Abstract
Purpose
The aim of this study was to determine the expression and function of miR-509-5p in renal cell carcinoma (RCC).
Materials and methods
In this research, we have conducted quantitative real-time polymerase chain reaction (qRT-PCR) assay to determine the expression level of miR-509-5p in tissues and plasma from renal cell carcinoma patients. We preformed in vitro migration scratch assay, flow cytometry analysis and 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay to determine the exact function of miR-509-5p.
Results
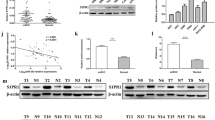
We evaluated the expression level of miR-509-5p in RCC tissues and paired adjacent normal tissues from 42 patients and found that miR-509-5p expression in 42 RCC specimens was significantly down-regulated compared to that in adjacent normal tissue. Furthermore, the level of miR-509-5p in RCC patients’ plasma was significantly lower than that in control plasma. In addition, the overexpression of miR-509-5p suppressed the proliferation of RCC cell (786-0), induced cell apoptosis and inhibited cell migration in vitro.
Conclusion
In this study, we have shown that miR-509-5p played an important role in RCC by inhibiting cell proliferation and migration and by promoting cell apoptosis. In addition, miR-509-5p expression was significantly lower in RCC patient plasma compared to that in normal individuals.


Similar content being viewed by others
References
Rini BI, Campbell SC, Escudier B (2009) Renal cell carcinoma. Lancet 373:1119–1132
Slaby O, Svoboda M, Michalek J, Vyzula R (2009) DNA and microRNA microarray technologies in diagnostics and prediction for patients with renal cell carcinoma. Klin Onkol 22:202–209
Kovacs G (1996) Molecular genetics of human renal cell tumours. Nephrol Dial Transpl 11(Suppl 6):62–65
Wang GK, Zhu JQ, Zhang JT et al (2010) Circulating microRNA: a novel potential biomarker for early diagnosis of acute myocardial infarction in humans. Eur Heart J 31:659–666
Caporali A, Meloni M, Vollenkle C et al (2011) Deregulation of microRNA-503 contributes to diabetes mellitus-induced impairment of endothelial function and reparative angiogenesis after limb ischemia. Circulation 123:282–291
Garzon R, Calin GA, Croce CM (2009) MicroRNAs in Cancer. Annu Rev Med 60:167–179
Kim VN (2005) MicroRNA biogenesis: coordinated cropping and dicing. Nat Rev Mol Cell Biol 6:376–385
Ha TY (2011) MicroRNAs in human diseases: from cancer to cardiovascular disease. Immune Netw 11:135–154
Gottardo F, Liu CG, Ferracin M et al (2007) Micro-RNA profiling in kidney and bladder cancers. Urol Oncol 25:387–392
Huang Y, Dai Y, Yang J et al (2009) Microarray analysis of microRNA expression in renal clear cell carcinoma. Eur J Surg Oncol 35:1119–1123
Chow TF, Mankaruos M, Scorilas A et al (2010) The miR-17-92 cluster is over expressed in and has an oncogenic effect on renal cell carcinoma. J Urol 183:743–751
Schaefer A, Stephan C, Busch J, Yousef GM, Jung K (2010) Diagnostic, prognostic and therapeutic implications of microRNAs in urologic tumors. Nat Rev Urol 7:286–297
White NM, Yousef GM (2010) MicroRNAs: exploring a new dimension in the pathogenesis of kidney cancer. BMC Med 8:65
White NM, Bui A, Mejia-Guerrero S et al (2010) Dysregulation of kallikrein-related peptidases in renal cell carcinoma: potential targets of miRNAs. Biol Chem 391:411–423
Zhou B, Wu H, Li W, Liu W, Luo D (2011) Microarray analysis of microRNA expression in skin of Xpc (+)/(−) mice and wild-type mice. Ir J Med Sci 180:721–726
Heinzelmann J, Henning B, Sanjmyatav J et al (2011) Specific miRNA signatures are associated with metastasis and poor prognosis in clear cell renal cell carcinoma. World J Urol 29:367–373
Porkka KP, Pfeiffer MJ, Waltering KK, Vessella RL, Tammela TL, Visakorpi T (2007) MicroRNA expression profiling in prostate cancer. Cancer Res 67:6130–6135
Ljungberg B (2007) Prognostic markers in renal cell carcinoma. Curr Opin Urol 17:303–308
Neal CS, Michael MZ, Rawlings LH, Van der Hoek MB, Gleadle JM (2010) The VHL-dependent regulation of microRNAs in renal cancer. BMC Med 8:64
Saini S, Yamamura S, Majid S et al (2011) MicroRNA-708 induces apoptosis and suppresses tumorigenicity in renal cancer cells. Cancer Res 71:6208–6219
Boguslawska J, Wojcicka A, Piekielko-Witkowska A, Master A, Nauman A (2011) MiR-224 targets the 3′UTR of type 1 5′-iodothyronine deiodinase possibly contributing to tissue hypothyroidism in renal cancer. PLoS One 6:e24541
Filipowicz W, Bhattacharyya SN, Sonenberg N (2008) Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet 9:102–114
Mitchell PS, Parkin RK, Kroh EM et al (2008) Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA 105:10513–10518
Chim SS, Shing TK, Hung EC et al (2008) Detection and characterization of placental microRNAs in maternal plasma. Clin Chem 54:482–490
Hunter MP, Ismail N, Zhang X et al (2008) Detection of microRNA expression in human peripheral blood microvesicles. PLoS One 3:e3694
Zhu W, Liu X, He J, Chen D, Hunag Y, Zhang YK (2011) Overexpression of members of the microRNA-183 family is a risk factor for lung cancer: a case control study. BMC Cancer 11:393
Liu J, Gao J, Du Y et al (2012) Combination of plasma microRNAs with serum CA19-9 for early detection of pancreatic cancer. Int J Cancer 131:683–691
Lakomy R, Sana J, Hankeova S et al (2011) MiR-195, miR-196b, miR-181c, miR-21 expression levels and O-6-methylguanine-DNA methyltransferase methylation status are associated with clinical outcome in glioblastoma patients. Cancer Sci 102:2186–2190
Acknowledgments
This work was supported by the Natural Science Foundation of Hubei (No. 2011cdb513).
Conflict of interest
The authors have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
W.-B. Zhang and Z.-Q. Pan contributed equally to this work.
Rights and permissions
About this article
Cite this article
Zhang, W.B., Pan, Z.Q., Yang, Q.S. et al. Tumor suppressive miR-509-5p contributes to cell migration, proliferation and antiapoptosis in renal cell carcinoma. Ir J Med Sci 182, 621–627 (2013). https://doi.org/10.1007/s11845-013-0941-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11845-013-0941-y