Environmental Microbiology, 75, 1688–1696.
Marzorati, M., de Ferra, F., Van Raemdonck, H., Borin, S., Allifranchini, E., Carpani, G., et al. (2007). A novel reductive dehalogenase, identified in a contaminated groundwater enrichment culture and in Desulfitobacterium dichloroeliminans Strain DCA1, is linked to dehalogenation of 1,2-dichloroethane. Applied and Environmental Microbiology, 73, 2990–2999.
Olaniran, A. O., Bhola, V., & Pillay, B. (2008). Aerobic biodegradation of a mixture of chlorinated organics in contaminated water. African Journal of Biotechnology, 7, 2217–2220.
Horvath, R. S. (1972). Microbial co-metabolism and the degradation of organic compounds in nature. Bacteriology Reviews, 36, 146–155.
Krooneman, J., Sliekers, A. O., Gomes, T. M. P., Forney, L. J., & Gottschal, J. C. (2000). Characterization of 3-chlorobenzoate degrading aerobicbacteria isolated under various environmental conditions. FEMS Microbiology Ecology, 32, 53–59.
Chikere, C. B., Okpokwasili, G. C., & Chikere, B. O. (2009). Bacterial diversity in a tropical crude oil-polluted soil undergoing bioremediation. African Journal of Biotechnology, 8, 2535–2540.
Tomás-Gallardo, L., Canosa, I., Santero, E., Camafeita, E., Calvo, E., López, J. A., et al. (2006). Proteomic and transcriptional characterization of aromatic degradation pathways in Rhodococcus sp. strain TFB. Proteomics, 6, S119–S132.
Heitkamp, A., & Cerniglia, C. E. (1989). Polycyclic aromatic hydrocarbon degradation by a Mycobacterium sp. in microcosms containing sediment and water from a pristine ecosystem. Applied and Environmental Microbiology, 55, 1968–1973.
Miller, M. A., & Lipscomb, J. D. (1996). Homoprotocatechuate 2,3-dioxygenase from Brevibacterium fuscum. Journal of Biological Chemistry, 271, 5524–5535.
Baek, K. H., Yoon, B. D., Lee, I. S., Oh, H. M., & Kim, H. S. (2006). Biodegradation of aliphatic aromatic hydrocarbons by Nocardia sp. H17-1. Geomicrobiology Journal, 23, 253–259.
Kim, S.-J., Kweon, O., Jones, R. C., Edmondson, R. D., & Cerniglia, C. E. (2008). Genomic analysis of polycyclic aromatic hydrocarbon degradation in Mycobacterium vanbaalenii PYR-1. Biodegradation, 19, 859–881.
Nies, D. H. (2003). Efflux-mediated heavy metal resistance in prokaryotes. FEMS Microbiology Reviews, 27, 313–339.
Baumgartner, L. K., Reid, R. P., Dupraz, C., Decho, A. W. D., Buckley, H. J., Spear, R., et al. (2006). Sulfate reducing bacteria in microbial mats: Changing paradigms, new discoveries. Sediment Geology, 185, 131–145.
Acknowledgments
The authors wish to thank the National Research Foundation for financial support and TIA (Dr James Sakwa and colleagues) for the pyrosequencing.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Abbai, N.S., Govender, A., Shaik, R. et al. Pyrosequence Analysis of Unamplified and Whole Genome Amplified DNA from Hydrocarbon-Contaminated Groundwater. Mol Biotechnol 50, 39–48 (2012). https://doi.org/10.1007/s12033-011-9412-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-011-9412-8