Abstract
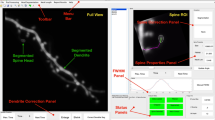
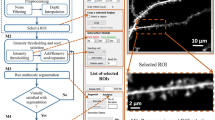
The rapid development of microscopic imaging techniques has greatly facilitated time-lapse imaging of neuronal morphology. However, analysis of structural dynamics in the vast amount of 4-Dimensional data generated by in vivo or ex vivo time-lapse imaging still relies heavily on manual comparison, which is not only laborious, but also introduces errors and discrepancies between individual researchers and greatly limits the research pace. Here we present a supervised 4D Structural Plasticity Analysis (4D SPA) computer method to align and match 3-Dimensional neuronal structures across different time points on a semi-automated basis. We demonstrate 2 applications of the method to analyze time-lapse data showing gross morphological changes in dendritic arbor morphology and to identify the distribution and types of branch dynamics seen in a series of time-lapse images. Analysis of the dynamic changes of neuronal structure can be done much faster and with greatly improved consistency and reliability with the 4D SPA supervised computer program. Users can format the neuronal reconstruction data to be used for this analysis. We provide file converters for Neurolucida and Imaris users. The program and user manual are publically accessible and operate through a graphical user interface on Windows and Mac OSX.




Similar content being viewed by others
References
Al-Kofahi, O., Radke, R. J., et al. (2006). Automated semantic analysis of changes in image sequences of neurons in culture. IEEE Transactions on Biomedical Engineering, 53(6), 1109–1123.
Antonini, A., & Stryker, M. P. (1993). Rapid remodeling of axonal arbors in the visual cortex. Science, 260(5115), 1819–1821.
Besl, P. J., & MaKay, N. D. (1992). A Method for Registration of 3-D Shapes. IEEE Transactions PAMI, 14(2), 239–256.
Bestman, J. E., & Cline, H. T. (2008). The RNA binding protein CPEB regulates dendrite morphogenesis and neuronal circuit assembly in vivo. Proceedings of the National Academy of Sciences of the United States of America, 105(51), 20494–20499.
Bestman, J. E., & Cline, H. T. (2009). The relationship between dendritic branch dynamics and CPEB-labeled RNP granules captured in vivo. Front Neural Circuits, 3, 10.
Bestman, J. E., Lee-Osbourne, J., et al. (2012). In vivo time-lapse imaging of cell proliferation and differentiation in the optic tectum of Xenopus laevis tadpoles. The Journal of Comparative Neurology, 520(2), 401–433.
Cantallops, I., Haas, K., et al. (2000). Postsynaptic CPG15 promotes synaptic maturation and presynaptic axon arbor elaboration in vivo. Nature Neuroscience, 3(10), 1004–1011.
Carmeliet, P. (2003). Angiogenesis in health and disease. Nature Medicine, 9(6), 653–660.
Chen, J. L., Lin, W. C., et al. (2011). Structural basis for the role of inhibition in facilitating adult brain plasticity. Nature Neuroscience, 14(5), 587–594.
Chiu, S. L., Chen, C. M., et al. (2008). Insulin receptor signaling regulates synapse number, dendritic plasticity, and circuit function in vivo. Neuron, 58(5), 708–719.
Cline, H. T. (1999). Dendrite development in dendrites. London: Oxford Univ. Press.
Cohen-Cory, S. (2002). The developing synapse: construction and modulation of synaptic structures and circuits. Science, 298(5594), 770–776.
De Paola, V., Holtmaat, A., et al. (2006). Cell type-specific structural plasticity of axonal branches and boutons in the adult neocortex. Neuron, 49(6), 861–875.
Donte, D., Foggia, P., et al. (2004). Thirty years of graph matching in pattern recognition. IJPRAI, 18, 265–298.
Dumay, A. C. M., R. J. Geest, et al. (1992). “Consistent inexact graph matching applied to labeling coronary segments in arteriograms.” Proc.Int. Conf. Pattern Recognition Conf.: 439–442.
Haas, K., Li, J., et al. (2006). AMPA receptors regulate experience-dependent dendritic arbor growth in vivo. Proceedings of the National Academy of Sciences of the United States of America, 103(32), 12127–12131.
Halavi, M., Hamilton, K. A., et al. (2012). Digital reconstructions of neuronal morphology: three decades of research trends. Frontiers in Neuroscience, 6, 49.
Haris, K., Efstratiadis, S. N., et al. (1999). Model-based morphological segmentation and labeling of coronary angiograms. IEEE Transactions on Medical Imaging, 18(10), 1003–1015.
He, H. Y., & Cline, H. T. (2011). Diadem X: automated 4 dimensional analysis of morphological data. Neuroinformatics, 9(2–3), 107–112.
Helmchen, F., & Denk, W. (2005). Deep tissue two-photon microscopy. Nature Methods, 2(12), 932–940.
Hochman, D. W. (2012). The extracellular space and epileptic activity in the adult brain: explaining the antiepileptic effects of furosemide and bumetanide. Epilepsia, 53(Suppl 1), 18–25.
Lu, S. M., Tremblay, M. E., et al. (2011). HIV-1 Tat-induced microgliosis and synaptic damage via interactions between peripheral and central myeloid cells. PloS One, 6(9), e23915.
Reh, T. A., & Constantine-Paton, M. (1985). Eye-specific segregation requires neural activity in three-eyed Rana pipiens. Journal of Neuroscience, 5(5), 1132–1143.
Ruthazer, E. S., Akerman, C. J., et al. (2003). Control of axon branch dynamics by correlated activity in vivo. Science, 301(5629), 66–70.
Thal, D. R., L. T. Grinberg, et al. (2012). “Vascular dementia: Different forms of vessel disorders contribute to the development of dementia in the elderly brain.” Experimental Gerontology.
Tremblay, M., Fugere, V., et al. (2009). Regulation of radial glial motility by visual experience. Journal of Neuroscience, 29(45), 14066–14076.
Tremblay, M. E., Lowery, R. L., et al. (2010). Microglial interactions with synapses are modulated by visual experience. PLoS Biology, 8(11), e1000527.
Turney, S. G., & Lichtman, J. W. (2008). Chapter 11: imaging fluorescent mice in vivo using confocal microscopy. Methods in Cell Biology, 89, 309–327.
Wilt, B. A., Burns, L. D., et al. (2009). Advances in light microscopy for neuroscience. Annual Review of Neuroscience, 32, 435–506.
Zhang, S., Boyd, J., et al. (2005). Rapid reversible changes in dendritic spine structure in vivo gated by the degree of ischemia. Journal of Neuroscience, 25(22), 5333–5338.
Zudaire, E., Gambardella, L., et al. (2011). A computational tool for quantitative analysis of vascular networks. PloS One, 6(11), e27385.
Acknowledgments
This work was funded by support from the National Institutes of Health (EY011261), the Hahn Family Foundation and the Nancy Lurie Marks Family Foundation to HTC, and the National Science Council, Taiwan, Grants 98-2221-E-009-118-MY3 and 101-2221-E-009-143-MY3 to Y-TC. We thank Dr. Shu-Ling Chiu for fostering this productive collaboration. P-CL would like to thank Dr. M. Giugliano for his help implementing the file converter.
Conflicts of Interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Ping-Chang Lee and Hai-yan He contributed equally.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
(AVI 46835 kb)
(MPG 3628 kb)
(MPG 20586 kb)
(MPG 12874 kb)
Rights and permissions
About this article
Cite this article
Lee, PC., He, Hy., Lin, CY. et al. Computer Aided Alignment and Quantitative 4D Structural Plasticity Analysis of Neurons. Neuroinform 11, 249–257 (2013). https://doi.org/10.1007/s12021-013-9179-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12021-013-9179-0