Abstract
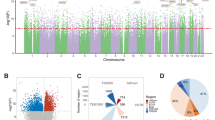
Schizophrenia (SCZ) is a complex psychiatric disease with a lifetime morbidity rate of 0.5–1.0 %. To date, aberrant DNA methylation in SCZ has been reported in several studies. However, no comprehensive studies using medication-free subjects with SCZ have been conducted. In addition, most of these studies have been limited to the analysis of the CpG sites in CpG islands (CGIs) in the gene promoter regions, so little is known about the DNA methylation signatures across the whole genome in SCZ. Genome-wide DNA methylation profiling (485,764 CpG sites) of peripheral leukocytes was conducted in the first set of samples (24 medication-free patients with SCZ and 23 non-psychiatric controls) using Infinium HumanMethylation450 Beadchips. Second, a monozygotic twin study was performed using three pairs of monozygotic twins that were discordant for SCZ. Finally, the data from these two independent cohorts were compared. A total of 234 differentially methylated CpG sites that were common between these two cohorts were identified. Of the 234 CpG sites, 153 sites (65.4 %) were located in the CGIs and in the regions flanking CGIs (CGI: 40.6 %; CGI shore: 13.3 %; CGI shelf: 11.5 %). Of the 95 differently methylated CpG sites in the CGIs, most of them were located in the promoter regions (promoter: 75.8 %; gene body: 14.7 %; 3′-UTR: 2.1 %). Aberrant DNA methylation in SCZ was identified at numerous loci across the whole genome in peripheral leukocytes using two independent sets of samples. These findings support the notion that altered DNA methylation could be involved in the pathophysiology of SCZ.


Similar content being viewed by others
References
Abdolmaleky, H. M., Cheng, K. H., Faraone, S. V., Wilcox, M., Glatt, S. J., Gao, F., et al. (2006). Hypomethylation of MB-COMT promoter is a major risk factor for schizophrenia and bipolar disorder. Human Molecular Genetics, 15, 3132–3145.
Ball, M. P., Li, J. B., Gao, Y., Lee, J., LeProust, E. M., Park, I., et al. (2009). Targeted and genome-scale strategies reveal gene-body methylation signatures in human cells. Nature Biotechnology, 27, 361–368.
Bhugra, D. (2005). The global prevalence of schizophrenia. PLoS Medicine, 2, e151–e175.
Bhutani, N., Burns, D. M., & Blau, H. M. (2011). DNA demethylation dynamics. Cell, 146, 866–872.
Bibikova, M., Barnes, B., Tsan, C., Ho, V., Klotzle, B., Le, J. M., et al. (2011). High density DNA methylation array with single CpG site resolution. Genomics, 98, 288–295.
Cardno, A. G., & Gottesman, I. I. (2000). Twin studies of schizophrenia: From bow-and-arrowconcordances to star wars Mx and functional genomics. American Journal of Medical Genetics, 97, 12–17.
Carrard, A., Salzmann, A., Malafosse, A., & Karege, F. (2011). Increased DNA methylation status of the serotonin receptor 5HTR1A gene promoter in schizophrenia and bipolar disorder. Journal of Affective Disorders, 132, 450–453.
Chen, Y., Zhang, J., Zhang, L., Shen, Y., Xu, Q. (2011). Effects of MAOA promoter methylation on susceptibility to paranoid schizophrenia. Human Genetics [Epub ahead of print].
Colantuoni, C., Lipska, B. K., Ye, T., Hyde, T. M., Tao, R., Leek, J. T., et al. (2011). Temporal dynamics and genetic control of transcription in the human prefrontal cortex. Nature, 478, 519–523.
Datta, S. R., McQuillin, A., Rizig, M., Blaveri, E., Thirumalai, S., Kalsi, G., et al. (2010). A threonine to isoleucine missense mutation in the pericentriolar material 1 gene is strongly associated with schizophrenia. Molecular Psychiatry, 15, 615–628.
Deaton, A. M., Webb, S., Kerr, A. R. W., Illingworth, R. S., Guy, J., Andrews, R., et al. (2011). Cell type-specific DNA methylation at intragenic CpG islands in the immune system. Genome Research, 21, 1074–1086.
Dedeurwaerder, S., Defrance, M., Calonne, E., & Sotiriou, C. (2011). Evaluation of the Infinium 450 K technology. Epigenomics, 3, 771–784.
Dempster, E. L., Pidsley, R., Schalkwyk, L. C., Owens, S., Georgiades, A., Kane, F., et al. (2011). Disease-associated epigenetic changes in monozygotic twins discordant for schizophrenia and bipolar disorder. Human Molecular Genetics, 20, 4786–4796.
Dong, E., Nelson, M., Grayson, D. R., Costa, E., & Guidotti, A. (2008). Clozapine and sulpiride but not haloperidol or olanzapine activate brain DNA demethylation. Proceedings of National Academy of Sciences of the United States of America, 105, 13614–13619.
Gervin, K., Vigeland, M. D., Mattingsdal, M., Hammerø, M., Nygård, H., Olsen, A. O., et al. (2012). DNA methylation and gene expression changes in monozygotic twins discordant for psoriasis: Identification of epigenetically dysregulated genes. PLoS Genetics, 8, e1002454.
Grayson, D. R., Jia, X., Chen, Y., Sharma, R. P., Mitchell, C. P., Guidotti, A., et al. (2005). Reelin promoter hypermethylation in schizophrenia. Proceedings of National Academy of Sciences of the United States of America, 10, 9341–9346.
Gurling, H. M., Critchley, H., Datta, S. R., McQuillin, A., Blaveri, E., Thirumalai, S., et al. (2006). Genetic association and brain morphology studies and the chromosome 8p22 pericentriolar material 1 (PCM1) gene in susceptibility to schizophrenia. Archives of General Psychiatry, 63, 844–854.
Harrison, P. J., & Weinberger, D. R. (2005). Schizophrenia genes, gene expression, and neuropathology: On the matter of their convergence. Molecular Psychiatry, 10, 40–68.
Illingworth, R. S., Gruenewald-Schneider, U., Webb, S., Kerr, A. R., James, K. D., Turner, D. J., et al. (2010). Orphan CpG islands identify numerous conserved promoters in the mammalian genome. PLoS Genetics, 6, e1001134.
International Schizophrenia Consortium. (2008). Rare chromosomal deletions and duplications increase risk of schizophrenia. Nature, 455, 237–241.
Irizarry, R. A., Ladd-Acosta, C., Wen, B., Wu, Z., Montano, C., Onyango, P., et al. (2009). The human colon cancer methylome shows similar hypo- and hypermethylation at conserved tissue-specific CpG island shores. Nature Genetics, 41, 178–186.
Iwamoto, K., Bundo, M., Yamada, K., Takao, H., Iwayama-Shigeno, Y., Yoshikawa, T., et al. (2005). DNA methylation status of SOX10 correlates with its downregulation and oligodendrocyte dysfunction in schizophrenia. The Journal of Neuroscience, 25, 5376–5381.
Javierre, B. M., Fernandez, A. F., Richter, J., Al-Shahrour, F., Martin-Subero, J. I., Rodriguez-Ubreva, J., et al. (2010). Changes in the pattern of DNA methylation associate with twin discordance in systemic lupus erythematosus. Genome Research, 20, 170–179.
Kähler, A. K., Djurovic, S., Rimol, L. M., Brown, A. A., Athanasiu, L., Jönsson, E. G., et al. (2011). Candidate gene analysis of the human natural killer-1 carbohydrate pathway and perineuronal nets in schizophrenia: B3GAT2 is associated with disease risk and cortical surface area. Biological Psychiatry, 69, 90–96.
Kaminsky, Z., Tochigi, M., Jia, P., Pal, M., Mill, J., Kwan, A., et al. (2012). A multi-tissue analysis identifies HLA complex group 9 gene methylation differences in bipolar disorder. Molecular Psychiatry, 17(7), 728–740.
Kim, T., Park, J. K., Kim, H. J., Chung, J. H., & Kim, J. W. (2010). Association of histone deacetylase genes with schizophrenia in Korean population. Psychiatry Research, 178, 266–269.
Kuratomi, G., Iwamoto, K., Bundo, M., Kusumi, I., Kato, N., Iwata, N., et al. (2008). Aberrant DNA methylation associated with bipolar disorder identified from discordant monozygotic twins. Molecular Psychiatry, 13, 429–441.
Lavedan, C., Licamele, L., Volpi, S., Hamilton, J., Heaton, C., Mack, K., et al. (2009). Association of the NPAS3 gene and five other loci with response to the antipsychotic iloperidone identified in a whole genome association study. Molecular Psychiatry, 14, 804–819.
Leek, J. T., & Storey, J. D. (2007). Capturing heterogeneity in gene expression studies by surrogate variable analysis. PLoS Genetics, 3, 1724–1735.
Lett, T. A., Wallance, T. J. M., Chowdhur, N. I., Tiwari, A. K., Kennedy, J. L., & Muller, D. J. (2011). Pharmacogenetics of antipsychotic-induced weight gain: Review and clinical implications. Molecular Psychiatry, 17, 242–266.
Maunakea, A. K., Nagarajan, R. P., Bilenky, M., Ballinger, T. J., D’souza, C., Fouse, S. D., et al. (2010). Conserved role of intragenic DNA mehtylaion in regulating alternative promoters. Nature, 466, 253–257.
Melas, P. A., Rogdaki, M., Osby, U., Challing, M., Lavebratt, C., & Ekstrom, T. J. (2012). Epigenetic aberrations in leukocytes of patients with schizophrenia: Association of global DNA methylation with antipsychotic drug treatment and disease onset. The FASEB Journal, 26, 2712–2718.
Mill, J., Tang, T., Kaminsky, Z., Khare, T., Yazdanpanah, S., Bouchard, L., et al. (2008). Epigenomic profiling reveals DNA-methylation changes associated with major psychosis. American Journal of Human Genetics, 82, 696–711.
Moskvina, V., Craddock, P., Nikolov, I., Pahwa, J. S., Green, E., Wellcome Trust Case Control Consortium, et al. (2009). Gene-wide analysis of genome-wide association data sets: Evidence for multiple common risk alleles for schizophrenia and bipolar disorder and for overlap in genetic risk. Molecular Psychiatry, 14, 252–260.
Nohesara, S., Ghadirivasfi, M., Mostafavi, S., Eskandari, M. R., Ahmadkhaniha, H., Thiagalingam, S., et al. (2011). DNA hypomethylation of MB-COMT promoter in the DNA derived from saliva in schizophrenia and bipolar disorder. Journal of Psychiatic Research, 45, 1432–1438.
Numata, S., Ye, T., Hyde, T. M., Guitart-Navarro, X., Tao, R., Wininger, M., et al. (2012). DNA methylation signatures in development and aging of the human prefrontal cortex. American Journal of Human Genetics, 90, 260–272.
Ono, S., Imamura, A., Tasaki, S., Kurotaki, N., Ozawa, H., Yoshiura, K., et al. (2010). Failure to confirm CNVs as of aetiological significance in twin pairs discordant for schizophrenia. Twin Research and Human Genetics, 13, 455–460.
Petronis, A., Gottesman, I. I., Crow, T. J., DeLisi, L. E., Klar, A. J., Macciardi, F., et al. (2000). Psychiatric epigenetics: A new focus for the new century. Molecular Psychiatry, 5, 34234–34236.
Purcell, S. M., Wray, N. R., Stone, J. L., Visscher, P. M., O’Donovan, M. C., Sullivan, P. F., et al. (2009). Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature, 460, 748–752.
Rakyan, V. K., Beyan, H., Down, T., Hawa, M. I., Maslau, S., Aden, D., et al. (2011). Identification of type 1 diabetes–associated DNA methylation variable positions that precede disease diagnosis. PLoS Genetics, 7, e1002300.
Rees, E., Moskvina, V., Owen, M. J., O’Donovan, M. C., & Kirov, G. (2011). De novo rates and selection of schizophrenia-associated copy number variations. Biological Psychiatry, 70, 1109–1114.
Sandoval, J., Heyn, H., Moran, S., Serra-Musach, J., Pujana, M. A., Bibikova, M., et al. (2011). Validation of a DNA methylation microarray for 450,000 CpG sites in the human genome. Epigenetics, 6, 692–702.
Shann, Y. J., Cheng, C., Chiao, C. H., Chen, D. T., Li, P. H., & Hsu, M. T. (2008). Genome-wide mapping and characterization of hypomethylated sites in human tissues and breast cancer cell lines. Genome Research, 18, 791–801.
Shi, J., Levinson, D. F., Duan, J., Sanders, A. R., Zheng, Y., Pe’er, I., et al. (2009). Common variants on chromosome 6p22.1 are associated with schizophrenia. Nature, 460, 753–757.
Shimabukuro, M., Jinno, Y., Fuke, C., & Okazaki, Y. (2006). Haloperidol treatment induces tissue- and sex-specific changes in DHA methylation: A control study using rats. Behavioral and Brain Functions, 2, 37.
Siegmund, K. D., Connor, C. M., Campan, M., Long, T. I., Weisenberger, D. J., Biniszkiewicz, D., et al. (2007). DNA methylation in the human cerebral cortex is dynamically regulated throughout the life span and involves differentiated neurons. PLoS ONE, 2(9), e895.
Souza, R. P., de Luca, V., Remington, G., Lieberman, J. A., Meltzer, H. Y., Kennedy, J. L., et al. (2010a). Glial cell line-derived neurotrophic factor alpha 2 (GFRA2) gene is associated with tardive dyskinesia. Psychopharmacology (Berl), 210, 347–354.
Souza, R. P., Romano-Silva, M. A., Lieberman, J. A., Meltzer, H. Y., MacNeil, L. T., Culotti, J. G., et al. (2010b). Genetic association of the GDNF alpha-receptor genes with schizophrenia and clozapine response. Journal of Psychiatry Research, 44, 700–706.
Stefansson, H., Ophoff, R. A., Steinberg, S., Andreassen, O. A., Cichon, S., Rujescu, D., et al. (2009). Common variants conferring risk of schizophrenia. Nature, 460, 744–747.
Sullivan, P. F., Kendler, K. S., & Neale, M. C. (2003). Schizophrenia as a complex trait: Evidence from a meta-analysis of twin studies. Archives of General Psychiatry, 60, 1187–1192.
Tremolizzo, L., Doueiri, M. S., Dong, E., Grayson, D. R., Davis, J., Pinna, G., et al. (2005). Valproate corrects the schizophrenia-like epigenetic behavioral modifications induced by methionine in mice. Biological Psychiatry, 57, 500–550.
Acknowledgments
The authors would like to thank all the volunteers who understood our study purpose and participated in this study, and the physicians who helped us to collect clinical data and blood samples at the mental hospitals. The authors would also like to thank Mrs. Akemi Okada for her technical assistance. This work was supported by Japan Science and Technology Agency, CREST and by a Grants-in-Aid for Scientific Research from the Japanese Ministry of Education, Culture, Sports, Science and Technology. The all authors report no biomedical financial interests or potential conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Makoto Kinoshita and Shusuke Numata contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kinoshita, M., Numata, S., Tajima, A. et al. DNA Methylation Signatures of Peripheral Leukocytes in Schizophrenia. Neuromol Med 15, 95–101 (2013). https://doi.org/10.1007/s12017-012-8198-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12017-012-8198-6