Abstract
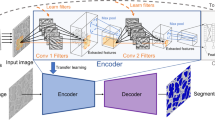
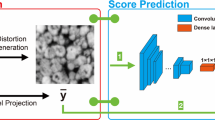
Four state-of-the-art Deep Learning-based Convolutional Neural Networks (DCNN) were applied to automate the semantic segmentation of a 3D Transmission x-ray Microscopy (TXM) nanotomography image data. The standard U-Net architecture as baseline along with UNet++, PSPNet, and DeepLab v3+ networks were trained to segment the microstructural features of an AA7075 micropillar. A workflow was established to evaluate and compare the DCNN prediction dataset with the manually segmented features using the Intersection of Union (IoU) scores, time of training, confusion matrix, and visual assessment. Comparing all model segmentation accuracy metrics, it was found that using pre-trained models as a backbone along with appropriate training encoder–decoder architecture of the Unet++ can robustly handle large volumes of x-ray radiographic images in a reasonable amount of time. This opens a new window for handling accurate and efficient image segmentation of in situ time-dependent 4D x-ray microscopy experimental datasets.









Similar content being viewed by others
References
Q. Zhang, S. Niverty, A.S.S. Singaravelu, J.J. Williams, E. Guo, T. Jing, and N. Chawla, Mater. Charact. 148, 52. (2019).
J.M. Yu, N. Wanderka, A. Rack, R. Daudin, E. Boller, H. Markötter, A. Manzoni, F. Vogel, T. Arlt, I. Manke, and J. Banhart, Acta Mater. 129, 194. (2017).
Q. Krol, and H. Löwe, Acta Mater. 151, 478. (2018).
N. Limodin, L. Salvo, E. Boller, M. Suéry, M. Felberbaum, S. Gailliègue, and K. Madi, Acta Mater. 57, 2300. (2009).
C.S. Kaira, V. De Andrade, S. Singh, C. Kantzos, A. Kirubanandham, F. De Carlo, and N. Chawla, Adv. Mater. 29, 1703482. (2017).
E. Boulard, C. Denoual, A. Dewaele, A. King, Y. Le Godec, and N. Guignot, Acta Mater. 192, 30. (2020).
S. Niverty, C. Kale, K.N. Solanki, and N. Chawla, Corros. Sci. 185, 109429. (2021).
M.B. Kelly, S. Niverty, and N. Chawla, J. Alloys Compds. 818, 152918. (2020).
A.S.S. Singaravelu, J.J. Williams, H.D. Goyal, S. Niverty, S.S. Singh, T.J. Stannard, X. Xiao, and N. Chawla, Metall. Mater. Trans. A 51, 28. (2020).
M.B. Kelly, S. Niverty, and N. Chawla, Acta Mater. 189, 118. (2020).
V. Mazars, O. Caty, G. Couégnat, A. Bouterf, S. Roux, S. Denneulin, J. Pailhès, and G.L. Vignoles, Acta Mater. 140, 130. (2017).
S. Niverty, (2020).
S.S. Singh, T.J. Stannard, X. Xiao, and N. Chawla, JOM 69, 1404. (2017).
J. Samei, C. Pelligra, M. Amirmaleki, and D.S. Wilkinson, Mater. Lett. 269, 127664. (2020).
A. Isaac, F. Sket, W. Reimers, B. Camin, G. Sauthoff, and A.R. Pyzalla, Mater. Sci. Eng. A 478, 108. (2008).
T. Lacondemine, J. Réthoré, É. Maire, F. Célarié, P. Houizot, C. Roux-Langlois, C.M. Schlepütz, and T. Rouxel, Acta Mater. 179, 424. (2019).
H.A. Bale, A. Haboub, A.A. MacDowell, J.R. Nasiatka, D.Y. Parkinson, B.N. Cox, D.B. Marshall, and R.O. Ritchie, Nature Mater. 12, 40. (2013).
A.S.S. Singaravelu, J.J. Williams, J. Ruppert, M. Henderson, C. Holmes, and N. Chawla, J. Mater. Sci. (2020).
B.M. Patterson, L. Kuettner, T. Shear, K. Henderson, M.J. Herman, A. Ionita, N. Chawla, J. Williams, T. Sun, K. Fezzaa, X. Xiao, and C. Welch, J. Mater. Sci. 55, 11353. (2020).
C.S. Kaira, C.R. Mayer, V. De Andrade, F. De Carlo, and N. Chawla, Microsc. Microanal. 22, 808. (2016).
X. Yang, D. Gürsoy, C. Phatak, V. De Andrade, E.B. Gulsoy, and F. De Carlo, Microsc. Microanal. 22, 240. (2016).
C.S. Kaira, V. De Andrade, S.S. Singh, C. Kantzos, F. De Carlo, and N. Chawla, Microsc. Microanal. 23, 2220. (2017).
C. Shashank Kaira, X. Yang, V. De Andrade, F. De Carlo, W. Scullin, D. Gursoy, and N. Chawla, Mater. Charact. 142, 203. (2018).
Y. Wang, J. Gao, Y. Ren, V. De Andrade, and A.J. Shahani, JOM 72, 2965. (2020).
L.J. Ausderau, H.J. Gonzalez Malabet, J.R. Buckley, V. De Andrade, Y. Liu, and G.J. Nelson, JOM 69, 1478. (2017).
V. De Andrade, A. Deriy, M.J. Wojcik, D. Gürsoy, D. Shu, K. Fezzaa and F. De Carlo, SPIE Newsroom, (2016).
D. Gürsoy, F. De Carlo, X. Xiao, and C. Jacobsen, J. Synchrotron Radiat. 21, 1188. (2014).
J.J. Williams, Z. Flom, A.A. Amell, N. Chawla, X. Xiao, and F. De Carlo, Acta Mater. 58, 6194. (2010).
J.C.E. Mertens, J.J. Williams, and N. Chawla, Nucl. Instrum. Methods Phys. Res. A 800, 82. (2015).
C. Gobert, A. Kudzal, J. Sietins, C. Mock, J. Sun, and B. McWilliams, Add. Manuf. 36, 101460. (2020).
A. Kumar, and G.K.H. Pang, IEEE Trans. Syst. Man Cybernet. B 32, 553. (2002).
P.I. Guntoro, G. Tiu, Y. Ghorbani, C. Lund, and J. Rosenkranz, Miner. Eng. 142, 105882. (2019).
C. Bishop, Pattern Recognition and Machine Learning (Springer, New York, 2006).
T.F. Gonzalez, Handbook of Approximation Algorithms and Metaheuristics 1 (Taylor & Francis, London, 2007).
B. Ma, X. Ban, H. Huang, Y. Chen, W. Liu, and Y. Zhi, Symmetry 10, 107. (2018).
T. Stan, Z.T. Thompson, and P.W. Voorhees, Mater. Charact. 160, 110119. (2020).
A. Tekawade, B.A. Sforzo, K.E. Matusik, A.L. Kastengren and C.F. Powell, in Developments in X-Ray Tomography XII, ed. B. Müller and G. Wang (SPIE, 2019), p. 67.
S. Evsevleev, S. Paciornik, and G. Bruno, Adv. Eng. Mater. 22, 1. (2020).
D. Chen, D. Guo, S. Liu, and F. Liu, Symmetry 12, 639. (2020).
X. Yang, F. De Carlo, C. Phatak, and D. Gürsoy, J. Synchrotron Radiat. 24, 469. (2017).
X. Yang, V. De Andrade, W. Scullin, E.L. Dyer, N. Kasthuri, F. De Carlo, and D. Gürsoy, Sci. Rep. 8, 2575. (2018).
I. Rizwan, I. Haque, and J. Neubert, Inform. Med. Unlock. 18, 100297. (2020).
Z. Zhou, R. Siddiquee, N. Tajbakhsh, and J. Liang, 1 (n.d.).
W. Zhang, X. He, W. Li, Z. Zhang, Y. Luo, L. Su, and P. Wang, Image Vis. Comput. 93, 103824. (2020).
X. Liu, Z. Deng, and Y. Yang, Artif. Intell. Rev. 52, 1089. (2019).
Q. Liu, A.B. Salberg and R. Jenssen, International Geoscience and Remote Sensing Symposium (IGARSS) 2018-July, 6943 (2018).
I. Goodfellow, Y. Benjio, and A. Courville, Deep Learning (MIT, Cambridge, 2016).
Z. Zhou, M. M. Rahman Siddiquee, N. Tajbakhsh, and J. Liang, in (2018), pp. 3–11.
L. C. Chen, Y. Zhu, G. Papandreou, F. Schroff and H. Adam, Proceedings of the European Conference on Computer Vision (ECCV) 801 (2018).
L.C. Chen, G. Papandreou, I. Kokkinos, K. Murphy, and A.L. Yuille, IEEE Trans. Pattern Anal. Mach. Intell. 40, 834. (2018).
H. Zhao, J. Shi, X. Qi, X. Wang and J. Jia, Proceedings - 30th IEEE Conference on Computer Vision and Pattern Recognition, CVPR 2017 2017-Jan, 6230 (2017).
S.S. Singh, C. Schwartzstein, J.J. Williams, X. Xiao, F. De Carlo, and N. Chawla, J. Alloys Compds. 602, 163. (2014).
M. Gao, C.R. Feng, and R.P. Wei, Metall. Mater. Trans. A 29, 1145. (1998).
S. Dey, M.K. Gunjan, and I. Chattoraj, Corros. Sci. 50, 2895. (2008).
W. Tian, S. Li, B. Wang, J. Liu, and M. Yu, Corros. Sci. 113, 1. (2016).
O. Ronneberger, P. Fischer, and T. Brox, in (2015), pp. 234–241.
G. Shi, S. Guan and X. Yang, (2011).
K. Jamart, Z. Xiong, G. D. Maso Talou, M. K. Stiles and J. Zhao, Front. Cardiovasc. Med., 7, (2020).
C.S. Kaira, C. Kantzos, J.J. Williams, V. De Andrade, F. De Carlo, and N. Chawla, Acta Mater. 144, 419. (2018).
J. Bertels, T. Eelbode, M. Berman, D. Vandermeulen, F. Maes, R. Bisschops,and M. B. Blaschko, in (2019), pp. 92–100.
S. Jadon, 2020 IEEE Conference on Computational Intelligence in Bioinformatics and Computational Biology, CIBCB 2020 (2020).
Acknowledgements
H.T., S.N., and N.C. are grateful for financial support from the Office of Naval Research (ONR) under Contract No. N00014-10-1-0350 (Dr. W. Mullins, Program Manager). We acknowledge the use of resources at Beamline 32-ID-C of the Advanced Photon Source, a U.S. Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Torbati-Sarraf, H., Niverty, S., Singh, R. et al. Machine-Learning-based Algorithms for Automated Image Segmentation Techniques of Transmission X-ray Microscopy (TXM). JOM 73, 2173–2184 (2021). https://doi.org/10.1007/s11837-021-04706-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11837-021-04706-x