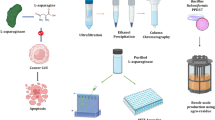
Abstract
A highly potent glycolipid was isolated from a crude oil-contaminated soil bacterium Bacillus licheniformis SV1 (NCBI GenBank Acc. No. KX130852) when grown on a modified mineral salt medium supplemented with 2% oleic acid. The maximum reduction in surface tension of cell-free broth from 71 ± 0.812 to 25.919 ± 0.984 mN/m with 89 ± 0.346% emulsification activity was recorded after 120 h of growth. Glycolipid was purified using chromatographic techniques and the presence of aliphatic chain (C–H stretch) and OH-band was revealed by NMR, GC–MS and FTIR analysis. Stability of glycolipid up to 250 °C and its complete decomposition at 507 °C was recorded by thermogravimetric analysis (TGA). MTT assay (IC50 = 0.473 ± 0.048 mg/ml) along with 4′,6-diamidino-2-phenylindole (DAPI) and acridine orange and ethidium bromide (AO/EtBr) staining validated that glycolipid induces cell apoptosis in human prostate cancer cell line (PC-3). Furthermore, potent anti-cancerous compounds such as 9-octadecanoic acid (22.55%), linoleic acid methyl ester (2.2%) and palmitic acid (1.18%) were also detected in the GC–MS spectra of the purified glycolipid.






Similar content being viewed by others
Abbreviations
- MSM:
-
Mineral salt medium
- AO/EtBr:
-
Acridine orange and ethidium bromide
- DAPI:
-
4′,6-Diamidino-2-phenylindole
References
Antoniou E, Fodelianakis S, Korkakaki E, Kalogerakis N. Biosurfactant production from marine hydrocarbon-degrading consortia and pure bacterial strains using crude oil as carbon source. Front Microbiol. 2015;6:1–14.
Gudiña EJ, Pereira JFB, Costa R, Coutinho JAP, Teixeira JA, Rodrigues LR. Biosurfactant-producing and oil-degrading Bacillus subtilis strains enhance oil recovery in laboratory sand-pack columns. J Hazard Mater. 2013;261:106–13.
Batista SB, Mounteer AH, Amorim FR, Totola MR. Isolation and characterization of biosurfactant/bioemulsifier-producing bacteria from petroleum contaminated sites. Bioresour Technol. 2006;97:868–75.
Uzoigwe C, Burgess JG, Ennis CJ, Rahman PKSM. Bioemulsifiers are not biosurfactants and require different screening approaches. Front Microbiol. 2015;6:1–6.
Banat IM, Satpute SK, Cameotra SS, Patil R, Nyayanit NV. Cost effective technologies and renewable substrates for biosurfactants production. Front Microbiol. 2014;5:1–18.
Rahman KSM, Rahman TJ, Kourkoutas Y, Petsas I, Marchant R, Banat IM. Enhanced bioremediation of n-alkane in petroleum sludge using bacterial consortium amended with rhamnolipid and micronutrients. Bioresour Technol. 2003;90:159–68.
Christova N, Tuleva B, Kril A, Georgieva M, Konstantinov S, Terziyski I, Nikolova B, Stoineva I. Chemical structure and in vitro antitumor activity of rhamnolipids from Pseudomonas aeruginosa BN10. Appl Biochem Biotechnol. 2013;170:676–89.
Jiang L, Shen C, Long X, Zhang G, Meng Q. Rhamnolipids 354 elicit the same cytotoxic sensitivity between cancer cell and normal cell by reducing surface tension of culture medium. Appl Microbiol Biotechnol. 2014;98:10187–96.
Gudiña EJ, Teixeira JA, Rodrigues LR. Biosurfactants produced by marine microorganisms with therapeutic applications. Mar Drugs. 2016;14(2):1–15.
Jemal A, Siegel R, Ward E, Xu J. Cancer statistics. CA Cancer J Clin. 2010;60:277–300.
Pemmaraju SC, Sharma D, Singh N, Panwar R, Cameotra SS, Pruthi V. Production of microbial surfactants from oily sludge contaminated soil by Bacillus subtilis DSVP23. Appl Biochem Biotechnol. 2012;167:1119–31.
Keskin NOS, Han D, Ozkan AD, Angun P, Umu OCO, Tekinay T. Production and structural characterization of biosurfactant produced by newly isolated Staphylococcus xylosus STF1 from petroleum contaminated soil. J Petrol Sci Eng. 2015;133:689–94.
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol. 2013;30:2725–9.
Mukherjee S, Das P, Sivapathasekaran C, Sen R. Enhanced production of biosurfactant by a marine bacterium on statistical screening of nutritional parameters. Bio Eng J. 2008;42:254–60.
Sim L, Ward OP, Li Z-Y. Production and characterization of a biosurfactant isolated from Pseudomonas aeruginosa UW-1. J Ind Microbiol Biotechnol. 1997;19:232–8.
Heyd M, Kohnert A, Tan TH, Nusser M, Kirschhöfer F, Brenner-Weiss G, Franzreb M, Berensmeier S. Development and trends of biosurfactant analysis and purification using rhamnolipids as an example. Anal Bioanal Chem. 2008;391:1579–90.
Abbasi H, Hamedi MM, Lotfabad TB, Zahiri HS, Sharafi H, Masoomi F, Moosavi- Movahedi AA, Ortiz A, Amanlou M, Noghabi KA. Biosurfactant-producing bacterium, Pseudomonas aeruginosa MA01 isolated from spoiled apples: physicochemical and structural characteristics of isolated biosurfactant. J Biosci Bioeng. 2012;113:211–9.
Lan G, Fan Q, Liu Y, Chen C, Li G, Liu Y, Yin X. Rhamnolipid production from waste cooking oil using Pseudomonas SWP-4. Biochem Eng J. 2015;101:44–54.
Tai M, Stephanopoulos G. Engineering the push and pull of lipid biosynthesis in oleaginous yeast Yarrowia lipolytica for biofuel 382 production. Metab Eng. 2013;15:1–9.
Claus H. Rapid identification of fatty acid methyl esters using a multidimensional gas chromatography mass spectroscopy database. J Chromatogr A. 2008;1177(1):159–69.
Jain R, Mody K, Mishra A. Production and structural characterization of biosurfactant produced by an alkaliphilic bacterium Klebsiella sp.: Evaluation of different carbon sources. Colloids Surf B. 2013;108:199–204.
Kumar N, Sharan S, Singh AK, Chakraborty A, Roy P. Anticancer activities of pterostilbene isothiocyanate conjugate in breast cancer cells: involvement of PPARc. PLoS ONE. 2014;9:1–17.
Ribble D, Goldstein NB, Norris DA, Shellman YG. A simple technique for quantifying apoptosis in 96-well plates. BMC Biotechnol. 2005;5:1–7.
Dadrasnia A, Usman MM, Wei KSC, Velappan RD, Jamali H, Mohebali N, Ismail S. Native soil bacterial isolate in Malaysia exhibit promising supplements on degrading organic pollutants. Process Saf Environ. 2016;100:264–71.
Liu B, Liu J, Ju M, Li X, Yu Q. Purification and characterization of biosurfactant produced by Bacillus licheniformis Y-1 and its application in remediation of petroleum contaminated soil. Marine Poll Bull. 2016;107(1):1–6.
Wei YH, Chou CL, Chang JS. Rhamnolipid production by indigenous Pseudomonas aeruginosa J4 originating from petrochemical wastewater. Biochem Eng J. 2005;27:146–54.
Aparna A, Srinikethan G, Smitha H. Production and characterization of biosurfactant produced by a novel Pseudomonas sp. Colloids Surf B. 2012;95:23–9.
Yoo Y, Shin B, Hong J, Lee J, Chee H, Song K, Lee K. Isolation of fatty acids with anticancer activity from Protaetia brevitarsis larva. Arch Pharm Res. 2007;30:361–5.
Lee KW, Lee HJ, Cho HY, Kim YJ. Role of the conjugated linoleic acid in the prevention of cancer. Crit Rev Food Sci. 2007;45:135–44.
Janak T, Radwanska A, Lukaszwicz M. Lipopeptide biosurfactant pseudofactin II induced apoptosis of melanoma A 375 cells by specific interaction with the plasma membrane. PLoS ONE. 2013;8:1–9.
Janek T, Łukaszewicz M, Rezanka T, Krasowska A. Isolation and characterization of two new lipopeptide biosurfactants produced by Pseudomonas fluorescens BD5 isolated from water from the Arctic Archipelago of Svalbard. Bioresour Technol. 2010;101(15):6118–23.
Acknowledgements
Authors are highly grateful to the Department of Science and Technology, INSPIRE (DST, INSPIRE) Govt. of India (Grant No. IF130678) for providing financial assistance to carry out the present work. Authors are also thankful to Institute Instrumentation Centre (IIC), IIT Roorkee and Department of Chemical Engineering, IIT Roorkee for providing the instrumentation facility.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
About this article
Cite this article
Gupta, S., Varshney, R., Jha, R. et al. In Vitro Apoptosis Induction in a Human Prostate Cancer Cell Line by Thermotolerant Glycolipid from Bacillus licheniformis SV1. J Surfact Deterg 20, 1141–1151 (2017). https://doi.org/10.1007/s11743-017-1986-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11743-017-1986-0