Abstract
The biochemical and structural characterization of ubiquitin-conjugating enzymes (E2s) over the past 30 years has fostered important insights into ubiquitin transfer mechanisms. Although many of these enzymes share high sequence and structural conservation, their functional roles in the cell are decidedly diverse. Here, we report that the mono-ubiquitinating E2 UBE2W forms a homodimer using two distinct protein surfaces. Dimerization is primarily driven by residues in the ß-sheet region and Loops 4 and 7 of the catalytic domain. Mutation of two residues in the catalytic domain of UBE2W is capable of disrupting UBE2W homodimer formation, however, we find that dimerization of this E2 is not required for its ubiquitin transfer activity. In addition, residues in the C-terminal region, although not compulsory for the dimerization of UBE2W, play an ancillary role in the dimer interface. In all current E2 structures, the C-terminal helix of the UBC domain is at least 15Å away from the primary dimerization surface shown here for UBE2W. This leads to the proposal that the C-terminal region of UBE2W adopts a noncanonical position that places it closer to the UBC ß-sheet, providing the first indication that at least some E2s adopt C-terminal conformations different from the canonical structures observed to date.






Similar content being viewed by others
Notes
E2s are organized into classes: Class I E2s contain only the ~150-residue catalytic domain (“UBC” domain); Class II, III, and IV E2s also contain N-terminal, C-terminal extension, or both. Currently, little is known regarding the function of such extensions and there is little structural information available on any extensions. All available E2 structures, regardless of the class they belong to, are of the UBC domain.
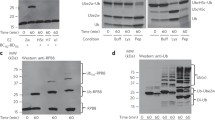
Of the ~120 resonances in the UBE2W-‘KK’ 1H, 15N-HSQC spectrum, we were able to unambiguously assign 115. From this dataset, we were able to transfer assignments to the ‘WT’ spectrum and identify 80 of the ~90 resonances present.
References
Pickart, C. M. (2001). Mechanisms underlying ubiquitination. Annual Review of Biochemistry, 2001(70), 195–201.
Ye, Y., & Rape, M. (2009). Building ubiquitin chains: E2 enzymes at work. Nature Reviews Molecular Cell Biology, 10, 755–764.
Christensen, D. E., Brzovic, P. S., & Klevit, R. E. (2007). E2-BRCA1 RING interactions dictate synthesis of mono- or specific polyubiquitin chain linkages. Nature Structural and Molecular Biology, 14, 941–948.
Rodrigo-Brenni, M. C., Foster, S. A., & Morgan, D. O. (2010). Catalysis of lysine 48-specific ubiquitin chain assembly by residues in E2 and ubiquitin. Molecular Cell, 39, 548–559.
Alpi, A. F., Pace, P. E., Babu, M. M., & Patel, K. J. (2008). Mechanistic insight into site-restricted monoubiquitination of FANCD2 by Ube2t, FANCL, and FANCI. Molecular Cell, 32, 767–777.
Scaglione, K. M., Zavodszky, E., Todi, S. V., Patury, S., Xu, P., Rodríguez-Lebrón, E., et al. (2011). Ube2w and Ataxin-3 Coordinately Regulate the Ubiquitin Ligase CHIP. Molecular Cell, 43, 599–612.
Pickart, C. M., & Raasi, S. (2005). Controlled synthesis of polyubiquitin chains. Methods in Enyzmology, 399, 21–36.
Delaglio, F., Grzesiek, S., Vuister, G. W., Zhu, G., Pfeifer, J., & Bax, A. (1995). NMRPipe: A multidimensional spectral processing system based on UNIX pipes. Journal of Biomolecular NMR, 6, 277–293.
Johnson, B. A., & Blevins, R. A. (1994). NMR view: A computer program for the visualization and analysis of NMR data. Journal of Biomolecular NMR, 4, 603–614.
Kelley, L. A., & Sternberg, M. J. E. (2009). Protein structure prediction on the web: A case study using the Phyre server. Nature Protocols, 4, 363–371.
Juan, Y. C., Landry, M. C., Sanches, M., Vittal, V., Leung, C. C., Ceccarelli, D. F., et al. (2012). OTUB1 co-opts Lys48-linked ubiquitin recognition to suppress E2 enzyme function. Molecular Cell, 45, 384–397.
Sheng, Y., Hong, J.H., Doherty, R., Srikumar, T., Shloush, J., Avvakumov, G.V., et al. (2012). A human ubiquitin conjugating enzyme (E2)-HECT E3 ligase structure-function screen. Molecular Cell Proteomics. 2012. 11, 329–341.
Brzovic, P. S., Lissounov, A., Christensen, D. E., Hoyt, D. W., & Klevit, R. E. (2006). A UbcH5c/ubiquitin noncovalent complex is required for processive BRCA1-direct ubiquitination. Molecular Cell, 21, 873–880.
Sakata, E., Satoh, T., Yamamoto, S., Yamaguchi, Y., Yagi-Utsumi, M., Tanaka, K., et al. (2010). (2010) Crystal structure of UbcH5b ~ ubiquitin intermediate: Insight into the formation of the self-assembled E2 ~ Ub conjugates. Structure, 18, 138–147.
Page, R. C., Pruneda, J. N., Amick, J., Klevit, R. E., & Misra, S. (2012). Structural insights into the conformation and oligomerization of E2 ~ ubiquitin conjugates. Biochemistry, 51, 4175–4187.
Das, R., Mariano, J., Tsai, Y. C., Kalathur, R. C., Kostova, Z., Li, J., et al. (2009). Allosteric activation of E2-RING finger-mediate ubiquityation by a structurally defined specific E2-binding region of gp78. Molecular Cell, 34, 674–685.
Li, W., Tu, D., Li, L., Wollert, T., Ghirlando, R., Brunger, et al. (2009). Mechanistic insights into active site-associated polyubiquitination by the ubiquitin-conjugating enzyme Ube2g2. Proceedings of the National Academy of Sciences of the United States of America. 106, 3722–3727.
Hibbert, R. G., Huang, A., Boelens, R., & Sixma, T. K. (2011). E3 ligase Rad18 promotes monubiquitination rather than ubiquitin chain formation by E2 enzyme Rad6. Proceedings of the National Academy of Sciences of the United States of America, 108, 5590–5595.
VanDermark, A. P., Hofmann, R. M., Tsui, C., Pickart, C. M., & Wolberger, C. (2001). Molecular insights into polyubiquitin chain assembly: Crystal structure of the Mms2/Ubc13 heterodimer. Cell, 105, 711–720.
Moraes, T. F., Edwards, R. A., McKenna, S., Pastushok, L., Xiao, W., Glover, J. N., et al. (2001). Crystal structure of the human ubiquitin conjugating enzyme complex, hMms2-hUbc13. Natural Structural Biology, 8, 669–673.
Pickart, C. E., & Rose, I. A. (1985). Functional heterogeneity of ubiquitin carrier proteins. Journal of Biological Chemistry, 260, 1573–1581.
Silver, E. T., Gwozd, T. J., Ptak, C., Goebl, M., & Ellison, M. J. (1992). A chimeric ubiquitin conjugating enzyme that combins the cell cycle properties of CDC34 (UBC3) and the DNA repair properties of RAD6 (UBC2): Implications for the structure, function and evolution of the E2s. EMBO Journal, 11, 3091–3098.
Girod, P. A., & Vierstra, R. D. (1993). A major ubiquitin conjugation system in wheat germ extracts involves a 15-kDa ubiquitin-conjugating enzyme (E2) homologous to the years UBC4/UBC5 gene products. Journal of Biological Chemistry, 268, 955–960.
Ptak, C., Prendergast, J. A., Hodgins, R., Kay, C. M., Chau, V., & Ellison, M. J. (1994). Functional and physical characterization of the cell cycle ubiquitin-conjugating enzyme CDC34 (UBC3). Identification of a functional determinant within the tail that facilitates CDC34 self-association. Journal of Biological Chemistry, 269, 26539–26545.
Haldeman, M. T., Xia, G., Kasperek, E. M., & Pickart, C. M. (1997). Structure and function of ubiquitin conjugating enzyme E2–25K: The tail is a core-dependent activity element. Biochemistry, 36, 10526–10537.
Gazdoiu, S., Yamoah, K., Wu, K., Escalante, C. R., Tappin, I., Bermudez, V., et al. (2005). Proximity-induced activation of human Cdc34 through heterologous dimerization. Proceedings of the National Academy of Sciences of the United States of America, 102(2005), 15053–15058.
Varelas, X., Ptak, C., & Ellison, M. J. (2003). Cdc34 self-association is facilitated by ubiquitin thiolester formation and is required for its catalytic activity. Molecular and Cellular Biology, 23, 5388–5400.
Skowyra, D., Graig, K. L., Tyers, M., Elledge, S. J., & Harper, J. W. (1997). F-box proteins are receptors that recruit phosphorylated substrates to the SCF ubiquitin-ligase complex. Cell, 91, 209–219.
Deffenbaugh, A. E., Scaglione, K. M., Zhang, L., Moore, J. M., Buranda, T., Sklar, L. A., et al. (2003). Release of ubiquitin-charged Cdc34-S – Ub from the RING domain is essential for ubiquitination of the SCF(Cdc4)-bound substrate Sic1. Cell, 114, 611–622.
Acknowledgments
The authors acknowledge D. Christensen and C. Eakin for their initial observations on the UBE2W-KK mutant. This study was supported by the National Institute of General Medical Sciences Grants R01 GM088055 (R.E.K.). V. V. was supported in part by the Hurd Fellowship Fund and PHS NRSA 2T32 GM007270.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.


Rights and permissions
About this article
Cite this article
Vittal, V., Wenzel, D.M., Brzovic, P.S. et al. Biochemical and Structural Characterization of the Ubiquitin-Conjugating Enzyme UBE2W Reveals the Formation of a Noncovalent Homodimer. Cell Biochem Biophys 67, 103–110 (2013). https://doi.org/10.1007/s12013-013-9633-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-013-9633-5