Abstract
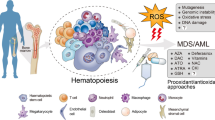
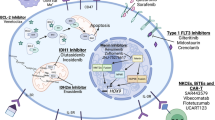
Cancer cells alter their metabolism by switching from glycolysis to oxidative phosphorylation (OXPHOS), regardless of oxygen availability. Metabolism may be a molecular target in acute myeloid leukemia (AML), where mutations in metabolic genes have been described. This study evaluated glycolysis and OXPHOS as therapeutic targets. The sensitivity to 2-deoxy-d-glucose (2-DG; glycolysis inhibitor) and oligomycin (OXPHOS inhibitor) was tested in six AML cell lines (HEL, HL-60, K-562, KG-1, NB-4, THP-1). These cells were characterized for IDH1/2 exon 4 mutations, reactive oxygen species, and mitochondrial membrane potential. Metabolic activity was assessed by resazurin assay, whereas cell death and cell cycle were assessed by flow cytometry. Glucose uptake and metabolism-related gene expression were analyzed by 18F-FDG and RT-PCR/qPCR, respectively. No IDH1/2 exon 4 mutations were detected. HEL cells had the highest 18F-FDG uptake and peroxides/superoxide anion levels, whereas THP-1 showed the lowest. 2-DG reduced metabolic activity in all cell lines with HEL, KG-1, and NB-4 being the most sensitive cells. Oligomycin decreased metabolic activity in a cell line-dependent manner, the THP-1 resistant and HL-60 being the most sensitive. Both inhibitors induced apoptosis and cell cycle arrest in a cell line- and compound-dependent manner. 2-DG decreased 18F-FDG uptake in HEL, HL-60, KG-1, and NB-4, while oligomycin increased the uptake in K-562. Metabolism gene expression had different responses to treatments. In conclusion, HEL and KG-1 show to be more glycolytic, whereas HL-60 was more OXPHOS dependent. Results suggest that AML cells reprogram their metabolism to overcome OXPHOS inhibition suggesting that glycolysis may be a better therapeutic target.




Similar content being viewed by others
Data availability
The data used and/or analyzed during the current study are available by contacting the corresponding author with reasonable request.
References
Ghaffari P, Mardinoglu A, Nielsen J. Cancer metabolism: a modeling perspective. Front. Physiol. 2015;6:382. https://doi.org/10.3389/fphys.2015.00382.
Schulze A, Harris AL. How cancer metabolism is tuned for proliferation and vulnerable to disruption. Nature. 2012;491(7424):364–73. https://doi.org/10.1038/nature11706.
Koppenol WH, Bounds PL, Dang CV. Otto Warburg's contributions to current concepts of cancer metabolism. Nat Rev Cancer. 2011;11(5):325–37. https://doi.org/10.1038/nrc3038.
Kalyanaraman B. Teaching the basics of cancer metabolism: Developing antitumor strategies by exploiting the differences between normal and cancer cell metabolism. Redox Biol. 2017;12:833–42. https://doi.org/10.1016/j.redox.2017.04.018.
Warburg O. On the origin of cancer cells. Science. 1956;123(3191):309–14. https://doi.org/10.1126/science.123.3191.309.
Cairns RA, Harris IS, Mak TW. Regulation of cancer cell metabolism. Nat Rev Cancer. 2011;11(2):85–95. https://doi.org/10.1038/nrc2981.
Zheng J. Energy metabolism of cancer: glycolysis versus oxidative phosphorylation (review). Oncol Lett. 2012;4(6):1151–7. https://doi.org/10.3892/ol.2012.928.
Pavlova NN, Thompson CB. The emerging hallmarks of cancer metabolism. Cell Metab. 2016;23(1):27–47. https://doi.org/10.1016/j.cmet.2015.12.006.
Yeung SJ, Pan J, Lee MH. Roles of p53, MYC and HIF-1 in regulating glycolysis: the seventh hallmark of cancer. Cell Mol Life Sci CMLS. 2008;65(24):3981–99. https://doi.org/10.1007/s00018-008-8224-x.
Courtnay R, Ngo DC, Malik N, Ververis K, Tortorella SM, Karagiannis TC. Cancer metabolism and the Warburg effect: the role of HIF-1 and PI3K. Mol Biol Rep. 2015;42(4):841–51. https://doi.org/10.1007/s11033-015-3858-x.
Danhier P, Banski P, Payen VL, Grasso D, Ippolito L, Sonveaux P, et al. Cancer metabolism in space and time: beyond the Warburg effect. Biochem Biophys Acta. 2017;1858(8):556–72. https://doi.org/10.1016/j.bbabio.2017.02.001.
Hagland H, Nikolaisen J, Hodneland LI, Gjertsen BT, Bruserud O, Tronstad KJ. Targeting mitochondria in the treatment of human cancer: a coordinated attack against cancer cell energy metabolism and signalling. Expert Opin Ther Targets. 2007;11(8):1055–69. https://doi.org/10.1517/14728222.11.8.1055.
Prada-Arismendy J, Arroyave JC, Rothlisberger S. Molecular biomarkers in acute myeloid leukemia. Blood Rev. 2017;31(1):63–766. https://doi.org/10.1016/j.blre.2016.08.005.
Chan SM, Majeti R. Role of DNMT3A, TET2, and IDH1/2 mutations in pre-leukemic stem cells in acute myeloid leukemia. Int J Hematol. 2013;98(6):648–57. https://doi.org/10.1007/s12185-013-1407-8.
Sun Y, Chen BR, Deshpande A. Epigenetic regulators in the development, maintenance, and therapeutic targeting of acute myeloid leukemia. Front Oncol. 2018;8:41. https://doi.org/10.3389/fonc.2018.00041.
Eriksson A, Lennartsson A, Lehmann S. Epigenetic aberrations in acute myeloid leukemia: early key events during leukemogenesis. Exp Hematol. 2015;43(8):609–24. https://doi.org/10.1016/j.exphem.2015.05.009.
Suganuma K, Miwa H, Imai N, Shikami M, Gotou M, Goto M, et al. Energy metabolism of leukemia cells: glycolysis versus oxidative phosphorylation. Leuk Lymphoma. 2010;51(11):2112–9. https://doi.org/10.3109/10428194.2010.512966.
Jorge J, Petronilho S, Alves R, Coucelo M, Goncalves AC, Nascimento Costa JM, et al. Apoptosis induction and cell cycle arrest of pladienolide B in erythroleukemia cell lines. Investig New Drugs. 2020;38(2):369–77. https://doi.org/10.1007/s10637-019-00796-2.
Brito AF, Abrantes AM, Ribeiro M, Oliveira R, Casalta-Lopes J, Goncalves AC, et al. Fluorine-18 fluorodeoxyglucose uptake in hepatocellular carcinoma: correlation with glucose transporters and p53 expression. J Clin Exp Hepatol. 2015;5(3):183–9. https://doi.org/10.1016/j.jceh.2015.05.003.
Miwa H, Shikami M, Goto M, Mizuno S, Takahashi M, Tsunekawa-Imai N, et al. Leukemia cells demonstrate a different metabolic perturbation provoked by 2-deoxyglucose. Oncol Rep. 2013;29(5):2053–7. https://doi.org/10.3892/or.2013.2299.
Goto M, Miwa H, Shikami M, Tsunekawa-Imai N, Suganuma K, Mizuno S, et al. Importance of glutamine metabolism in leukemia cells by energy production through TCA cycle and by redox homeostasis. Cancer Invest. 2014;32(6):241–7. https://doi.org/10.3109/07357907.2014.907419.
Liu H, Kurtoglu M, Cao Y, Xi H, Kumar R, Axten JM, et al. Conversion of 2-deoxyglucose-induced growth inhibition to cell death in normoxic tumor cells. Cancer Chemother Pharmacol. 2013;72(1):251–62. https://doi.org/10.1007/s00280-013-2193-y.
Xi H, Kurtoglu M, Liu H, Wangpaichitr M, You M, Liu X, et al. 2-Deoxy-d-glucose activates autophagy via endoplasmic reticulum stress rather than ATP depletion. Cancer Chemother Pharmacol. 2011;67(4):899–910. https://doi.org/10.1007/s00280-010-1391-0.
Xiong W, Jiao Y, Huang W, Ma M, Yu M, Cui Q, et al. Regulation of the cell cycle via mitochondrial gene expression and energy metabolism in HeLa cells. Acta Biochim Biophys Sin. 2012;44(4):347–58. https://doi.org/10.1093/abbs/gms006.
Kurtoglu M, Gao N, Shang J, Maher JC, Lehrman MA, Wangpaichitr M, et al. Under normoxia, 2-deoxy-d-glucose elicits cell death in select tumor types not by inhibition of glycolysis but by interfering with N-linked glycosylation. Mol Cancer Ther. 2007;6(11):3049–58. https://doi.org/10.1158/1535-7163.MCT-07-0310.
Quentmeier H, MacLeod RA, Zaborski M, Drexler HG. JAK2 V617F tyrosine kinase mutation in cell lines derived from myeloproliferative disorders. Leukemia. 2006;20(3):471–6. https://doi.org/10.1038/sj.leu.2404081.
Marty C, Lacout C, Droin N, Le Couedic JP, Ribrag V, Solary E, et al. A role for reactive oxygen species in JAK2 V617F myeloproliferative neoplasm progression. Leukemia. 2013;27(11):2187–95. https://doi.org/10.1038/leu.2013.102.
Seo M, Crochet RB, Lee Y-H. Targeting altered metabolism-emerging cancer therapeutic. Strategies. 2014;12:427–48. https://doi.org/10.1016/b978-0-12-396521-9.00014-0.
Zhang D, Li J, Wang F, Hu J, Wang S, Sun Y. 2-Deoxy-D-glucose targeting of glucose metabolism in cancer cells as a potential therapy. Cancer Lett. 2014;355(2):176–83. https://doi.org/10.1016/j.canlet.2014.09.003.
Mueckler M, Thorens B. The SLC2 (GLUT) family of membrane transporters. Mol Aspects Med. 2013;34(2–3):121–38. https://doi.org/10.1016/j.mam.2012.07.001.
Calvo MB, Figueroa A, Pulido EG, Campelo RG, Aparicio LA. Potential role of sugar transporters in cancer and their relationship with anticancer therapy. Int J Endocrinol. 2010. https://doi.org/10.1155/2010/205357.
Nagle DG, Zhou Y-D. Natural products as probes of selected targets in tumor cell biology and hypoxic signaling. In: Liu H-W, Mander L, editors. Comprehensive natural products II. Oxford: Elsevier; 2010. p. 651–683.
Shchepina LA, Pletjushkina OY, Avetisyan AV, Bakeeva LE, Fetisova EK, Izyumov DS, et al. Oligomycin, inhibitor of the F0 part of H+-ATP-synthase, suppresses the TNF-induced apoptosis. Oncogene. 2002;21(53):8149–57. https://doi.org/10.1038/sj.onc.1206053.
Comelli M, Di Pancrazio F, Mavelli I. Apoptosis is induced by decline of mitochondrial ATP synthesis in erythroleukemia cells. Free Radic Biol Med. 2003;34(9):1190–9. https://doi.org/10.1016/S0891-5849(03)00107-2.
Gemin A, Sweet S, Preston TJ, Singh G. Regulation of the cell cycle in response to inhibition of mitochondrial generated energy. Biochem Biophys Res Commun. 2005;332(4):1122–32. https://doi.org/10.1016/j.bbrc.2005.05.061.
Ho PA, Kopecky KJ, Alonzo TA, Gerbing RB, Miller KL, Kuhn J, et al. Prognostic implications of the IDH1 synonymous SNP rs11554137 in pediatric and adult AML: a report from the Children's Oncology Group and SWOG. Blood. 2011;118(17):4561–6. https://doi.org/10.1182/blood-2011-04-348888.
Xu Q, Li Y, Lv N, Jing Y, Xu Y, Li W, et al. Correlation between isocitrate dehydrogenase gene aberrations and prognosis of patients with acute myeloid leukemia: a systematic review and meta-analysis. Clin Cancer Res. 2017;23(15):4511–22. https://doi.org/10.1158/1078-0432.CCR-16-2628.
Willander K, Falk IJ, Chaireti R, Paul E, Hermansson M, Green H, et al. Mutations in the isocitrate dehydrogenase 2 gene and IDH1 SNP 105C>T have a prognostic value in acute myeloid leukemia. Biomark Res. 2014;2:18. https://doi.org/10.1186/2050-7771-2-18.
Acknowledgements
The present work was supported by Center of Investigation on Environment, Genetics and Oncobiology (CIMAGO), Faculty of Medicine, University of Coimbra, Portugal and by National Funds via Foundation for Science and Technology (FCT) through the Strategic Project UID/NEU/04539/2019, COMPETE-FEDER (POCI-01-0145-FEDER-007440), UIDB/04539/2020, and UIDP/04539/2020 (CIBB). RA was supported by FCT trough a PhD Grant (SFRH/BD/51994/2012).
Funding
The present work was supported by the Center of Investigation on Environment, Genetics and Oncobiology (CIMAGO), Faculty of Medicine, University of Coimbra, Portugal and by National Funds via Foundation for Science and Technology (FCT) through the Strategic Project UID/NEU/04539/2019, COMPETE-FEDER (POCI-01-0145-FEDER-007440), UIDB/04539/2020, and UIDP/04539/2020 (CIBB). RA was supported by the Portuguese Foundation to Science and Technology (FCT) through a PhD Grant (SFRH/BD/51994/2012).
Author information
Authors and Affiliations
Contributions
ACG and ABSR designed the experiments. BL and ACG drafted the manuscript. BL, ACG, RA, JJ, ASP, AMA, AA, MFB, and MC performed or interpreted the experiments. BL, JJ, and RA executed the statistical analyses. JMNC and ABSR revised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interests.
Ethical approval
This research project was approved by the Institutional Review Board of the Medical School of the University of Coimbra (reference: 072-CE-2017).
Consent to participate
Not applicable.
Consent for publication
The authors declare that they consent to the manuscript publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lapa, B., Gonçalves, A.C., Jorge, J. et al. Acute myeloid leukemia sensitivity to metabolic inhibitors: glycolysis showed to be a better therapeutic target. Med Oncol 37, 72 (2020). https://doi.org/10.1007/s12032-020-01394-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-020-01394-6