Abstract
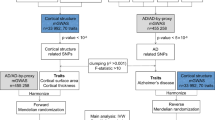
The disproportionate cortical atrophy is an established biomarker for the pathophysiological process of Alzheimer’s disease (AD). However, the genetic basis underlying the cortical atrophy remains poorly defined. Herein, we aim to illustrate the effect of the Wnt target genes on the cortical volumes of AD patients. 82 sporadic AD patients were recruited. All the subjects had history survey, blood biochemical examination, cognitive assessment, MRI morphometry and whole exome sequencing. This report focused on 84 common variants (minor allele frequency > 0.01) of 32 Wnt target genes, including the APC, DAAM1, DACT1, DISC1, LATS2, TLR2, WDR61, and the AXIN, DVL, FZD, LRP, TCF/LEF, WNT family genes. The Wnt target genes showed asymmetric effects on the cortical volumes of AD patients. The right temporal/parietal/occipital cortices were more affected than left temporal/parietal/occipital cortices. Nevertheless, the reverse applied to the frontal cortex. The DACT1 affected the cortical thickness most, followed by the TCF3 and APC. The DACT1 rs698025-GG genotype displayed greater right temporal pole and left medial orbito-frontal gyrus than rs698025-GA genotype (2.4 ± 0.4 vs. 2.0 ± 0.6, P = 0.005; 5.2 ± 0.6 vs. 5.0 ± 0.6, P = 0.001). The brain region most influenced by the Wnt target genes was the right calcarine cortex. In conclusion, the common variants of the Wnt target genes exert asymmetric effects on the cortical volumes of AD patients. The Wnt signaling pathway may play a role in the cortical atrophy of AD patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alzheimer's disease (AD) is pathologically characterized by amyloid-beta (Aβ) accumulation and phosphorylated tau deposition. The disproportionate cortical atrophy on structural magnetic resonance imaging (MRI) is an established biomarker for the pathophysiological process of AD (McKhann et al. 2011). The cortical atrophy typically occurs early in the medial temporal lobe and extends along a temporal–parietal–frontal trajectory (Pini et al. 2016). However, the genetic architecture underlying the cortical atrophy remains poorly defined.
The APOE-ɛ4 is the strongest genetic risk factor for AD. With a systematic review and meta-analysis of 14 cross-sectional studies, Liu et al., concluded that the APOE-ɛ4 is primarily relevant to hippocampal volume (Liu et al. 2015). With the data from Alzheimer’s Disease Neuroimaging Initiative and Allen Human Brain Atlas, Acosta et al., suggested that the APOE-ɛ4 did not contribute to the regional atrophy of AD (Acosta et al. 2018). Anyway, more than half of AD patients do not harbor the APOE-ɛ4 (Ward et al. 2012). Therefore, more research is required to explore the genetic basis of cortical atrophy in AD.
In recent years, the Wnt signaling pathway is supposed to play a role in AD pathogenesis. It can prevent the amyloidogenic processing of APP by inhibiting the BACE1 transcription. It can suppress the tau phosphorylation by decreasing the GSK3β activity, which is a kinase related to tau phosphorylation. Besides, the Wnt signaling pathway facilitates neuronal survival and neurogenesis, promotes synaptic plasticity, modulates microglial activity and neuroinflammation, enhances the integrity and function of blood–brain barrier (Inestrosa and Varela-Nallar 2014; Jia et al. 2019). Therefore, the inhibition of Wnt signaling pathway can lead to Aβ42 accumulation, tau phosphorylation, and eventually the occurrence of AD.
In 2020, Grasby et al., deployed a genome-wide association study (GWAS) of brain imaging data from 51,665 individuals. They found that the genetic loci affecting the cortical structures clustered near the genes which were related to the Wnt signaling pathway, such as the DACT1, DAAM1 and WNT2B (Grasby et al. 2020).
We hypothesize that the Wnt target genes exert effects on the cortical thickness of AD patients. In this study, firstly, we will identify the Wnt target genes with literature review, Gene Expression Omnibus (GEO) and Uniprot database analysis. Secondly, we will investigate the effect of Wnt target genes on the cortical volumes in AD patients. Ultimately, we expect to better understand the role of Wnt signaling pathway in AD.
Method
Participants and Inclusion Criteria
82 unrelated participants were recruited. The inclusion criteria were as follows: 1. Intact data on history survey, blood biochemical examination, cognitive assessment, brain MRI morphometry and whole exome sequencing. 2. Right handedness. 3. Diagnosed with probable AD based on the 2011 diagnostic criteria from National Institute on Aging and the Alzheimer’s Association (McKhann et al. 2011). 4. Sporadic cases with no dementia-causing variants were detected. 5. Written informed consent was acquired.
Gene Sequencing and Wnt Target Genes
The Genomic DNA were obtained from the peripheral blood. The fragmented DNA were captured by the Agilent SureSelect Human All ExonV6 Kit (Agilent Technologies, Santa Clara, CA, USA) and sequenced on the Illumina Novaseq 6000 platform (Illumina Inc., San Diego, CA, USA).
This report focused on the common variants of the Wnt target genes. The common variants met the following criteria: 1. Minor allele frequency > 0.01 in 1000genomes, GnomAD, ESP6500, ExAC dababases; 2. Located in the exons or adjacent introns (within 10 bp of splicing junction).
The identification of the Wnt target genes were performed with literature review, GEO and Uniprot database analysis. Three GEO datasets were downloaded, including GSE132903, GSE33000 and GSE63061 (https://www.ncbi.nlm.nih.gov/gds/). The differentially expressed genes (DEGs) between AD and control groups were achieved by GEO2R online tool, with adjusted P < 0.01 as the cutoff value. The overlapping DEGs from the three datasets were identified by Venn diagram. According to the uniprot database (https://www.uniprot.org), the overlapping DEGs involved in the Wnt signaling pathway were included in the study.
MRI Morphometry and Target Cortical Regions
MRI scan was performed on the 3.0 Tesla scanner (Discovery MR750, GE) with an 8-channel head coil. The structural images were acquired by sagittal T1-weighted three-dimensional fast spoiled gradient echo sequence (repetition time = 8.2 ms, echo time = 3.2 ms, flip angle = 12°, Prep Time = 400 ms, matrix = 256 × 256). Morphometry analysis was performed by SPM 12 techniques. According to the tutorial of voxel-based morphometry, the original images were segmented into gray matter, white matter and cerebrospinal fluid. The segmentation images were normalized to Montreal Neurological Institute space. The normalized gray matter images were smoothed with an 8mm3 full width at half maximum filter.
This report focused on 56 cortical regions, including 22 temporal (bilateral hippocampus, parahippocampus, entorhinal cortex, insula, fusiform, superior/middle/inferior/transverse temporal gyrus, bank of superior temporal sulcus, temporal pole), 16 frontal (bilateral superior frontal gyrus, anterior/posterior middle frontal gyrus, frontal pole, orbital gyrus, medial/lateral orbitofrontal gyrus, precentral gyrus), 10 parietal (bilateral postcentral, supramarginal gyrus, superior/inferior parietal lobule, precuneus), and 8 occipital areas (bilateral cuneus, lingual, calcarine, lateral occipital cortex).
Statistics
As for each single nucleotide polymorphism (SNP), the MRI morphometry data were compared by analysis of covariate and post hoc Bonferroni correction. Age, gender, disease course, APOE status (ε4 carriers or non-carriers) and whole brain volume were included in the model as fixed factors or covariates.
Result
Demographic Characteristics
As shown in Table 1, 36.6% (30/82) participants were males and 63.4% (52/82) were females. The average age was 73.6 ± 9.6 years old. The average disease course was 3.4 ± 2.4 years. The APOE-ε4 allele frequency in the cohort was 0.26. 58.5% (48/82) cases were APOE-ε4 non-carriers, 31.7% (26/82) were ε4 heterozygotes, and 9.8% (8/82) were ε4 homozygotes. All the participants were right handedness.
Wnt Target Genes and SNPs
Literature review indicated that several genes were involved in the Wnt signaling pathway, including the APC, CTNNB1, DAAM1, DACT1, GSK3B, LRP5-6, as well as the AXIN, DVL, FZD, TCF/LEF and WNT family genes (Cheyette et al. 2002; Jia et al. 2019; Li et al. 2005; Liu et al. 2008; Tapia-Rojas et al. 2016; Zhang et al. 2006).
The GSE132903, GSE33000 and GSE63061 were RNA expression profiles from middle temporal gyrus, prefrontal cortex and peripheral blood, respectively. There were 9762 DEGs between AD and control groups in GSE132903, 14,628 DEGs in GSE33000, and 1001 DEGs in GSE63061. A total of 431 overlapping DEGs from the three datasets were identified (Fig. 1; Supplement Table S1). According to the uniprot database, six genes were implicated in the Wnt signaling pathway, including the CSNK1G2, DISC1, LATS2, PFDN5, TLR2 and WDR61 (https://www.uniprot.org).
Volcano plots and Venn diagram of GSE132903, GSE33000, GSE63061. The first three volcano plots were from GSE132903, GSE33000, GSE63061, respectively. The differentially expressed genes between AD and control groups were achieved by GEO2R online tool, with adjusted P < 0.01 as the cutoff value. The fourth was the Venn diagram for the overlapping differentially expressed genes of the three datasets
This report focused on the common variants of the above Wnt target genes. Some genes were excluded since no common variants were available in the cohort, such as the CTNNB1, GSK3B, CSNK1G2, PFDN5, etc. Finally, 84 SNPs of 32 Wnt target genes were included in this study. The 32 Wnt genes included the APC, AXIN1, AXIN2, DAAM1, DACT1, DISC1, DVL2, FZD1, FZD3, FZD6, FZD8, FZD10, LATS2, LEF1, LRP5, LRP6, TCF3, TCF4, TCF7, TCF7L2, TCF12, TLR2, WDR61, WNT2B, WNT7A, WNT8A, WNT8B, WNT9A, WNT9B, WNT10B, WNT11 and WNT16. The 84 Wnt target SNPs were shown in Table 2.
Wnt SNP Frequency
Table 2 demonstrated the alternate allele frequencies of the 84 SNPs in the cohort. Compared with the East Asian population from GnomAD database, the AD patients in this study showed lower allele frequencies in three TCF3 SNPs, including rs1052692 (0.09 vs. 0.23), rs1052696 (0.10 vs. 0.23) and rs2074888 (0.27 vs.0.41). The allele frequencies of the remaining 81 SNPs in this AD group were close to those of the East Asian population (Table 2; Supplement Table S2).
Effect of Wnt SNPs on MRI Morphometry
Among the 84 Wnt target SNPs, 45.2% (38/84) SNPs showed effects on cortical volumes (Fig. 2; Supplement Table S3). The raw values were shown below (mm3).
Effect of Wnt target SNPs on cortical volumes of AD patients. This illustrated the effect of 38 Wnt target SNPs on 56 cortical regions of AD patients. Each square represents the specific effect of the SNP of the horizontal axis on the cortex of the vertical axis. The different colors of these squares represent different P-values. P < 0.05; P < 0.01, the subjects with the alternate alleles show smaller cortical volumes than those with reference alleles. P < 0.05; P < 0.01, the subjects with the alternate alleles show greater cortical volumes than those with reference alleles. P ≥ 0.05. The remaining 46 Wnt SNPs with no effects on cortical thickness were not shown in this figure
Among the APC and AXIN SNPs, the 351,771-A allele was related to smaller right calcarine cortex (GG vs. GA, AA: 2.7 ± 0.7 vs. 2.3 ± 0.5, 2.1 ± 0.4, P = 0.004). So were the alternate alleles of rs41115, rs42427, rs866006 and rs465899. The rs459552-TA genotype showed greater right superior parietal lobule than rs459552-AA (11.4 ± 1.4 vs. 10.6 ± 1.5, P = 0.004). The rs9915936-G allele was relevant to greater right parahippocampus, insula and superior temporal gyrus (AA vs. AG, GG: 2.7 ± 0.5 vs. 3.4 ± 0.7, 3.4 ± 0.6, P = 0.001; 4.8 ± 0.6 vs. 6.1 ± 0.8, 6.0 ± 0.8, P < 0.001; 8.1 ± 2.4 vs. 10.1 ± 1.4, 10.2 ± 1.3, P = 0.009). The rs2240307-C allele was associated with greater right orbital gyrus (TT vs. TC, CC: 2.4 ± 0.4 vs. 2.6 ± 0.4, 2.7 ± 0.4, P = 0.001).
Among the DACT1, DISC1 and LAST2 SNPs, the rs698025-GG genotype displayed greater right temporal pole and left medial orbito-frontal gyrus than rs698025-GA genotype (2.4 ± 0.4 vs. 2.0 ± 0.6, P = 0.005; 5.2 ± 0.6 vs. 5.0 ± 0.6, P = 0.001). So was the rs863091-CC genotype relative to rs863091-CT. Also, the rs7317471-AG genotype exhibited greater left medial orbito-frontal gyrus relative to rs7317471-GG (5.4 ± 0.6 vs. 5.1 ± 0.6, P = 0.005). In addition, the rs3738401-G allele was associated with greater left middle temporal gyrus (GG, GA vs. AA: 9.4 ± 1.5, 9.3 ± 1.5 vs. 7.1 ± 1.9, P = 0.005). The rs59928188-GG genotype showed smaller left temporal pole than rs59928188-GA (2.4 ± 0.3 vs. 2.7 ± 0.4, P = 0.005).
Among the LRP and TCF SNPs, the rs2277268-GG phenotype showed smaller left superior frontal gyrus than rs2277268-GA (19.0 ± 2.4 vs. 20.4 ± 2.9, P = 0.005). So was the rs4988322-TT genotype relative to rs4988322-TC. The rs2302685-GA and rs2240590-CT genotypes exhibited smaller left insula than rs2302685-AA and rs2240590-CC, respectively (5.5 ± 0.7 vs. 5.9 ± 0.8, P = 0.004; 5.5 ± 0.7 vs. 5.9 ± 0.7, P = 0.003).
Among the WNT SNPs, the rs2908004-G allele correlated with greater right lateral orbitofrontal gyrus (GG, GA vs. AA: 6.5 ± 0.9, 6.3 ± 0.7 vs. 6.0 ± 0.7, P = 0.003). So was the rs2707466-C allele. The rs2908004-GG genotype had smaller right cuneus than rs2908004-GA (2.7 ± 0.4 vs. 2.8 ± 0.5, P = 0.002). So was the rs2707466-CC genotype relative to rs2707466-CT. The rs3762719-T allele was relevant to smaller right bank of superior temporal sulcus (TT, TC vs. CC: 1.4 ± 0.4, 1.4 ± 0.3 vs. 1.7 ± 0.3, P = 0.002). Also, the rs12639607-GA exhibited smaller right bank of superior temporal sulcus than rs12639607-AA (1.4 ± 0.3 vs. 1.7 ± 0.3, P = 0.003). Besides, the rs4968281-TT genotype displayed smaller left superior frontal gyrus relative to rs4968281-TC (18.4 ± 2.2 vs. 20.1 ± 2.4, P = 0.002).
Effect of APOE-ε4 on MRI Morphometry
As illustrated in Fig. 3, the APOE-ε4 was associated with greater frontal cortical volumes. The raw values were shown below (mm3). Compared with the APOE-ε4 non-carriers, the ε4 heterozygotes showed greater left lateral orbitofrontal gyrus (6.5 ± 0.9 vs. 6.2 ± 0.7, P = 0.014) and left precentral gyrus (13.1 ± 1.5 vs. 12.4 ± 1.5, P = 0.041). The ε4 homozygotes exhibited greater right superior frontal gyrus than ε4 non-carriers (20.8 ± 4.3 vs. 17.9 ± 2.6, P = 0.014).
The post hoc analysis with LSD correction demonstrated the smaller right hippocampus in ε4 homozygotes relative to ε4 non-carriers (3.0 ± 0.7 vs. 3.5 ± 0.7, P = 0.030). However, no statistical significance was found after Bonferroni correction.
Discussion
This report focuses on the common variants of 32 Wnt target genes. Of them, the APC, AXIN, DVL, FZD, LRP, TCF/LEF and WNT families encode the vital elements of the Wnt signaling pathway. During the pathway, the WNT binds to LRP5/6 and FZD, which activates DVL and inhibits GSK-3β, AXIN, APC. Subsequently, the phosphorylation of ß-catenin is blocked, which contributes to the formation of TCF/LEF complex, and then the activation of the pathway (Jia et al. 2019; Li et al. 2005; Tapia-Rojas et al. 2016). The DAAM1, DACT1, DISC1, LATS2, TLR2 and WDR61 encode the regulators of the pathway (www.uniprot.org). The DAAM1 mediates DVL complex formation (Liu et al. 2008). The DACT1 interacts with DVL and regulates the degradation of β-catenin (Cheyette et al. 2002), (Zhang et al. 2006). The DISC1 modulates GSK-3β activity and CTNNB1 abundance (Mao et al. 2009). The LATS2 inhibits the β-catenin/BCL9 interaction (Li et al. 2013). The TLR2 participates in the WNT5A expression (Blumenthal et al. 2006). The WDR61 is the component of the PAF1 complex, which is required for the transcription of Wnt target genes (www.uniprot.org).
With the data from 51,665 individuals, Grasby et al., revealed that the loci affecting the cortical structures clustered near the genes related to the Wnt signaling pathway (Grasby et al. 2020). It is the first time that we describes the association between the Wnt target genes and cortical volumes in AD patients. In this study, 45.2% (38/84) Wnt target SNPs show effects on the cortical thickness of AD patients. We believe that the Wnt target genes affect the cortical thickness in both general population and AD patients.
Moreover, the Wnt target SNPs exert asymmetric effects on bilateral cortices of AD patients. The right temporal/parietal/occipital cortices are more affected than left temporal/parietal/occipital cortices. Nevertheless, the reverse applies to the frontal cortex. It is possibly owing to the asymmetric expression of Wnt target genes in the cortex. Miao et al., discovered that the predicted targets associated with the Wnt signaling pathway exhibited differential expression between the two hemispheres, such as the WNT, FZD and AXIN2 (Miao et al. 2020). With a thorough review, Hüsken et al., concluded that the Wnt signaling pathway played a vital role in the establishment of brain asymmetry and laterality (Husken and Carl 2013).
In this study, the DACT1 affects the cortical thickness most, followed by the TCF3 and APC. The two SNPs with the most effects on the cortical volumes are rs698025 (DACT1) and rs863091 (DACT1). Grasby et al., also observed that the loci near the DACT1 affected the cortical volume (Grasby et al. 2020). These findings emphasize the correlation between the DACT1 gene and cortical thickness. Therefore, we advise to focus on the DACT1 gene while exploring the genetic basis of cortical atrophy in AD patients.
The APC was identified as a novel AD susceptibility gene from a recent GWAS study (Prokopenko et al. 2021). However, no studies elucidated the relevance of the APC gene for AD phenotype. Herein, we find the APC gene is closely related to right posterior cortex. Five APC SNPs correlate with right calcarine cortex, and one SNP is relevant to right superior parietal lobule.
The brain region most influenced by the Wnt target genes is the right calcarine cortex. Eight SNPs show effects on the right calcarine cortex. This is different from the APOE-ε4, which is mainly associated with smaller hippocampal volume (Liu et al. 2015). In this report, we additionally find that the APOE-ε4 is related to greater frontal volume. This is consistent with two previous studies, which implies the region-specific effects of APOE-ε4 on the cortical atrophy of AD (Cacciaglia et al. 2018; Ten et al. 2016). We suppose that the Wnt target genes and APOE affect brain cortices of AD patients with different patterns.
The AD patients in this study show lower alternate allele frequencies in three TCF3 SNPs (rs1052692, rs1052696, rs2074888) relative to the East Asian population. This may be due to the ethnic differences. The allele frequencies of these SNPs in Chinese population and AD patients could be further investigated.
In conclusion, the common variants of the Wnt target genes exert asymmetric effects on the cortical volumes of AD patients. The Wnt signaling pathway may play a role in the disproportionate cortical atrophy of AD patients. The main limitation of this report is the small sample size. The future research could focus on the DACT1, APC and TCF3 genes with an expanded sample size.
Data Availability
The original contributions are included in the article, further dataset are available from the corresponding author on reasonable request.
References
Acosta D, Powell F, Zhao Y, Raj A (2018) Regional vulnerability in Alzheimer’s disease: The role of cell-autonomous and transneuronal processes. Alzheimers Dement 14(6):797–810. https://doi.org/10.1016/j.jalz.2017.11.014
Blumenthal A, Ehlers S, Lauber J, Buer J, Lange C, Goldmann T et al (2006) The Wingless homolog WNT5A and its receptor Frizzled-5 regulate inflammatory responses of human mononuclear cells induced by microbial stimulation. Blood 108(3):965–973. https://doi.org/10.1182/blood-2005-12-5046
Cacciaglia R, Molinuevo JL, Falcon C, Brugulat-Serrat A, Sanchez-Benavides G, Gramunt N et al (2018) Effects of APOE-epsilon4 allele load on brain morphology in a cohort of middle-aged healthy individuals with enriched genetic risk for Alzheimer’s disease. Alzheimers Dement 14(7):902–912. https://doi.org/10.1016/j.jalz.2018.01.016
Cheyette BN, Waxman JS, Miller JR, Takemaru K, Sheldahl LC, Khlebtsova N et al (2002) Dapper, a Dishevelled-associated antagonist of beta-catenin and JNK signaling, is required for notochord formation. Dev Cell 2(4):449–461. https://doi.org/10.1016/s1534-5807(02)00140-5
Grasby KL, Jahanshad N, Painter JN, Colodro-Conde L, Bralten J, Hibar DP et al (2020) The genetic architecture of the human cerebral cortex. Science 367(6484):eaay6690. https://doi.org/10.1126/science.aay6690
Husken U, Carl M (2013) The wnt/beta-catenin signaling pathway establishes neuroanatomical asymmetries and their laterality. Mech Develop 130(6–8):330–335. https://doi.org/10.1016/j.mod.2012.09.002
Inestrosa NC, Varela-Nallar L (2014) Wnt signaling in the nervous system and in alzheimer’s disease. J Mol Cell Biol 6(1):64–74. https://doi.org/10.1093/jmcb/mjt051
Jia L, Pina-Crespo J, Li Y (2019) Restoring wnt/beta-catenin signaling is a promising therapeutic strategy for alzheimer’s disease. Mol Brain 12(1):104. https://doi.org/10.1186/s13041-019-0525-5
Li F, Chong ZZ, Maiese K (2005) Vital elements of the wnt-frizzled signaling pathway in the nervous system. Curr Neurovasc Res 2(4):331–340. https://doi.org/10.2174/156720205774322557
Li J, Chen X, Ding X, Cheng Y, Zhao B, Lai ZC et al (2013) LATS2 suppresses oncogenic Wnt signaling by disrupting beta-catenin/BCL9 interaction. Cell Rep 5:1650–1663. https://doi.org/10.1016/j.celrep.2013.11.037
Liu W, Sato A, Khadka D, Bharti R, Diaz H, Runnels LW et al (2008) Mechanism of activation of the formin protein daam1. P Natl Acad Sci USA 105(1):210–215. https://doi.org/10.1073/pnas.0707277105
Liu Y, Yu JT, Wang HF, Han PR, Tan CC, Wang C et al (2015) Apoe genotype and neuroimaging markers of alzheimer’s disease: systematic review and meta-analysis. J Neurol Neurosur Ps 86(2):127–134. https://doi.org/10.1136/jnnp-2014-307719
Mao Y, Ge X, Frank CL, Madison JM, Koehler AN, Doud MK et al (2009) Disrupted in schizophrenia 1 regulates neuronal progenitor proliferation via modulation of gsk3beta/beta-catenin signaling. Cell 136(6):1017–1031. https://doi.org/10.1016/j.cell.2008.12.044
McKhann GM, Knopman DS, Chertkow H, Hyman BT, Jack CJ, Kawas CH et al (2011) The diagnosis of dementia due to alzheimer’s disease: recommendations from the national institute on aging-alzheimer’s association workgroups on diagnostic guidelines for alzheimer’s disease. Alzheimers Dement 7(3):263–269. https://doi.org/10.1016/j.jalz.2011.03.005
Miao N, Lai X, Zeng Z, Cai W, Chen W, Sun T (2020) Differential expression of micrornas in the human fetal left and right cerebral cortex. Mol Biol Rep 47(9):6573–6586. https://doi.org/10.1007/s11033-020-05708-9
Pini L, Pievani M, Bocchetta M, Altomare D, Bosco P, Cavedo E et al (2016) Brain atrophy in alzheimer’s disease and aging. Ageing Res Rev 30:25–48. https://doi.org/10.1016/j.arr.2016.01.002
Prokopenko D, Morgan SL, Mullin K, Hofmann O, Chapman B, Kirchner R et al (2021) Whole-genome sequencing reveals new alzheimer’s disease-associated rare variants in loci related to synaptic function and neuronal development. Alzheimers Dement. https://doi.org/10.1002/alz.12319
Tapia-Rojas C, Burgos PV, Inestrosa NC (2016) Inhibition of wnt signaling induces amyloidogenic processing of amyloid precursor protein and the production and aggregation of amyloid-beta (abeta)42 peptides. J Neurochem 139(6):1175–1191. https://doi.org/10.1111/jnc.13873
Ten KM, Sanz-Arigita EJ, Tijms BM, Wink AM, Clerigue M, Garcia-Sebastian M et al (2016) Impact of apoe-varepsilon4 and family history of dementia on gray matter atrophy in cognitively healthy middle-aged adults. Neurobiol Aging 38:14–20. https://doi.org/10.1016/j.neurobiolaging.2015.10.018
Ward A, Crean S, Mercaldi CJ, Collins JM, Boyd D, Cook MN et al (2012) Prevalence of apolipoprotein e4 genotype and homozygotes (apoe e4/4) among patients diagnosed with alzheimer’s disease: a systematic review and meta-analysis. Neuroepidemiology 38(1):1–17. https://doi.org/10.1159/000334607
Zhang L, Gao X, Wen J, Ning Y, Chen YG (2006) Dapper 1 antagonizes wnt signaling by promoting dishevelled degradation. J Biol Chem 281(13):8607–8612. https://doi.org/10.1074/jbc.M600274200
Funding
Dr. Jing Gao was supported by grants from National Key Research and Development Program of China (No. 2020YFA0804500 and 2016YFC1306300), CAMS Innovation fund for Medical Sciences (No. 2021–1-I2M-1–020), National Natural Science Foundation of China (No. 81550021 and 30470618), and the strategic priority research program (pilot study) “Biological basis of aging and therapeutic strategies” of the Chinese Academy of Sciences (No. XDPB10).
Author information
Authors and Affiliations
Contributions
LD and BH contributed to acquisition, analysis, interpretation of the data, and draft of the work. JG and FF contributed to acquisition, analysis, interpretation of the data, and conception, revision of the work. CL, CM, XH, LS, SC, BP and LC contributed to acquisition of the data. All authors approved the submitted version.
Corresponding authors
Ethics declarations
Ethical Approval
This study was approved by ethics committee of PUMCH (No. JS-1836). Written informed consent was obtained.
Conflict of Interest
The authors have no conflict of interest to report.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dong, L., Hou, B., Liu, C. et al. Association Between Wnt Target Genes and Cortical Volumes in Alzheimer’s Disease. J Mol Neurosci 73, 1010–1016 (2023). https://doi.org/10.1007/s12031-023-02122-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-023-02122-1