Abstract
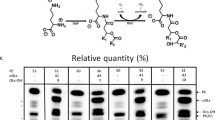
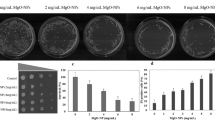
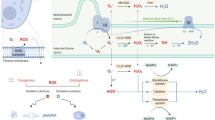
Cadmium (Cd) is a potent toxic element used in several industries and in the process contaminates air, soil, and water. Exposure of Saccharomyces cerevisiae to Cd increases the major phospholipids, and profound increase was observed in phosphatidylethanolamine (PE). In yeast, there are four different pathways contributing to the biosynthesis of PE, and contribution to PE pool through phosphatidylserine decarboxylase2 (psd2) is not significant in normal conditions. Upon Cd exposure, psd2Δ strain showed a significant decrease in major phospholipids including PE. When exposed to Cd, wild-type (WT) cells depicted an increase in ER stress and autophagy, whereas in psd2, ER stress was noted but autophagy process was impaired. The supplementation of ethanolamine did not overcome the Cd stress and also the autophagy process, whereas overexpression of PSD2 in psd2Δ increased the cellular tolerance, PE levels, and the autophagy process against Cd stress. From our studies, we can suggest that PSD2 of S. cerevisiae has an important role in PE synthesis and in autophagy process under Cd stress.








Similar content being viewed by others
References
Klionsky, D. J. (2005). The molecular machinery of autophagy: Unanswered questions. Journal of Cell Science, 118, 7–18.
Yorimitsu, T., Nair, U., Yang, Z., & Klionsky, D. J. (2006). Endoplasmic reticulum stress triggers autophagy. Journal of Biological Chemistry, 281, 30299–30304.
Chrestensen, C. A., Starke, D. W., & Mieyal, J. J. (2000). Acute cadmium exposure inactivates thiol transferase (Glutaredoxin), inhibits intracellular reduction of protein glutathionyl mixed disulfides, and initiates apoptosis. Journal of Biological Chemistry, 275, 26556–26565.
Stohs, S. J., & Bagchi, D. (1995). Oxidative mechanisms in the toxicity of metal ions. Free Radical Biology & Medicine, 18, 321–336.
Faller, P., Kienzler, K., & Krieger-Liszkay, A. (2005). Mechanism of Cd2+ toxicity: Cd2+ inhibits photo activation of Photosystem II by competitive binding to the essential Ca2+ site. Biochimica et Biophysica Acta, 1706, 158–164.
Nargund, A. M., Avery, S. V., & Houghton, J. E. (2008). Cadmium induces a heterogeneous and caspase dependent apoptotic response in Saccharomyces cerevisiae. Apoptosis, 13, 811–821.
Yokouchi, M., Hiramatsu, N., Hayakawa, K., Kasai, A., Takano, Y., Yao, J., et al. (2007). A typical, bidirectional regulation of cadmium-induced apoptosis via distinct signaling of unfolded protein response. Cell Death and Differentiation, 14, 1467–1474.
Biagioli, M., Pifferi, S., Ragghianti, M., Bucci, S., Rizzuto, R., & Pinton, P. (2008). Endoplasmic reticulum stress and alteration in calcium homeostasis are involved in cadmium induced apoptosis. Cell Calcium, 43, 184–195.
Muthukumar, K., & Nachiappan, V. (2010). Cadmium-induced oxidative stress in Saccharomyces cerevisiae. Indian Journal of Biochemistry & Biophysics, 47, 383–387.
Howlett, N. G., & Avery, S. V. (1997). Induction of lipid peroxidation during heavy metal stress in Saccharomyces cerevisiae and influence of plasma membrane fatty acid unsaturation. Applied and Environment Microbiology, 63, 2971–2976.
Girotti, W. (1985). Mechanisms of lipid peroxidation. Free Radical Biology & Medicine, 1, 87–95.
Vance, J. E. (2008). Phosphatidylserine and phosphatidylethanolamine in mammalian cells: Two metabolically related amino phospholipids. Journal of Lipid Research, 49, 1377–1387.
Trotter, P. J., Pedretti, J., & Voelker, D. R. (1993). Phosphatidylserine decarboxylase from Saccharomyces cerevisiae. Isolation of mutants, cloning of the gene, and creation of a null allele. Journal of Biological Chemistry, 268(28), 21416–21424.
Trotter, P. J., & Voelker, D. R. (1995). Identification of a non-mitochondrial phosphatidylserine decarboxylase activity in the yeast Saccharomyces cerevisiae. Journal of Biological Chemistry, 270, 6062–6070.
Kennedy, E. P., & Weiss, S. B. (1956). The function of cytidine coenzymes in the biosynthesis of phospholipids. Journal of Biological Chemistry, 222, 193–214.
Riekhof, W. R., Wu, J., Jones, J. L., & Voelker, D. R. (2007). Identification and characterization of the major lysophosphatidylethanolamine acyltransferase in Saccharomyces cerevisiae. Journal of Biological Chemistry, 282, 28344–28352.
Gulshan, K., Shahi, P., & Scott Moye-Rowley, W. (2010). Compartment-specific synthesis of phosphatidylethanolamine is required for normal heavy metal resistance. Molecular Biology of the Cell, 21, 443–455.
Yoshimori, T. (2004). Autophagy: A regulated bulk degradation process inside cells. Biochemical and Biophysical Research Communications, 313, 453–458.
Carman, G. M., & Han, G. S. (2009). Regulation of phospholipid biosynthesis in yeast. Journal of Lipid Research, 50, 69–73.
Iwanyshyn, W. M., Han, G. S., & Carman, G. M. (2004). Regulation of phospholipid synthesis in Saccharomyces cerevisiae by zinc. Journal of Biological Chemistry, 279, 21976–21983.
Ghosh, A. K., Ramakrishnan, G., & Rajasekharan, R. (2008). YLR099C (ICT1) encodes a soluble acyl CoA dependent lysophosphatidic acid acyltransferase responsible for enhanced phospholipid synthesis on organic solvent stress in Saccharomyces cerevisiae. Journal of Biological Chemistry, 283, 9768–9775.
Bligh, E. G., & Dyer, W. J. (1959). A rapid method of total lipid extraction and purification. Canadian Journal of Biochemistry and Physiology, 37, 911–917.
Wagner, S., & Paltauf, F. (1994). Generation of glycerophospholipid molecular species in the yeast Saccharomyces cerevisiae. Fatty acid pattern of phospholipid classes and selective acyl turnover at sn-1 and sn-2 positions. Yeast, 10, 1429–1437.
Siakotos, A. N., Rouser, G., & Fleischer, S. (1966). Phospholipid composition of human, bovine and frog myelin isolated on a large scale from brain and spinal cord. Lipids, 1, 85–86.
Gietz, R. D., & Schiestl, R. H. (1995). Transforming Yeast with DNA. Methods in Molecular and Cellular Biology, 5, 255–269.
Kirisako, T., Ichimura, Y., Okada, H., Kabeya, Y., Mizushima, N., Yoshimori, T., et al. (2000). The reversible modification regulates the membrane-binding state of Apg8/Aut7 essential for autophagy and the cytoplasm to vacuole targeting pathway. Journal of Cell Biology, 151, 263–276.
Yen, W. L., Legakis, J. E., Nair, U., & Klionsky, D. J. (2007). Atg27 is required for autophagy dependent cycling of Atg9. Molecular Biology of the Cell, 18, 581–593.
Normington, K., Kohno, K., Kozutsumi, Y., Gething, M. J., & Sambrook, J. (1989). S. cerevisiae encodes an essential protein homologous in sequence and function to mammalian BiP. Cell, 57, 1223–1236.
Nebauer, R., Rosenberger, S., & Daum, G. (2007). Phosphatidylethanolamine, a limiting factor of autophagy in yeast strains bearing a defect in the carboxypeptidase Y pathway of vacuolar targeting. Journal of Biological Chemistry, 282, 16736–16743.
Storey, M. K., Clay, K. L., Kutateladze, T., Murphy, R. C., Overduin, M., & Voelker, D. R. (2001). Phosphatidylethanolamine has an essential role in Saccharomyces cerevisiae that is independent of its ability to form hexagonal phase structures. Journal of Biological Chemistry, 276, 48539–48548.
Muthukumar, K., Rajakumar, S., Sarkar, M. N., & Nachiappan, V. (2011). Glutathione peroxidase3 of Saccharomyces cerevisiae protects phospholipids during cadmium-induced oxidative stress. Antonie van Leeuwenhoek, 99, 761–771.
Vijayaraj, P., Sabarirajan, J., & Nachiappan, V. (2011). Enhanced phospholipase B activity and alteration of phospholipids and neutral lipids in Saccharomyces cerevisiae exposed to N nitrosonornicotine. Antonie van Leeuwenhoek, 99, 567–577.
Ron, D., & Walter, P. (2007). Signal integration in the endoplasmic reticulum unfolded protein response. Nature Reviews Molecular Cell Biology, 8, 519–529.
Schuck, S., Prinz, W. A., Thorn, K. S., Voss, C., & Walter, P. (2009). Membrane expansion alleviates endoplasmic reticulum stress independently of the unfolded protein response. Journal of Cell Biology, 187, 525–536.
Miura, S., Zou, W., Ueda, M., & Tanaka, A. (2000). Screening of genes involved in isooctane tolerance in Saccharomyces cerevisiae by using mRNA differential display. Applied and Environment Microbiology, 66, 4883–4889.
Luo, J., Matsuo, Y., Gulis, G., Hinz, H., Patton-Vogt, J., & Marcus, S. (2009). Phosphatidylethanolamine is required for normal cell morphology and cytokinesis in the fission yeast Schizosaccharomyces pombe. Eukaryotic Cell, 8, 790–799.
Bogdanov, M., Umeda, M., & Dowhan, W. (1999). Phospholipid-assisted refolding of an integral membrane protein. Minimum structural features for phosphatidylethanolamine to act as a molecular chaperone. Journal of Biological Chemistry, 274, 12339–12345.
Opekarova, M., Robl, I., & Tanner, W. (2002). Phosphatidylethanolamine is essential for targeting the arginine transporter Can1p to the plasma membrane of yeast. Biochimica et Biophysica Acta, 1564, 9–13.
Birner, R., Nebauer, R., Schneiter, R., & Daum, G. (2003). Synthetic lethal interaction of the mitochondrial phosphatidylethanolamine biosynthetic machinery with the prohibitin complex of Saccharomyces cerevisiae. Molecular Biology of the Cell, 14, 370–383.
Gardarin, A., Chédin, S., Lagniel, G., Aude, J. C., Godat, E., Catty, P., et al. (2010). Endoplasmic reticulum is a major target of cadmium toxicity in yeast. Molecular Microbiology, 76, 1034–1048.
Cox, J. S., Shamu, C. E., & Walter, P. (1993). Transcriptional induction of genes encoding endoplasmic reticulum resident proteins requires a transmembrane protein kinase. Cell, 73, 1197–1206.
Mori, K., Ma, W., Gething, M. J., & Sambrook, J. (1993). A transmembrane protein with a cdc2/CDC28-related kinase activity is required for signaling from the ER to the nucleus. Cell, 74, 743–756.
Kimata, Y., Kimata, Y. I., Shimizu, Y., Abe, H., Farcasanu, I. C., Takeuchi, M., et al. (2003). Genetic evidence for a role of BiP/Kar2 that regulates Ire1 in response to accumulation of unfolded proteins. Molecular Biology of the Cell, 14, 2559–2569.
Fagone, P., & Jackowski, S. (2009). Membrane phospholipid synthesis and endoplasmic reticulum function. Journal of Lipid Research, Suppl, S311–S316.
Xie, Z., Nair, U., & Klionsky, D. J. (2008). Atg8 controls phagophore expansion during autophagosome formation. Molecular Biology of the Cell, 19, 3290–3298.
Nakatogawa, H., Ichimura, Y., & Ohsumi, Y. (2007). Atg8, a ubiquitin-like protein required for autophagosome formation, mediates membrane tethering and hemifusion. Cell, 130, 165–178.
Klionsky, D. J., Cueva, R., & Yaver, D. S. (1992). Aminopeptidase I of Saccharomyces cerevisiae is localized to the vacuole independent of the secretory pathway. Journal of Cell Biology, 119, 287–299.
Nakatogawa, H., Ohbayashi, S., Nakatogawa, M. S., Kakuta, S., Suzuki, S. W., Kirisako, H., et al. (2012). The autophagy-related protein kinase Atg1 interacts with the ubiquitin-like protein Atg8 via the Atg8 family interacting motif to facilitate autophagosome formation. Journal of Biological Chemistry, 287, 28503–28507.
Tsukada, M., & Ohsumi, Y. (1993). Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Letters, 333, 169–174.
Acknowledgments
The financial support from Bharathidasan University, Tiruchirappalli, Council of Scientific & Industrial Research—CSIR, India and instrumentation facility provided by Department of Science & Technology—DST under DST-PURSE programme are gratefully acknowledged. The authors also thank Prof. Dennis Voelker (National Jewish Health, Denver, CO, USA) for providing the YEp352-PSD2 plasmids, Prof. Gunther Daum, (Graz University, Austria) for providing mutant strains for this work, Prof. Daniel J. Klionsky (University of Michigan, Michigan, USA), Prof. Yoshinori Ohsumi, Prof. Kuninori Suzuki (Tokyo Institute of Technology, Yokohama, Japan), Prof. Jeffrey Brodsky (University of Pittsburgh, PA, USA), and Prof. Marco Foiani, (University of Milan, Italy) for providing Ape1, Atg8, kar2, and pgk1 antiserum, respectively.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Muthukumar, K., Nachiappan, V. Phosphatidylethanolamine from Phosphatidylserine Decarboxylase2 is Essential for Autophagy Under Cadmium Stress in Saccharomyces cerevisiae . Cell Biochem Biophys 67, 1353–1363 (2013). https://doi.org/10.1007/s12013-013-9667-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-013-9667-8