Abstract
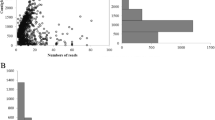
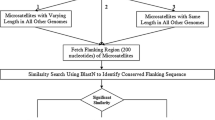
The genomic resources of Porphyra yezoensis expressed sequence tags (ESTs) were utilized to identify simple sequence repeats (SSRs), or microsatellites. This method took the advantage of using ESTs and microsatellites either for the establishment of gene identities or for the acquisition of high polymorphism. The microsatellites can be used as gene markers when microsatellites are tagged to genes. Revealed by bioinformatics analysis, 1162 out of 21 954 ESTs contained microsatellites and cluster analysis indicated that 984 of these ESTs fell into 112 contigs, while the other 178 ESTs were singletons. A total of 290 unique SSR-containing genes were identified. The AAC SSRs were the most populous type of microsatellites. GC-rich microsatellites were predominant among all the microsatellites.
Similar content being viewed by others
References
Asamizu, E., M. Nakajima, Y. Kitade, N. Saga, Y. Nakamura, et al., 2003. Comparison of RNA expression profiles between the two generations of Porphyra yezoensis (Rhodophyta), based on expressed sequence tag frequency analysis. J. Phycol., 39: 923–930.
Blair, M. W., F. Pedraza, H. F. Buendia, Gaitán-Solís, S. E. Beede, et al., 2003. Development of a genome-wide anchored microsatellite map for common bean (Phaseolus vulgaris L.). Theor. Appl. Genet., 107: 1362–1374.
Cagigas, M. E., E. Vazquez, G. Blanco, and J. A. Sanchez, 1999. Combined assessment of genetic variability in populations of brown trout (Salmo trutta L.) based on allozymes, microsatellites, and rapid markers. Mar. Biotech., 1: 286–296.
Delghandi, M., A. Mortensen, and J. I. Westgaard, 2003. Simultaneous analysis of six microsatellite markers in Atlantic cod (Gadus morhua): a novel multiplex assay system for use in selective breeding studies. Mar. Biotech., 5: 141–148.
Eujay, I., M. E. Sorrells, M. Baum, P. Wolters, and W. Powell, 2002. Isolation of EST-derived microsatellite markers for genotyping the A and B genomes of wheat. Theor. Appl. Genet., 104: 399–407.
Gabor, T., G. Zoltan, and J. Jerzy, 2000. Microsatellites in different eukaryotic genomes: survey and analysis. Genome Res., 10: 967–981.
Gupta, P. K., and R. K. Varshney, 2000. The development and use of microsatellite markers for genetic analysis and plant breeding with emphasis on bread wheat. Euphytica, 113: 163–185.
Han, K., L. Li, G. M. Leclere, A. M. Hays, and B. Ely, 2000. Isolation and characterization of microsatellite loci for striped bass (Morone saxatilis). Mar. Biotechnol., 2: 405–408.
Kantety, R. V., R. M. La, D. E. Matthews, and M. E. Sorrells, 2002. Data mining for simple sequence repeats in expressed sequence tags from barley, maize, rice, sorghum and wheat. Plant. Mol. Biol., 48: 501–510.
Liu, B. Q., Q. G. Zeng, Q. J. Luo, Y. J. Wang, and S. H. Li, 2005. Isolation of microstatellite loci from dbEST of algae Porphyra yezoensis and primer amplification of interspecies transfer. Oceanol. Limnol. Sinica, 36(3): 248–254.
Liu, Z. J., G. Tan, H. Kucuktas, P. Li, A. Karsi, et al., 1999. High levels of conservation at microsatellite loci among Ictalurid catfishes. J. Hered., 90: 307–312.
Morgante, M., and A. M. Olivieri, 1993. PCR-Amplified microsatellites as markers in plant genetics. Plant J., 3: 175–182.
Pang, G. X., G. C. Wang, S. N. Hu, and C. K. Tseng, 2005. Acquirement and analysis of expressed sequence tags of filamentous sporophyte of Porphyra haitanensis. Oceanol. Limnol. Sinica, 36(5): 452–457.
Powell, W., G. C. Machray, and J. Provan, 1996. Polymorphism revealed by simple sequence repeats. Trends. Plant Sci., 1: 215–222.
Sahoo, D., X. R. Tang, and C. Yarish, 2002. Porphyra — the economic seaweed as a new experimental system. Curr. Sci., 83(11): 1313–1316.
Sekino, M., M. Hamaguchi, F. Aranishi, and K. Okoshi, 2003. Development of novel microsatellite DNA markers from the Pacific oyster Crassostrea gigas. Mar. Biotechnol., 5: 227–233.
Temnykh, S., C. De, A. Lukashova, L. Lipovich, S. Cartinhour, et al., 2001. Computational and experimental analysis of microsatellites in rice (Oryza sativa L.): Frequency, length variation, transposon associations, and genetic marker potential. Genome Res., 11: 1441–1452.
Thiel, T., W. Michalek, R. K. Varshney, and A. Graner, 2003. Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor. Appl. Genet., 106: 411–422.
Thoquet, P., M. Ghrardi, E. P. Journet, A. Kereszt, J. M. An, et al., 2002. The molecular genetic linkage map of the model legume Medicago truncatula: an essential tool for comparative legume genomics and the isolation of agronomically important genes. BMC Plant Biol., 2: 1–13.
Varshney, R. K., A. Graner, and M. E. Sorrells, 2005. Genic microsatellite markers in plants: features and applications. Trends Biotech., 23(1): 48–55.
Varshney, R. K., T. Thiel., N. Stein, P. Langridge, and A. Graner, 2002. In silico analysis of frequency and distribution of microsatellites in ESTs of some cereal species. Cell Mol. Biol. Lett., 7: 537–546.
Xu, M. J., Y. X. Mao, X. C. Zhang, X. J. Zhou, Z. H. Sui, et al., 2006. Bioinformatic analysis of expressed sequence tags from sporophyte of Porphyra yezoensis. Prog. Nat. Sci., 16(1): 39–49.
Yang, G. P., H. S. Shen, and P. Xu, 2002. Analysis of expressed sequence tags of filamentous sporophyte of Porphyra yezoensis. High Tech. Lett., 2: 93–97.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, M., Hu, J., Zhuang, Y. et al. In Silico screening for microsatellite markers from expressed sequence tags of Porphyra yezoensis (Bangiales, Rhodophyta). J Ocean Univ. China 6, 161–166 (2007). https://doi.org/10.1007/s11802-007-0161-z
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s11802-007-0161-z