Abstract
Purpose of Review
Common genetic variants that associate with type 2 diabetes risk are markedly enriched in pancreatic islet transcriptional enhancers. This review discusses current advances in the annotation of islet enhancer variants and their target genes.
Recent Findings
Recent methodological advances now allow genetic and functional mapping of diabetes causal variants at unprecedented resolution. Mapping of enhancer-promoter interactions in human islets has provided a unique appreciation of the complexity of islet gene regulatory processes and enabled direct association of noncoding diabetes risk variants to their target genes.
Summary
The recently improved human islet enhancer annotations constitute a framework for the interpretation of diabetes genetic signals in the context of pancreatic islet gene regulation. In the future, integration of existing and yet to come regulatory maps with genetic fine-mapping efforts and in-depth functional characterization will foster the discovery of novel diabetes molecular risk mechanisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, clinical genomics and the investigation of noncoding genome functions, including microRNAs (miRNAs), long noncoding RNAs (lncRNAs) and transcriptional enhancers, have proved to be fertile ground to the discovery of genetic disease mechanisms [1]. Sequence variation leading to altered enhancer function is now appreciated as a driving mechanism in cancer [2,3,4] and a number of Mendelian disorders are caused by mutations affecting tissue-specific cis-regulatory elements [5], including non-syndromic pancreas agenesis [6]. The overwhelming majority of genetic variants associated with common human diseases and disease-associated traits are noncoding and concentrate within accessible chromatin regions of disease-relevant tissues [7]. This observation is also true for type 2 diabetes (T2D), for which common noncoding variants predominantly influence risk [8•], residing in pancreatic islet transcriptional enhancers [9,10,11,12, 13••, 14, 15], particularly within clustered enhancers [9,10,11,12,13]. Thus, most of the genetic susceptibility to T2D seems to arise from islet regulatory defects, which aligns well with the observation that many T2D-associated variants also affect measures of pancreatic β cell function [14,15,16]. Moreover, recent reports indicate that islet enhancers also harbour genetic variants that confer susceptibility to type 1 diabetes [17], even if not to the same extent as for T2D [18]. These observations combined with recent technological advances have prompted the production of evermore refined maps of human islet cis-regulatory elements in recent years, including whole islet, cell type-specific and disease state-specific maps, improving the functional annotation of diabetes risk variants. In this review, I cover the progress that has been made in the mapping of active islet enhancers and its implications for the identification of causal diabetes risk variants.
A Decade of Islet Regulomes: Restricting the Search Space for Diabetes Causal Variants
Since T2D risk variants are mostly noncoding [8•] and predominantly enriched in islet enhancers [9,10,11,12, 13••, 14, 15], defining the regulatory regions that are active in pancreatic islets is key to identifying truly causal variants. Several iterations of human islet cis-regulatory maps or regulomes have been built to date and a number of publicly accessible genome browsers now host human islet regulomes, including the Roadmap Epigenomics Project (https://egg2.wustl.edu/roadmap/web_portal/), the Islet Regulome Browser (www.isletregulome.org/) and the Diabetes Epigenome Atlas (https://www.t2depigenome.org/).
Although providing an exhaustive overview of the evolution of islet enhancer maps is not the aim of this review, a few general points are still worth mentioning. In earlier maps, islet enhancers were defined as broad domains enriched in typical active chromatin marks (H3K4me1 and H3K27ac), often spanning several kilobases, whereas currently enhancers are more commonly defined as accessible chromatin sites typically spanning 300–500 bp that overlap chromatin regions enriched in active chromatin features. Moreover, due to lower starting material requirements, ATAC-seq [19] was adopted in recent years as the standard method to map regulatory chromatin sites in human islets [12, 13••, 20, 21•, 22].
Improved understanding of the biochemical properties that distinguish active from inactive chromatin has enabled a progressive subcategorization of islet accessible chromatin regions and identification of specific enhancer subsets that are more relevant for diabetes risk. For example, Miguel-Escalada et al. have recently sub-classified active human islet enhancers into three classes according to their enrichment in active chromatin features (H3K27ac and the coactivator complex Mediator) [13••]. This subclassification revealed a subset of 13,635 strong islet enhancers that tended to regulate islet-specific genes and contribute more to diabetes and β cell-related trait heritability than other enhancer subsets [13••]. In another study, profiling of DNA methylation in human islets from 10 donors using whole-genome bisulphite sequencing revealed that T2D risk variants concentrate within open and hypomethylated regions [21•]. Future studies profiling additional enhancer features, including eRNA production [23] and enrichment in additional enhancer-associated histone modifications [24], have the potential to help obtain further granularity to recognize additional diabetes-relevant islet enhancer categories.
Strategies to Improve Causal Variant Mapping
A big challenge to transpose the knowledge arising from genetic association studies to disease mechanisms is the fact that the most significant variants identified in genome-wide association studies (GWAS) may not be the true causal ones, but simply be in high linkage disequilibrium with the causal variant. Several approaches may be taken to improve the resolution of causal variant mapping, including the analysis of larger numbers of individuals [25••] in diverse populations [26,27,28,29], and by applying genetic fine-mapping [18, 25••, 30,31,32] (reviewed in [33]). Remarkably, a recent meta-analysis of ~ 900,000 European descent individuals using genetic fine-mapping has expanded the repertoire of T2D risk loci to > 240 and isolated single causal variants at 18 association signals [25••]. Parallel efforts with > 400,000 East-Asian individuals implicated 56 additional loci, highlighting that the quest to identify T2D-associated loci is probably not over yet [34••].
It is reasonable to assume that for a given T2D locus where coding variants are not likely to be causal, variants residing within cis-regulatory elements are more likely to be driving the genetic risk (Fig. 1). This principle is the basis of FGWAS [35], in which the likelihood of a given GWAS variant being causal is weighted by functional annotations such as histone modification enrichment, chromatin accessibility or DNA methylation status [21•]. FGWAS has already been applied by a number of studies to fine-map T2D risk loci [8•, 21•, 25••, 36], with reported reductions in the order of 35% of the number of T2D-associated variants at specific loci [8•].
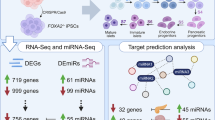
Overview of the workflow to prioritize noncoding variants and target gene investigation in T2D. Enhancer loss-of-function (LOF)1: enhancer LOF can be achieved by either indels at the core region (transcription factor binding sites), full deletion, or CRISPR-mediated inhibition (CRISPRi). Enhancer gain-of-function (GOF)2: CRISPR-mediated activation. Please note that the schematic does not provide an exhaustive list of all possible methods to prioritize genome-wide association study (GWAS) variants due to space limitations, providing instead the most frequently applied approaches
In order to correctly refine the sets of T2D variants using functional genomic data, not only sample purity and data quality are important, but also the type of functional data that is used to prioritize variants. For example, it has been observed that islet chromatin accessibility is more predictive of regulatory impact than DNA methylation [21•]. Moreover, clustered enhancers are more enriched in T2D variants than “orphan” enhancers [9, 10]. More recent analyses have also demonstrated that refining islet enhancer maps can help identify a restricted segment of the genome that is more relevant for islet gene regulation and, consequently, to T2D risk via β cell function impairment [13••]. In this study, an enhancer subset defined by strong enhancers that cluster in 3D (see the section “Enhancer Clusters in 3D” for details) was found to contribute more to the heritability of T2D and traits related of β cell function (HOMA-B and insulinogenic index from oral glucose tolerance test) compared to other classes of enhancers. Notably, this result was achieved not by covering a larger proportion of the genome with this annotation but, on the contrary, by restricting the annotation to open chromatin sites (marked by ATAC-seq) within regions that presented multiple features associated with tissue-specific regulation, such as stronger ChIP-seq enrichment in H3K27ac/Mediator, and being promoter-interacting.
Demonstrating Variant Causality
It is assumed that genetic variation in islet enhancers leads to changes in transcription factor binding and, consequently, enhancer activity and gene regulation. One way to measure this is by computing changes in transcription factor (TF) binding affinity using TF motif analysis, but this strategy is limited to known TF recognition sequences, which have only been identified for two thirds of human TFs [37]. Moreover, the presence of a particular TF motif in an enhancer does not necessarily reflect that it is a genuine binding site for that TF. In reality, in vivo TF binding is influenced by multiple factors, including cofactors, cooperativity, concentration and even chromatin shape [38]. A less biased approach is to directly interrogate sequence-dependent enhancer activity (Fig. 1). This strategy was successfully applied for a number of T2D-associated enhancer variants (see Table 1 for examples). Nonetheless, enhancer reporter assays also have limitations, as they interrogate candidate enhancer sequences outside of their native context. An alternative to address this question is to directly probe the epigenomic profiles from multiple individuals to identify genetic variants that confer allele-dependent chromatin activity (chromatin activity quantitative trait loci, QTLs) (Fig. 1). Three studies have already yielded encouraging results by performing chromatin accessibility QTL (caQTL) analysis on human islet ATAC-seq profiles from 17 [21•], 19 [20] and 23 individuals [22]. In one of these studies, Thurner et al. confidently reduced T2D GWAS signals at CAMK1D, KLHDC5 and ADCY5 to single regulatory variants by combining genetic fine-mapping, FGWAS and caQTL analysis [21•]. In another study, Khetan et al. identified nearly three thousand SNPs that yielded allele-dependent changes in islet chromatin accessibility, including 13 T2D-associated variants [20]. This number was later increased to 24 T2D variants by addition of four new islet samples [22]. Thus, future analyses with larger sample sizes and meta-analysis of existing human islet ATAC-seq datasets [12, 13, 20, 21•, 22, 39] hold promise for uncovering additional T2D risk variants that influence islet enhancer function. Similarly, allelic imbalance analysis of active chromatin histone modifications, such as H3K27ac, and islet TF occupancy may assist the identification of T2D regulatory variants, as it has been demonstrated at individual T2D loci [17, 40].
Enhancer Clusters in 3D
Three-dimensional (3D) chromatin studies with techniques such as Hi-C [41] and genome architecture mapping (GAM) [42], which allow identification of pairwise chromatin interactions in the entire genome, have put back in the spotlight 3D genome organization in the context of gene regulation and disease, and a number of recent reviews cover this topic extensively [43,44,45, 46•].
In one of the earlier maps of islet enhancers, the authors noted that islet-specific genes tended to be regulated not by one, but by multiple enhancer clusters [10]. In the same study, the authors detected long-range interactions between single promoters and multiple enhancers by performing circular chromosome conformation capture coupled with high-throughput sequencing (4C-seq) in nine loci [10]. These observations lend plausibility to the hypothesis that clustered enhancers form higher-order 3D regulatory structures to control cell-specific expression, and agree with well-characterized higher-order 3D regulatory structures detected in specific loci, including the HoxD cluster [47], and the α- and β-globin loci [48, 49]. However, the extent to which the formation of higher-order 3D regulatory structures is a general property of enhancers and enhancer clusters, and its contribution to the establishment of tissue-specific cis-regulatory programs could not be fully appreciated in these studies.
Recently, Ferrer and colleagues have addressed this question by generating a high-resolution map of promoter-mediated interactions in human pancreatic islets from four different donors using promoter capture Hi-C (pcHi-C) [13••]. In agreement with earlier observations [10], the authors observed that (1) islet genes often interact with multiple enhancers and clusters of enhancers and that (2) islet enhancers often interact with multiple target genes. Similar observations were made in a lower resolution map of 3D chromatin contacts obtained by performing Hi-C in three human islet samples [22]. Overall, this demonstrates that islet enhancers and their target promoters reside in restricted 3D chromatin domains, named enhancer hubs for their highly connected nature [13••]. Using a systematic approach, the authors defined > 1300 islet enhancer hubs, defining the portion of the genome that is particularly enriched in islet enhancers and islet enhancer-promoter interactions. Enhancer hubs include genes that tend to be more islet enriched and predominantly involved in islet cell function and diabetes risk [13••]. These results agree with work performed in other human cell lineages, where super-enhancers were found to regulate tissue-specific genes within frequently interacting regions (FIREs) [50]. Although the methods applied in these studies could not discern whether the detected pairwise chromatin interactions occur simultaneously or alternate, multi-way contacts between enhancers and promoters have been detected in other settings using techniques such as GAM [42], Tri-C [48] and multi-contact 4C [51]. It is thus likely that similar multi-way enhancer-promoter contacts exist in human pancreatic islet chromatin.
Beyond their role as genomic regions that passively harbour regulatory elements and genes important for tissue-specific regulation, there is evidence indicating that enhancer hubs also represent functional units that are important for islet gene regulation. Multiple lines of evidence support this notion, both in steady-state and in dynamic settings. First, the activity of enhancers and genes contained in the same hub tends to correlate more than for those located outside hubs [13••], an observation analogous to cis-regulatory domains recently defined in a large series of lymphoblastoid cell lines and primary fibroblasts [52]. Second, when comparing enhancer activity in two different glucose concentrations (4 and 11 mM), hub enhancers frequently showed coordinated changes in their activity, not only in terms of direction of effect, but also in effect size [13••]. Third, perturbations of single hub enhancers or promoters with CRISPR in the glucose-responsive pancreatic β cell line EndoC βH3 [53] frequently yielded changes in the expression of multiple genes in the same hub, demonstrating connectivity of hub cis-regulatory elements [13••]. There is therefore cumulative evidence indicating that 3D regulatory domains such as enhancer hubs are key for the establishment and maintenance of cell identity and function. Given the presence of many T2D risk variants in islet enhancers, particularly in strong hub enhancers [13••], these observations raise the question of whether individual T2D variants could be involved in dysregulation of multiple genes and gene pathways.
Harnessing 3D Genome Maps to Identify Diabetes Effector Transcripts
On its own, the observation that T2D risk variants predominantly locate to islet enhancers, will not bring us closer to identifying disease effector transcripts, as genome folding enables transcriptional enhancers to act over genes at varying distances in the linear genome, sometimes even over non-adjacent genes located hundreds of kilobases away (reviewed in [44]). The top obesity-associated locus illustrates well this model, in which the risk variant resides in an intronic enhancer at FTO and regulates the expression of two distal genes (IRX3 and IRX5), but not the FTO gene [54, 55•]. Thus, enhancer-promoter functional relationships cannot be by simply inferred by examining linear representations of the genome.
A few studies have aimed to link noncoding T2D risk variants to target genes using human islet expression quantitative trait loci (eQTL) analysis [13••, 56,57,58, 59•], the latest of which included pancreatic islet RNA-seq from 420 donors [59•] and identified candidate effector transcripts for 23 loci. While the results are encouraging, they still fall short from delivering a comprehensive assignment of regulatory variants to effector transcripts. Approaches such as chromosome conformation capture (3C) techniques, including Hi-C [41], ChIA-PET [60] and HiChIP [61], yield genome-scale maps of long-range functional chromatin interactions, but it should be noted that 3C methods are usually biased for detection of long-range interactions. Thus, eQTL and chromatin interaction studies should be taken as complementary, rather than alternative, approaches to identity disease effector transcripts (Fig. 1).
Islet 3D promoter-enhancer contacts were first used to define the regulatory landscapes for a select number of loci [10, 40, 62]. More recently, two studies aimed to associate islet enhancers with target genes using genome-scale 3D chromatin contact mapping in human pancreatic islets [13••, 22] and one in the pancreatic β cell line EndoC βH1 [63]. Greenwald et al. generated Hi-C maps in three human islet samples, identifying 3022 islet enhancers (6.7% of all enhancers mapped in the study) looping to at least one promoter [22]. As in other reports of enhancer-promoter 3D interactions, islet chromatin maps have revealed substantial number of very long-range interactions, sometimes spanning over 1 Mb [13••, 22]. Hi-C and ChIA-PET in EndoC βH1 cells demonstrated the overall suitability of this cell line to study islet cis-regulatory functions, with high correlation of interaction profiles across most loci [63]. Hi-C maps, however, offer limited resolution to identify specific functional interactions [64], and targeted methods that enrich Hi-C libraries for genomic regions of interest have the potential to uncover a larger number of functional 3D chromatin interactions, being therefore more informative on the target genes of disease-associated distal regulatory elements [65]. Following this logic, Miguel-Escalada et al. applied a variant of Hi-C, promoter capture Hi-C (pcHi-C), to enrich the maps of islet 3D chromatin interactions in promoter-mediated interactions [13••]. Using this approach, the authors substantially improved the mapping of enhancer-promoter interactions, detecting one or more interactions for ~ 40% of all annotated enhancers (18,031 enhancers). The highly connected nature of particular loci and the definition of enhancer hubs (see the section “Enhancer Clusters in 3D” for details) allowed the authors to infer target genes for a further 40% of the mapped islet enhancers, assigning in total 53 T2D- or fasting glycaemia-associated enhancers to at least one target gene. Surprisingly though, in 75% of the loci where at least one risk SNP falls in an islet enhancer, the authors linked it to one or more distal genes [13••]. This observation has been partially supported by human islet eQTL studies, where a few T2D-associated SNPs have been linked to genes other than their closest, including variants at CDC123 (linked to CAMK1D), ARAP1 (STARD10) and ZBED3 (PDE8B) [13••, 57, 58, 59•], and by CRISPR-mediated perturbations of diabetes-associated enhancers [13••].
At the ZBED3 locus, for example, CRISPR-mediated activation or repression of an islet enhancer carrying a regulatory T2D risk variant [22, 25••] in EndoC βH3 cells resulted in changes in the expression of multiple genes [13••]. Of the affected transcripts, PDE8B showed the strongest response to the enhancer perturbations. This result is quite interesting in light of the most recent human islet eQTL study, where PDE8B was identified as the effector transcript of this association signal [59•]. In this case, PDE8B seems like a good candidate to pursue with functional studies, as it encodes for a high-affinity cAMP-specific phosphodiesterase and its reduction has been proposed as an approach to enhance insulin response to glucose [66]. It must be noted however that changes in PDE8B levels may not make up the full story in this T2D locus, as the CRISPR enhancer perturbations in human β cells affected additional genes, including the lncRNA ZBED3-AS1 and ZBED3 itself [13••]. Long noncoding RNAs can have diverse functions and have been proposed as targets for modulation of diabetes-relevant genes in islets [67]. ZBED3 is a secreted protein, whose higher circulating levels have been associated with insulin resistance [68] and metabolic syndrome [69]. Furthermore, this T2D-associated enhancer is also active in adipose tissue [70]. Thus, genetic variation at ZBED3 could in theory contribute to diabetes risk via multiple cell-specific processes. Overall, these results illustrate how complex the genetic factors that contribute to T2D may be from a molecular standpoint, and exemplify the complementarity of islet 3D genome maps, eQTL and (epi)genome editing approaches to better define disease effector transcripts and gain insights into the molecular underpinnings of specific GWAS signals. More generally, these studies have important implications for the general interpretation of GWAS signals, which traditionally assign association signals to their nearest gene.
Islet Enhancer Landscapes Are Dynamic
The interaction between chromatin landscapes and T2D goes beyond the co-localization of risk variants with islet enhancers. In reality, islet enhancers are dynamically regulated, responding to external stimuli such as elevated glucose concentrations [13••] or exposure to pro-inflammatory cytokines [71••]. Thus, mapping of islet enhancers under different conditions may uncover new diabetes-related molecular mechanisms, as it has been recently shown in the context of type 1 diabetes [71••].
Studies comparing islets from donors with diabetes versus those from donors without the disease have demonstrated that T2D associates with changes at both transcriptional [56, 72] (single-cell studies addressing this question have been reviewed in [73•]) and chromatin levels [20, 21•, 39, 74, 75]. Significant changes were already detected when comparing the ATAC-seq profiles of islets from five donors with diabetes versus five donors without diabetes [20]. It must be noted however that the majority of T2D-associated changes were quantitative, not qualitative. In other words, the authors did not find evidence that T2D islets have a different set of active enhancers. Instead, the activity level of existing enhancers was found altered [20]. This observation seems sensible considering that T2D is progressive in nature and does not constitute a severe disease with strong transcriptional phenotypes, as observed in developmental disorders or cancer.
Further supporting the dynamic nature of islet chromatin landscapes, global loss of polycomb repression has been associated with β cell dedifferentiation, diabetes and age [74, 76], while other studies identified differentially methylated regions in T2D islets [21•, 75]. Altogether, these observations render several questions that should be addressed in future studies: do T2D risk variants affect islet enhancer dynamics, and do factors such as age and metabolic state change the regulatory impact of T2D variants?
Islet Cell-Specific Enhancers
As shown in the sections above, the study of whole human islets has provided important insights into the genetic architecture of T2D, fine-mapping noncoding sequences that are important for islet gene regulation. However, these studies were not designed to address the cellular heterogeneity of pancreatic islets [77], in which distinct endocrine cell lineages contribute to glucose homeostasis, including the glucagon-producing α, insulin-producing β and somatostatin-producing δ cells. Islet morphological and functional heterogeneity goes beyond the known endocrine cell lineages, as different β cell subpopulations have also been detected [78,79,80]. Furthermore, analysis of whole islets may not detect features restricted to rare, but nonetheless important, endocrine cell types. Reflecting the functional heterogeneity of islet cells, their transcriptomes differ substantially, as it has been observed in sorted cell populations [81], by deploying single-cell RNA-seq [73•] and, more recently, using single-molecule RNA FISH [82]. Transcriptional-level differences often reflect different chromatin landscapes. Indeed, even closely related cell lineages, such as pancreatic islet α and β cells, show differences at chromatin level, with differential methylation of cell-specific enhancer sites [83]. In an elegant study by Seung Kim and colleagues, the authors combined purification of specific pancreatic cell populations with ATAC-seq and ChIP-seq analysis to generate regulomes for human α, β, acinar and ductal cells [84••], identifying thousands of enhancers and promoters that are lineage-specific, including 3999 β and 5316 α cell-specific regulatory regions. In these cell-specific regulatory maps, endocrine lineage-specific regions tended to locate near genes generally involved in glucose homeostasis (e.g. INS, GCG, GLP1R, IRS1/2) and were enriched in T2D-associated variants, in contrast with non-endocrine lineages [84••].
More recently, single nuclei ATAC-seq (snATAC-seq) was deployed on human islets from three donors, revealing 13 cell clusters with different regulatory landscapes, which included the classic hormone expressing α, β and δ cells, but also usually less appreciated islet cell populations such as immune and endothelial cells [85••]. Reflecting the different regulatory landscapes that co-occur in islets, this study revealed that regulatory elements of different lineages are enriched in different sets of TF recognition sequences. Moreover, in agreement with reports of heterogeneity among β cells [78,79,80], the authors detected two β cell clusters characterized by different accessibility of the insulin promoter (INShigh and INSlow). Interestingly, although both INShigh and INSlow accessible sites were enriched in T2D- and fasting glycemia-associated variants, INShigh sites revealed a stronger enrichment [85••]. The value of these cell-specific regulatory maps is exemplified at DGKB, where a fine-mapped T2D risk variant that modulates regulatory activity [59•] was found in an INShigh-specific accessible region [85••]. Strikingly, this regulatory region was not previously annotated as active in bulk islet enhancer maps [59•]. Altogether, these studies provide a new layer of detail for the interpretation of T2D-associated variants and will aid the identification of affected cell-specific processes.
Perspectives and Future Directions
Great progress has been made in recent years to define the epigenomic landscape of human pancreatic islets, but the studies discussed in this review also highlight that the picture we currently have of the genetic architecture of polygenic diabetes is probably a lot more complex than previously anticipated. Future studies are likely to uncover even more common variants that confer diabetes risk. In particular, the greater investigation of non-European populations is likely to point to novel implicated loci [34••].
On the other hand, the advent of technologies that enable epigenomic profiling at single-cell resolution is expected to further advance the identification of cell-specific, as well as developmental-, metabolic- and disease stage-specific regulatory elements. A recently developed strategy proposed by the Buenrostro lab is particularly promising, having taken the throughput of single-cell ATAC-seq to the hundreds of thousands of cells with deeper genome coverage than previous methods [86]. Similar progress is being made to profile chromatin histone modifications at single-cell level, with single-cell ChIP-seq [87] and CUT&Tag [88]. It is therefore expected that in the near future more studies will address the regulatory landscapes of human pancreatic islets at single-cell level, hopefully obtaining important insights into the cis-regulatory networks that drive islet dysfunction in diabetes, as demonstrated by the first islet epigenomic maps with single-cell resolution [85••]. Greater attention is also expected to be given to the regulatory mechanisms that associate with β cell dysfunction during different diabetes disease trajectories [89•, 90].
In order to draw clinically relevant conclusions, genomic evidence of causality, including genetic association, epigenomic annotation, eQTL and caQTL, will have to pass through detailed functional validation in appropriate cellular and animal models (Fig. 1). Mechanistic studies of T2D-associated enhancers and putative effector transcripts should be designed taking into account the direction of effect of T2D variants, as not all T2D risk variants lead to loss-of-function (Table 1). Furthermore, future examination of risk variants should aim to go beyond regulatory and transcriptional outputs and also assess relevant cellular functions, as previously shown at ZMIZ1 [57], ADCY5 [40], ARAP1/STARD10 [36], and SLC30A8 [91] and at genome-scale [92]. β cell genome and epigenome editing at individual loci [13••, 40] and in the shape of large-scale screens [93] have already started to provide a better understanding of the regulatory mechanisms that operate at T2D-associated loci. It is thus expected that additional application of these methodologies to investigate putative causal variants in their genomic and cellular contexts will advance our understanding of diabetes genetic risk processes. Balboa et al. have recently provided a thorough overview of the applications of genome editing in human pancreatic β cell models [94••].
Despite the strong enrichment of T2D variants in islet enhancers, it is clear that not all risk variants act via islet dysfunction, as some correlate with insulin resistance [16, 95]. What is more, a subset of T2D-associated islet enhancers are also active in other diabetes-relevant tissues, such as adipose tissue and the liver [25••]. Thus, combining islet epigenomic maps with those of other relevant tissues will enable a more comprehensive characterization of risk variants.
Altogether, these studies have the potential to enable stratification of individuals by genetic risk of undergoing a specific disease trajectory [89•, 96]. At least five distinct pathways have been reported to drive T2D risk, including two related to β cell function [97•]. This concept has been recently tested by deploying islet enhancer maps to identify individuals at higher risk of developing T2D due to islet dysfunction [13••]. Individuals in this group tended to develop T2D at a younger age and with lower BMI, resembling to some extent individuals who present monogenic forms of diabetes, which is predominantly caused by mutations in islet TFs. Future studies with inclusion of even more refined enhancer maps for islets and other T2D-relevant tissues may therefore help deliver personalized medicine to at least a subset of patients with T2D.
Conclusions
Great progress has been made in recent years to assist efforts to define islet cell-specific diabetes risk mechanisms using enhancer maps. Nevertheless, the studies discussed in this review also highlight that the picture we currently have of the genetic architecture of polygenic diabetes is probably still incomplete. Future studies leveraging on lower sequencing costs, technological advances such as application of machine learning for noncoding variant prioritization, ATAC-seq on clinical specimens, single-cell genomics and genome-scale genetic screens, as well as on the wealth of epigenomic datasets that are already available, will further the discovery of diabetes risk mechanisms and aid patient stratification by molecular aetiology.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Gloss BS, Dinger ME. Realizing the significance of noncoding functionality in clinical genomics. Exp Mol Med. 2018;50:97. https://doi.org/10.1038/s12276-018-0087-0.
Hnisz D, Abraham BJ, Lee TI, Lau A, Saint-André V, Sigova AA, et al. Super-enhancers in the control of cell identity and disease. Cell. 2013;155:934–47. https://doi.org/10.1016/j.cell.2013.09.053.
Ongen H, Andersen CL, Bramsen JB, Oster B, Rasmussen MH, Ferreira PG, et al. Putative cis-regulatory drivers in colorectal cancer. Nature. 2014;512:87–90. https://doi.org/10.1038/nature13602.
Cohen AJ, Saiakhova A, Corradin O, Luppino JM, Lovrenert K, Bartels CF, et al. Hotspots of aberrant enhancer activity punctuate the colorectal cancer epigenome. Nat Commun. 2017;8:14400. https://doi.org/10.1038/ncomms14400.
Miguel-Escalada I, Pasquali L, Ferrer J. Transcriptional enhancers: functional insights and role in human disease. Curr Opin Genet Dev. 2015;33:71–6. https://doi.org/10.1016/j.gde.2015.08.009.
Weedon MN, Cebola I, Flanagan SE, et al. Recessive mutations in a distal PTF1A enhancer cause isolated pancreatic agenesis. Nat Genet. 2014;46:61–4. https://doi.org/10.1038/ng.2826.
Maurano MT, Humbert R, Rynes E, Thurman RE, Haugen E, Wang H, et al. Systematic localization of common disease-associated variation in regulatory DNA. Science. 2012;337:1190–5. https://doi.org/10.1126/science.1222794.
• Fuchsberger C, Flannick J, Teslovich TM, Mahajan A, Agarwala V, Gaulton KJ, et al. The genetic architecture of type 2 diabetes. Nature. 2016;536:41–7. https://doi.org/10.1038/nature18642 This study demonstrates that the genetic basis of T2D arises predominantly from common noncoding variants of small effect size.
Parker SCJ, Stitzel ML, Taylor DL, Orozco JM, Erdos MR, Akiyama JA, et al. Chromatin stretch enhancer states drive cell-specific gene regulation and harbor human disease risk variants. Proc Natl Acad Sci U S A. 2013;110:17921–6. https://doi.org/10.1073/pnas.1317023110.
Pasquali L, Gaulton KJ, Rodríguez-Seguí SA, Mularoni L, Miguel-Escalada I, Akerman İ, et al. Pancreatic islet enhancer clusters enriched in type 2 diabetes risk-associated variants. Nat Genet. 2014;46:136–43. https://doi.org/10.1038/ng.2870.
Cebola I, Pasquali L. Non-coding genome functions in diabetes. J Mol Endocrinol. 2016;56:R1–R20. https://doi.org/10.1530/JME-15-0197.
Varshney A, Scott LJ, Welch RP, Erdos MR, Chines PS, Narisu N, et al. Genetic regulatory signatures underlying islet gene expression and type 2 diabetes. Proc Natl Acad Sci U S A. 2017;114:2301–6. https://doi.org/10.1073/pnas.1621192114.
•• Miguel-Escalada I, Bonàs-Guarch S, Cebola I, et al. Human pancreatic islet three-dimensional chromatin architecture provides insights into the genetics of type 2 diabetes. Nat Genet. 2019;51:1137–48. https://doi.org/10.1038/s41588-019-0457-0 This study provides a genome-wide map of human islet 3D chromatin interactions, linking diabetes-associated enhancers to target genes, and exemplifies the integration of islet epigenomic annotations with 3D interaction maps, eQTLs and CRISPR validations.
Dupuis J, Langenberg C, Prokopenko I, Saxena R, Soranzo N, Jackson AU, et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat Genet. 2010;42:105–16. https://doi.org/10.1038/ng.520.
Scott RA, Lagou V, Welch RP, et al. Large-scale association analyses identify new loci influencing glycemic traits and provide insight into the underlying biological pathways. Nat Genet. 2012;44:991–1005. https://doi.org/10.1038/ng.2385.
Scott RA, Scott LJ, Magi R, et al. An expanded genome-wide association study of type 2 diabetes in Europeans. Diabetes. 2017;66:2888–902. https://doi.org/10.2337/db16-1253.
Aylward A, Chiou J, Okino M-L, Kadakia N, Gaulton KJ. Shared genetic risk contributes to type 1 and type 2 diabetes etiology. Hum Mol Genet. 2018. https://doi.org/10.1093/hmg/ddy314.
Onengut-Gumuscu S, Chen W-M, Burren O, et al. Fine mapping of type 1 diabetes susceptibility loci and evidence for colocalization of causal variants with lymphoid gene enhancers. Nat Genet. 2015;47:381–6. https://doi.org/10.1038/ng.3245.
Buenrostro JD, Giresi PG, Zaba LC, Chang HY, Greenleaf WJ. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat Methods. 2013;10:1213–8. https://doi.org/10.1038/nmeth.2688.
Khetan S, Kursawe R, Youn A, Lawlor N, Jillette A, Marquez EJ, et al. Type 2 diabetes-associated genetic variants regulate chromatin accessibility in human islets. Diabetes. 2018;67:2466–77. https://doi.org/10.2337/db18-0393.
• Thurner M, van de Bunt M, Torres JM, Mahajan A, Nylander V, Bennett AJ, et al. Integration of human pancreatic islet genomic data refines regulatory mechanisms at type 2 diabetes susceptibility loci. Elife. 2018;7:1363. https://doi.org/10.7554/eLife.31977 This study showcases how integration of multiple epigenomic datasets, including DNA methylation, can be used to functionally fine-map T2D risk variants.
Greenwald WW, Chiou J, Yan J, Qiu Y, Dai N, Wang A, et al. Pancreatic islet chromatin accessibility and conformation reveals distal enhancer networks of type 2 diabetes risk. Nat Commun. 2019;10:2078. https://doi.org/10.1038/s41467-019-09975-4.
Andersson R, Gebhard C, Miguel-Escalada I, et al. An atlas of active enhancers across human cell types and tissues. Nature. 2014;507:455–61. https://doi.org/10.1038/nature12787.
Pradeepa MM. Causal role of histone acetylations in enhancer function. Transcription. 2017;8:40–7. https://doi.org/10.1080/21541264.2016.1253529.
•• Mahajan A, Taliun D, Thurner M, et al. Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nat Genet. 2018;50:1505–13. https://doi.org/10.1038/s41588-018-0241-6 This study provides the most comprehensive picture of the genetic architecture of T2D to date, uncovering 403 independent association signals.
DIAbetes Genetics Replication And Meta-analysis (DIAGRAM) Consortium, Asian Genetic Epidemiology Network Type 2 Diabetes (AGEN-T2D) Consortium, South Asian Type 2 Diabetes (SAT2D) Consortium, et al. Genome-wide trans-ancestry meta-analysis provides insight into the genetic architecture of type 2 diabetes susceptibility. Nat Genet. 2014;46:234–44. https://doi.org/10.1038/ng.2897.
Liu C-T, Raghavan S, Maruthur N, Kabagambe EK, Hong J, Ng MC, et al. Trans-ethnic meta-analysis and functional annotation illuminates the genetic architecture of fasting glucose and insulin. Am J Hum Genet. 2016;99:56–75. https://doi.org/10.1016/j.ajhg.2016.05.006.
Magi R, Horikoshi M, Sofer T, et al. Trans-ethnic meta-regression of genome-wide association studies accounting for ancestry increases power for discovery and improves fine-mapping resolution. Hum Mol Genet. 2017;26:3639–50. https://doi.org/10.1093/hmg/ddx280.
Wojcik GL, Graff M, Nishimura KK, Tao R, Haessler J, Gignoux CR, et al. Genetic analyses of diverse populations improves discovery for complex traits. Nature. 2019;570:514–8. https://doi.org/10.1038/s41586-019-1310-4.
Gaulton KJ, Ferreira T, Lee Y, et al. Genetic fine mapping and genomic annotation defines causal mechanisms at type 2 diabetes susceptibility loci. Nat Genet. 2015;47:1415–25. https://doi.org/10.1038/ng.3437.
Horikoshi M, Mӓgi R, van de Bunt M, Surakka I, Sarin AP, Mahajan A, et al. Discovery and fine-mapping of glycaemic and obesity-related trait loci using high-density imputation. PLoS Genet. 2015;11:e1005230. https://doi.org/10.1371/journal.pgen.1005230.
Bonàs-Guarch S, Guindo-Martínez M, Miguel-Escalada I, Grarup N, Sebastian D, Rodriguez-Fos E, et al. Re-analysis of public genetic data reveals a rare X-chromosomal variant associated with type 2 diabetes. Nat Commun. 2018;9:321. https://doi.org/10.1038/s41467-017-02380-9.
Schaid DJ, Chen W, Larson NB. From genome-wide associations to candidate causal variants by statistical fine-mapping. Nat Rev Genet. 2018;19:491–504. https://doi.org/10.1038/s41576-018-0016-z.
•• Spracklen CN, Horikoshi M, Kim YJ, et al. Identification of type 2 diabetes loci in 433,540 East Asian individuals. bioRxiv. 2019. https://doi.org/10.1101/685172 This study exemplifies the potential of studying individuals of non-European ancestry to discover new genetic loci carrying T2D risk variants.
Pickrell JK. Joint analysis of functional genomic data and genome-wide association studies of 18 human traits. Am J Hum Genet. 2014;94:559–73. https://doi.org/10.1016/j.ajhg.2014.03.004.
Carrat GR, Hu M, Nguyen-Tu M-S, Chabosseau P, Gaulton KJ, van de Bunt M, et al. Decreased STARD10 expression is associated with defective insulin secretion in humans and mice. Am J Hum Genet. 2017;100:238–56. https://doi.org/10.1016/j.ajhg.2017.01.011.
Lambert SA, Jolma A, Campitelli LF, Das PK, Yin Y, Albu M, et al. The human transcription factors. Cell. 2018;172:650–65. https://doi.org/10.1016/j.cell.2018.01.029.
Slattery M, Zhou T, Yang L, Dantas Machado AC, Gordân R, Rohs R. Absence of a simple code: how transcription factors read the genome. Trends Biochem Sci. 2014;39:381–99. https://doi.org/10.1016/j.tibs.2014.07.002.
Bysani M, Agren R, Davegårdh C, Volkov P, Rönn T, Unneberg P, et al. ATAC-seq reveals alterations in open chromatin in pancreatic islets from subjects with type 2 diabetes. Sci Rep. 2019;9:7785. https://doi.org/10.1038/s41598-019-44076-8.
Roman TS, Cannon ME, Vadlamudi S, Buchkovich ML, Wolford BN, Welch RP, et al. A type 2 diabetes-associated functional regulatory variant in a pancreatic islet enhancer at the ADCY5 locus. Diabetes. 2017;66:2521–30. https://doi.org/10.2337/db17-0464.
Lieberman-Aiden E, van Berkum NL, Williams L, Imakaev M, Ragoczy T, Telling A, et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science. 2009;326:289–93. https://doi.org/10.1126/science.1181369.
Beagrie RA, Scialdone A, Schueler M, Kraemer DCA, Chotalia M, Xie SQ, et al. Complex multi-enhancer contacts captured by genome architecture mapping. Nature. 2017;543:519–24. https://doi.org/10.1038/nature21411.
Schmitt AD, Hu M, Ren B. Genome-wide mapping and analysis of chromosome architecture. Nat Rev Mol Cell Biol. 2016;17:743–55. https://doi.org/10.1038/nrm.2016.104.
Yu M, Ren B. The three-dimensional organization of mammalian genomes. Annu Rev Cell Dev Biol. 2017;33:265–89. https://doi.org/10.1146/annurev-cellbio-100616-060531.
Marti-Renom MA, Almouzni G, Bickmore WA, Bystricky K, Cavalli G, Fraser P, et al. Challenges and guidelines toward 4D nucleome data and model standards. Nat Genet. 2018;50:1352–8. https://doi.org/10.1038/s41588-018-0236-3.
• Stadhouders R, Filion GJ, Graf T. Transcription factors and 3D genome conformation in cell-fate decisions. Nature. 2019;569:345–54. https://doi.org/10.1038/s41586-019-1182-7 This review provides a comprehensive overview of the current state of the epigenomics field in the context of 3D genome architecture.
Montavon T, Soshnikova N, Mascrez B, Joye E, Thevenet L, Splinter E, et al. A regulatory archipelago controls Hox genes transcription in digits. Cell. 2011;147:1132–45. https://doi.org/10.1016/j.cell.2011.10.023.
Oudelaar AM, Davies JOJ, Hanssen LLP, Telenius JM, Schwessinger R, Liu Y, et al. Single-allele chromatin interactions identify regulatory hubs in dynamic compartmentalized domains. Nat Genet. 2018;50:1744–51. https://doi.org/10.1038/s41588-018-0253-2.
Tolhuis B, Palstra RJ, Splinter E, Grosveld F, de Laat W. Looping and interaction between hypersensitive sites in the active beta-globin locus. Mol Cell. 2002;10:1453–65.
Schmitt AD, Hu M, Jung I, Xu Z, Qiu Y, Tan CL, et al. A compendium of chromatin contact maps reveals spatially active regions in the human genome. Cell Rep. 2016;17:2042–59. https://doi.org/10.1016/j.celrep.2016.10.061.
Allahyar A, Vermeulen C, Bouwman BAM, Krijger PHL, Verstegen MJAM, Geeven G, et al. Enhancer hubs and loop collisions identified from single-allele topologies. Nat Genet. 2018;50:1151–60. https://doi.org/10.1038/s41588-018-0161-5.
Delaneau O, Zazhytska M, Borel C, Giannuzzi G, Rey G, Howald C, et al. Chromatin three-dimensional interactions mediate genetic effects on gene expression. Science. 2019;364:eaat8266–12. https://doi.org/10.1126/science.aat8266.
Benazra M, Lecomte M-J, Colace C, Müller A, Machado C, Pechberty S, et al. A human beta cell line with drug inducible excision of immortalizing transgenes. Mol Metab. 2015;4:916–25. https://doi.org/10.1016/j.molmet.2015.09.008.
Smemo S, Tena JJ, Kim K-H, Gamazon ER, Sakabe NJ, Gómez-Marín C, et al. Obesity-associated variants within FTO form long-range functional connections with IRX3. Nature. 2014;507:371–5. https://doi.org/10.1038/nature13138.
• Claussnitzer M, Dankel SN, Kim K-H, et al. FTO obesity variant circuitry and adipocyte browning in humans. N Engl J Med. 2015;373:895–907. https://doi.org/10.1056/NEJMoa1502214 This study combines detailed epigenomic and functional characterization of a GWAS locus.
Fadista J, Vikman P, Laakso EO, Mollet IG, Esguerra JL, Taneera J, et al. Global genomic and transcriptomic analysis of human pancreatic islets reveals novel genes influencing glucose metabolism. Proc Natl Acad Sci U S A. 2014;111:13924–9. https://doi.org/10.1073/pnas.1402665111.
van de Bunt M, Manning Fox JE, Dai X, Barrett A, Grey C, Li L, et al. Transcript expression data from human islets links regulatory signals from genome-wide association studies for type 2 diabetes and glycemic traits to their downstream effectors. PLoS Genet. 2015;11:e1005694. https://doi.org/10.1371/journal.pgen.1005694.
Khamis A, Canouil M, Siddiq A, Crouch H, Falchi M, Bulow M, et al. Laser capture microdissection of human pancreatic islets reveals novel eQTLs associated with type 2 diabetes. Mol Metab. 2019;24:98–107. https://doi.org/10.1016/j.molmet.2019.03.004.
• Viñuela A, Varshney A, van de Bunt M, et al. Influence of genetic variants on gene expression in human pancreatic islets – implications for type 2 diabetes. bioRxiv. 2019;50:1505. https://doi.org/10.1101/655670 This study provides the largest human islet eQTL dataset released to date.
Li G, Fullwood MJ, Xu H, Mulawadi FH, Velkov S, Vega V, et al. ChIA-PET tool for comprehensive chromatin interaction analysis with paired-end tag sequencing. Genome Biol. 2010;11:R22. https://doi.org/10.1186/gb-2010-11-2-r22.
Mumbach MR, Rubin AJ, Flynn RA, Dai C, Khavari PA, Greenleaf WJ, et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat Methods. 2016;13:919–22. https://doi.org/10.1038/nmeth.3999.
Jian X, Felsenfeld G. Insulin promoter in human pancreatic β cells contacts diabetes susceptibility loci and regulates genes affecting insulin metabolism. Proc Natl Acad Sci U S A. 2018;115:E4633–41. https://doi.org/10.1073/pnas.1803146115.
Lawlor N, Márquez EJ, Orchard P, Narisu N, Shamim MS, Thibodeau A, et al. Multiomic profiling identifies cis-regulatory networks underlying human pancreatic β cell identity and function. Cell Rep. 2019;26:788–801.e6. https://doi.org/10.1016/j.celrep.2018.12.083.
Schoenfelder S, Furlan-Magaril M, Mifsud B, Tavares-Cadete F, Sugar R, Javierre BM, et al. The pluripotent regulatory circuitry connecting promoters to their long-range interacting elements. Genome Res. 2015;25:582–97. https://doi.org/10.1101/gr.185272.114.
Javierre BM, Burren OS, Wilder SP, Kreuzhuber R, Hill SM, Sewitz S, et al. Lineage-specific genome architecture links enhancers and non-coding disease variants to target gene promoters. Cell. 2016;167:1369–1384.e19. https://doi.org/10.1016/j.cell.2016.09.037.
Dov A, Abramovitch E, Warwar N, Nesher R. Diminished phosphodiesterase-8B potentiates biphasic insulin response to glucose. Endocrinology. 2008;149:741–8. https://doi.org/10.1210/en.2007-0968.
Font-Cunill B, Arnes L, Ferrer J, Sussel L, Beucher A. Long non-coding RNAs as local regulators of pancreatic islet transcription factor genes. Front Genet. 2018;9:524. https://doi.org/10.3389/fgene.2018.00524.
Jia Y, Yuan L, Hu W, Luo Y, Suo L, Yang M, et al. Zinc-finger BED domain-containing 3 (Zbed3) is a novel secreted protein associated with insulin resistance in humans. J Intern Med. 2014;275:522–33. https://doi.org/10.1111/joim.12170.
Hu W, Tian B, Li X, Li L, Zhang L, Liu H, et al. Circulating Zbed3 levels in subjects with and without metabolic syndrome. Metab Syndr Relat Disord. 2017;15:207–12. https://doi.org/10.1089/met.2016.0122.
Roadmap Epigenomics Consortium, Kundaje A, Meuleman W, et al. Integrative analysis of 111 reference human epigenomes. Nature. 2015;518:317–30. https://doi.org/10.1038/nature14248.
•• Ramos-Rodríguez M, Raurell-Villa H, Colli ML, et al The impact of pro-inflammatory cytokines on the β-cell regulatory landscape provides new insights into the genetics of type 1 diabetes. Nat Genet. 2019;51:1588–1595. doi: https://doi.org/10.1038/s41588-019-0524-6. This study demonstrates that islet regulatory landscapes are largely dynamic and responsive to stimuli.
Morán I, Akerman I, van de Bunt M, Xie R, Benazra M, Nammo T, et al. Human β cell transcriptome analysis uncovers lncRNAs that are tissue-specific, dynamically regulated, and abnormally expressed in type 2 diabetes. Cell Metab. 2012;16:435–48. https://doi.org/10.1016/j.cmet.2012.08.010.
• Wang YJ, Kaestner KH. Single-cell RNA-Seq of the pancreatic islets--a promise not yet fulfilled? Cell Metab. 2019;29:539–44. https://doi.org/10.1016/j.cmet.2018.11.016 This review covers the currently available human islet single-cell RNA-seq datasets and sets the tone for future single-cell islet studies.
TT-H L, Heyne S, Dror E, et al. The polycomb-dependent epigenome controls β cell dysfunction, dedifferentiation, and diabetes. Cell Metab. 2018;27:1294–1308.e7. https://doi.org/10.1016/j.cmet.2018.04.013.
Volkov P, Bacos K, Ofori JK, Esguerra JLS, Eliasson L, Rönn T, et al. Whole-genome bisulfite sequencing of human pancreatic islets reveals novel differentially methylated regions in type 2 diabetes pathogenesis. Diabetes. 2017;66:1074–85. https://doi.org/10.2337/db16-0996.
Arda HE, Li L, Tsai J, Torre EA, Rosli Y, Peiris H, et al. Age-dependent pancreatic gene regulation reveals mechanisms governing human β cell function. Cell Metab. 2016;23:909–20. https://doi.org/10.1016/j.cmet.2016.04.002.
Grapin-Botton A, Serup P. Parsing the pancreas. N Engl J Med. 2017;376:886–8. https://doi.org/10.1056/NEJMcibr1616217.
Dorrell C, Schug J, Canaday PS, Russ HA, Tarlow BD, Grompe MT, et al. Human islets contain four distinct subtypes of β cells. Nat Commun. 2016;7:11756. https://doi.org/10.1038/ncomms11756.
Johnston NR, Mitchell RK, Haythorne E, Pessoa MP, Semplici F, Ferrer J, et al. Beta cell hubs dictate pancreatic islet responses to glucose. Cell Metab. 2016;24:389–401. https://doi.org/10.1016/j.cmet.2016.06.020.
Gutierrez GD, Gromada J, Sussel L. Heterogeneity of the pancreatic beta cell. Front Genet. 2017;8:3853. https://doi.org/10.3389/fgene.2017.00022.
Ackermann AM, Wang Z, Schug J, Naji A, Kaestner KH. Integration of ATAC-seq and RNA-seq identifies human alpha cell and beta cell signature genes. Mol Metab. 2016;5:233–44. https://doi.org/10.1016/j.molmet.2016.01.002.
Farack L, Golan M, Egozi A, Dezorella N, Bahar Halpern K, Ben-Moshe S, et al. Transcriptional heterogeneity of beta cells in the intact pancreas. Dev Cell. 2019;48:115–125.e4. https://doi.org/10.1016/j.devcel.2018.11.001.
Neiman D, Moss J, Hecht M, Magenheim J, Piyanzin S, Shapiro AMJ, et al. Islet cells share promoter hypomethylation independently of expression, but exhibit cell-type-specific methylation in enhancers. Proc Natl Acad Sci U S A. 2017;114:13525–30. https://doi.org/10.1073/pnas.1713736114.
•• Arda HE, Tsai J, Rosli YR, et al. A chromatin basis for cell lineage and disease risk in the human pancreas. Cell Syst. 2018;7:310–322.e4. https://doi.org/10.1016/j.cels.2018.07.007 This study provides an in-depth characterization of the regulatory landscapes of specific pancreatic cell types.
•• Chiou J, Zeng C, Cheng Z, et al. Single cell chromatin accessibility reveals pancreatic islet cell type- and state-specific regulatory programs of diabetes risk. bioRxiv. 2019;71:858. https://doi.org/10.1101/693671 This study provides the first human islet regulatory maps at single-cell resolution and demonstrates that specific T2D variants reside in cell-specific regulatory elements.
Lareau CA, Duarte FM, Chew JG, Kartha VK, Burkett ZD, Kohlway AS, et al. Droplet-based combinatorial indexing for massive-scale single-cell chromatin accessibility. Nat Biotechnol. 2019;37:916–24. https://doi.org/10.1038/s41587-019-0147-6.
Grosselin K, Durand A, Marsolier J, Poitou A, Marangoni E, Nemati F, et al. High-throughput single-cell ChIP-seq identifies heterogeneity of chromatin states in breast cancer. Nat Genet. 2019;51:1060–6. https://doi.org/10.1038/s41588-019-0424-9.
Kaya-Okur HS, Wu SJ, Codomo CA, Pledger ES, Bryson TD, Henikoff JG, et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat Commun. 2019;10:1930. https://doi.org/10.1038/s41467-019-09982-5.
• McCarthy MI. Painting a new picture of personalised medicine for diabetes. Diabetologia. 2017;60:793–9. https://doi.org/10.1007/s00125-017-4210-x This review proposes the ‘palette’ model to describe the multitude of possible diabetes disease trajectories.
Marchetti P, Schulte AM, Marselli L, Schoniger E, Bugliani M, Kramer W, et al. Fostering improved human islet research: a European perspective. Diabetologia. 2019;62:212–1516. https://doi.org/10.1007/s00125-019-4911-4.
Dwivedi OP, Lehtovirta M, Hastoy B, et al. Loss of ZnT8 function protects against diabetes by enhanced insulin secretion. Nat Genet. 2019;51:1596–1606. https://doi.org/10.1038/s41588-019-0513-9.
Thomsen SK, Ceroni A, van de Bunt M, Burrows C, Barrett A, Scharfmann R, et al. Systematic functional characterization of candidate causal genes for type 2 diabetes risk variants. Diabetes. 2016;65:3805–11. https://doi.org/10.2337/db16-0361.
Fang Z, Weng C, Li H, Tao R, Mai W, Liu X, et al. Single-cell heterogeneity analysis and CRISPR screen identify key β-cell-specific disease genes. Cell Rep. 2019;26:3132–3144.e7. https://doi.org/10.1016/j.celrep.2019.02.043.
•• Balboa D, Prasad RB, Groop L, Otonkoski T. Genome editing of human pancreatic beta cell models: problems, possibilities and outlook. Diabetologia. 2019;50:1505. https://doi.org/10.1007/s00125-019-4908-z This review provides a very up-to-date and critical overview on applying genome editing in β cells to model diabetes genetic risk variants, 1336.
Manning AK, Hivert M-F, Scott RA, et al. A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat Genet. 2012;44:659–69. https://doi.org/10.1038/ng.2274.
Gloyn AL, Drucker DJ. Precision medicine in the management of type 2 diabetes. Lancet Diabetes Endocrinol. 2018;6:891–900. https://doi.org/10.1016/S2213-8587(18)30052-4.
• Udler MS, Kim J, Grotthuss v M, et al. Type 2 diabetes genetic loci informed by multi-trait associations point to disease mechanisms and subtypes: a soft clustering analysis. PLoS Med. 2018;15:e1002654. https://doi.org/10.1371/journal.pmed.1002654 This study reveals that T2D patients can be clustered into different disease mechanisms.
Kycia I, Wolford BN, Huyghe JR, Fuchsberger C, Vadlamudi S, Kursawe R, et al. A common type 2 diabetes risk variant potentiates activity of an evolutionarily conserved islet stretch enhancer and increases C2CD4A and C2CD4B expression. Am J Hum Genet. 2018;102:620–35. https://doi.org/10.1016/j.ajhg.2018.02.020.
Fogarty MP, Cannon ME, Vadlamudi S, Gaulton KJ, Mohlke KL. Identification of a regulatory variant that binds FOXA1 and FOXA2 at the CDC123/CAMK1D type 2 diabetes GWAS locus. PLoS Genet. 2014;10:e1004633. https://doi.org/10.1371/journal.pgen.1004633.
Fogarty MP, Panhuis TM, Vadlamudi S, Buchkovich ML, Mohlke KL. Allele-specific transcriptional activity at type 2 diabetes-associated single nucleotide polymorphisms in regions of pancreatic islet open chromatin at the JAZF1 locus. Diabetes. 2013;62:1756–62. https://doi.org/10.2337/db12-0972.
Tuomi T, Nagorny CLF, Singh P, Bennet H, Yu Q, Alenkvist I, et al. Increased melatonin signaling is a risk factor for type 2 diabetes. Cell Metab. 2016;23:1067–77. https://doi.org/10.1016/j.cmet.2016.04.009.
Gaulton KJ, Nammo T, Pasquali L, Simon JM, Giresi PG, Fogarty MP, et al. A map of open chromatin in human pancreatic islets. Nat Genet. 2010;42:255–9. https://doi.org/10.1038/ng.530.
Acknowledgements
I would like to thank Goutham Atla (Centre for Genomic Regulation), Anthony Beucher and Vahid Elyasigomari (Imperial College London) for the valuable input on the original manuscript and all the human islet researchers who have deposited their research articles in bioRxiv ahead of publication, enabling this review to be as up-to-date as possible.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Inês Cebola declares that she has no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by the author.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Genetics
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Cebola, I. Pancreatic Islet Transcriptional Enhancers and Diabetes. Curr Diab Rep 19, 145 (2019). https://doi.org/10.1007/s11892-019-1230-6
Published:
DOI: https://doi.org/10.1007/s11892-019-1230-6