Abstract
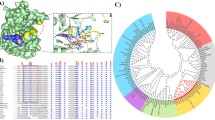
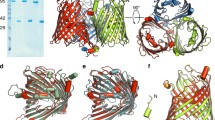
Vibrio harveyi chitinase A or VhChiA (EC.3.2.1.14) is a member of GH-18 chitinases that catalyzes chitin degradation from marine biomaterials. Our earlier structural data of VhChiA suggested that Tyr-435 marks the ending of subsite +2 and may influence binding of the interacting substrate at the aglycone binding sites. This study reports the effects of Tyr-435 using site-directed mutagenesis technique. Mutation of Tyr-435 to Ala (mutant Y435A) enhanced both binding and catalytic efficiency of VhChiA, whereas substitution of Tyr-435 to Trp (mutant Y435W) lessened the ability of the enzyme to bind and hydrolyze chitin substrates. The increased activity of Y435A can be explained by partial removal of a steric clash around subsite (+2), thereby allowing a chitin chain to move beyond or to access the enzyme’s active site from the aglycone side more straightforwardly.




Similar content being viewed by others
Abbreviations
- ChBD:
-
chitin binding domain
- DNS:
-
3,5-dinitrosalicylic acid
- GlcNAcn :
-
β-1-4 linked oligomers of N-acetyl-d-glucosamine residues, where n = 1–6
- IPTG:
-
isopropyl thio-β-d-galactoside
- PMSF:
-
phenylmethylsulphonylfluoride
- pNP:
-
p-nitrophenol
- VhChiA:
-
Vibrio harveyi chitinase A
- WT:
-
wild-type chitinase
References
Jayakumar, R., Prabaharan, M., Nair, S. V., & Tamura, H. (2010). Novel chitin and chitosan nanofibers in biomedical applications. Biotechnology Advances, 28, 142–150.
Ciardelli, G., & Chiono, V. (2006). Materials for peripheral nerve regeneration. Macromolecular Bioscience, 6, 13–26.
Yang, Y., Liu, M., Gu, Y., Lin, S., Ding, F., & Gu, X. (2009). Effect of chitooligosaccharide on neuronal differentiation of PC-12 cells. Cell Biology International, 33, 352–356.
You, Y., Park, W. H., Ko, B. M., & Min, B. M. (2004). Effects of PVA sponge containing chitooligosaccharide in the early stage of wound healing. Journal of Materials Science. Materials in Medicine, 15, 297–301.
Henrissat, B., & Bairoch, A. (1993). New families in the classification of glycosyl hydrolases based on amino-acid sequence similarities. Biochemistry Journal, 293, 781–788.
Henrissat, B., & Davies, G. (1997). Structural and sequence-based classification of glycosidehydrolases. Current Opinion in Structural Biology, 7, 637–644.
Suginta, W., Robertson, P. A., Austin, B., Fry, S. C., & Fothergill-Gilmore, L. A. (2000). Chitinases from Vibrios: activity screening and purification of chi A from Vibrio carchariae. Journal of Applied Microbiology, 89, 76–84.
Suginta, W. (2007). Identification of chitin binding proteins and characterization of two chitinase isoforms from Vibrio alginolyticus 283. Enzyme and Microbial Technology, 41, 212–220.
Suginta, W., Vongsuwan, A., Songsiriritthigul, C., Prinz, H., Estibeiro, P., Duncan, R. R., Svasti, J., & Fothergill-Gilmore, L. A. (2004). An endochitinase A from Vibrio carchariae: cloning, expression, mass and sequence analyses, and chitin hydrolysis. Archives of Biochemistry and Biophysics, 424, 171–180.
Songsiriritthigul, C., Pantoom, S., Aguda, A. H., Robinson, R. C., & Suginta, W. (2008). Crystal structures of Vibrio harveyi chitinase A complexed with chitooligosaccharides: implications for the catalytic mechanism. Journal of Structural Biology, 162, 491–499.
Suginta, W., Pantoom, S., & Prinz, H. (2009). Substrate binding modes and anomer selectivity of chitinase A from Vibrio harveyi. Journal of Chemical Biology, 2, 191–202.
Suginta, W., Songsiriritthigul, C., Kobdaj, A., Opassiri, R., & Svasti, J. (2007). Mutations of Trp275 and Trp397 altered the binding selectivity of Vibrio carchariae chitinase A. BBA-General Subjects, 1770, 1151–1160.
Pantoom, S., Songsiriritthigul, C., & Suginta, W. (2008). The effects of the surface-exposed residues on the binding and hydrolytic activities of Vibrio carchariae chitinase A. BMC Biochemistry, 9, 2.
Laemmli, U. K. (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature, 227, 680–685.
Bradford, M. M. (1976). A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Analytical Biochemistry, 72, 248–254.
Miller, G. L. (1959). Use of dinitrosalicylic acid reagent for determination of reducing sugar. Analytical Chemistry, 31, 426–428.
Hsu, S. C., & Lockwood, J. L. (1975). Powdered chitin agar as a selective medium for enumeration of actinomycetes in water and soil. Applied Microbiology, 29, 422–426.
Suginta, W., Vongsuwan, A., Songsiriritthigul, C., Svasti, J., & Prinz, H. (2005). Enzymatic properties of wild-type and active site mutants of chitinase A from Vibrio carchariae, as revealed by HPLC-MS. FEBS Journal, 272, 3376–3386.
Suginta, W., Chuenark, D., Mizuhara, M., & Fukamizo, T. (2010). Novel β-N-acetylglucosaminidases from Vibrio harveyi 650: cloning, expression, enzymatic properties, and subsite identification. BMC Biochemistry, 11, 40.
Eijsink, V. G., Vaaje-Kolstad, G., Vårum, K. M., & Horn, S. J. (2008). Towards new enzymes for biofuels: lessons from chitinase research. Trends in Biotechnology, 26, 228–235.
Zakariassen, H., Aam, B. B., Horn, S. J., Vårum, K. M., Sørlie, M., & Eijsink, V. G. (2009). Aromatic residues in the catalytic center of chitinase A from Serratia marcescens affect processivity, enzyme activity, and biomass converting efficiency. Journal of Biological Chemistry, 284, 10610–10617.
Aronson, N. N., Jr., Halloran, B. A., Alexyev, M. F., Amable, L., Madura, J. D., Pasupulati, L., Worth, C., & Van Roey, P. (2003). Family 18 chitinase-oligosaccharide substrate interaction: subsite preference and anomer selectivity of Serratia marcescens chitinase A. Biochemistry Journal, 376, 87–95.
Acknowledgments
This work was supported financially by The Thailand Research Fund through TRF-CHE Research Grant for Mid-Career University (Grant no RMU4980028) and Suranaree University of Technology, Thailand.
Conflict of interest
None declared.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. S1
Michaeli–Menten plots of VhChiA variants. For pNP-GlcNAc2, the reaction was set up with concentrations of the substrate varied from 0 to 500 μM. For chitooligosaccharides, the reaction mixture (200 μL), containing 0–500 μM GlcNAc3–6 and 0.8 nM enzyme in 0.1 M sodium acetate buffer, pH 5.0, was incubated at 37 °C for 15 min. After heating at 100 °C for 10 min, the entire reaction mixture was used in the DNS assay. For colloidal and glycol chitins, the reaction was carried out the same way as for chitooligosaccharides, but concentrations of the chitins were varied from 0% to 5% (w/v). The kinetic values were evaluated the nonlinear regression function obtained from the GraphPad Prism v5.0. (JPEG 99 kb)
Rights and permissions
About this article
Cite this article
Sritho, N., Suginta, W. Role of Tyr-435 of Vibrio harveyi Chitinase A in Chitin Utilization. Appl Biochem Biotechnol 166, 1192–1202 (2012). https://doi.org/10.1007/s12010-011-9504-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-011-9504-8