Abstract
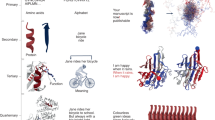
Natural proteins are concatenated amino acids with definite handedness or chirality, with their spatial orientation being preferentially left handed or L-chiral. This paper discusses the biophysics of stereo-chemical perturbation to proteins using D-(α) amino acid and its utility as an additional design alphabet while scripting novel protein structures.






Similar content being viewed by others
References
Allen BD, Mayo SL (2010) An efficient algorithm for multistate protein design based on FASTER. J Comput Chem 31(5):904–916
Anfinsen CB (1973) Principles that govern the folding of protein chains. Science 181(96):223–230
Anfinsen CB, Scheraga HA (1975) Experimental and theoretical aspects of protein folding. Adv Protein Chem 29:205–300
Avbelj F, Baldwin RL (2002) Role of backbone solvation in determining thermodynamic beta propensities of the amino acids. Proc Natl Acad Sci USA 99(3):1309–1313
Avbelj F, Moult J (1995) Role of electrostatic screening in determining protein main chain conformational preferences. Biochemistry 34(3):755–764
Baker D (2010) An exciting but challenging road ahead for computational enzyme design. Protein Sci 19(10):1817–1819
Baldwin RL (2007) Energetics of protein folding. J Mol Biol 371(2):283–301
Billmeyer FW (1984) Textbook of polymer science, 3rd edn. Wiley Interscience, New York
Billot-Klein D, Legrand R, Schoot B, Van Heijenoort J, Gutmann L (1997) Peptidoglycan structure of Lactobacillus casei, a species highly resistant to glycopeptide antibiotics. J Bacteriol 179(19):6208–6212
Bobde V, Beri S, Durani S (1993) Stable type II’ reverse turn-310 helix conformation of Boc-D-Glu-Ala-Gly-Lys-Ala-Leu-OMe in apolar solvents. Tetrahedron 49(24):5397–5406
Brant DA, Flory PJ (1965a) The configuration of random polypeptide chains. i. Experimental results. J Am Chem Soc 87(13):2788–2791. doi:10.1021/ja01091a002
Brant DA, Flory PJ (1965b) The configuration of random polypeptide chains. II. Theory. J Am Chem Soc 87(13):2791–2800. doi:10.1021/ja01091a003
Brueckner H, Becker D, Luepke M (1993) Chirality of amino acids of microorganisms used in food biotechnology. Chirality 5(5):385–392
Chothia C (1992) Proteins. One thousand families for the molecular biologist. Nature 357(6379):543–544
Chou PY, Fasman GD (1974) Prediction of protein conformation. Biochemistry 13(2):222–245
Cooper S, Khatib F, Treuille A, Barbero J, Lee J, Beenen M, Leaver-Fay A, Baker D, Popovic Z, Players F (2010) Predicting protein structures with a multiplayer online game. Nature 466(7307):756–760
Creamer TP, Rose GD (1992) Side-chain entropy opposes alpha-helix formation but rationalizes experimentally determined helix-forming propensities. Proc Natl Acad Sci USA 89(13):5937–5941
Daura X, van Gunsteren WF, Mark AE (1999) Folding–unfolding thermodynamics of a β-heptapeptide from equilibrium simulations. Proteins: structure, function, and bioinformatics 34:269–280. doi:10.1002/(sici)1097-0134(19990215)34:3<269:aid-prot1>3.0.co;2-3
Daura X, Gademann K, Schafer H, Jaun B, Seebach D, van Gunsteren WF (2001) The beta-peptide hairpin in solution: conformational study of a beta-hexapeptide in methanol by NMR spectroscopy and MD simulation. J Am Chem Soc 123(10):2393–2404
Dhanasekaran M, Fabiola F, Pattabhi V, Durani S (1999) A rationally designed turn-helix peptide. J Am Chem Soc 121(23):5575–5576. doi:10.1021/ja984237v
Donohue J (1953) hydrogen bonded helical configurations of the polypeptide chain. Proc Natl Acad Sci USA 39(6):470–478
Durani S (2008) Protein design with l- and d-alpha-amino acid structures as the alphabet. Acc Chem Res 41(10):1301–1308
Eisenberg D (2003) The discovery of the alpha-helix and beta-sheet, the principal structural features of proteins. Proc Natl Acad Sci USA 100(20):11207–11210
Fabiola F, Pattabhi V, Rawale S, Raju EB, Durani S (1997) Configurationally guided peptide conformational motifs: crystal structure of a l[small alpha]d[small beta]l[small beta]d[small alpha] d[small beta]l[small alpha] type hexapeptide fold. Chem Commun 15:1379–1380
Fitzkee NC, Rose GD (2004) Reassessing random-coil statistics in unfolded proteins. Proc Natl Acad Sci USA 101(34):12497–12502
Flory PJ (1969) Statistical Mechanics of Chain Molecules. Interscience, New York
Ghadiri MR, Granja JR, Buehler LK (1994) Artificial transmembrane ion channels from self-assembling peptide nanotubes. Nature 369(6478):301–304
Golovine S, Hecht SM, Dedkova L (2004) Methods of protein synthesis incorporating D-amino acids using modified ribosomes with 23S rRNA gene mutations. WO Patent 2003-US33108; 2004035757
Jack RW, Jung G (1998) Natural peptides with antimicrobial activity. Chimia 52(1–2):48–55
Joshi S, Rana S, Wangikar P, Durani S (2006) Computational design of proteins stereochemically optimized in size, stability, and folding speed. Biopolymers 83(2):122–134
Kohn JE, Millett IS, Jacob J, Zagrovic B, Dillon TM, Cingel N, Dothager RS, Seifert S, Thiyagarajan P, Sosnick TR, Hasan MZ, Pande VS, Ruczinski I, Doniach S, Plaxco KW (2004) Random-coil behavior and the dimensions of chemically unfolded proteins. Proc Natl Acad Sci USA 101(34):12491–12496
Korendovych IV, Senes A, Kim YH, Lear JD, Fry HC, Therien MJ, Blasie JK, Walker FA, Degrado WF (2010) De novo design and molecular assembly of a transmembrane diporphyrin-binding protein complex. J Am Chem Soc 132(44):15516–15518
Kumar A, Ramakrishnan V, Ranbhor R, Patel K, Durani S (2009) Homochiral stereochemistry: the missing link of structure to energetics in protein folding. J Phys Chem B 113(51):16435–16442
Leach AR (2001) Molecular modelling: principles and applications, 2nd edn. Prentice Hall, England
Leach SJ, Nemethy G, Scheraga HA (1966) Computation of the sterically allowed conformations of peptides. Biopolymers 4(4):369–407
Marahiel MA (2009) Working outside the protein-synthesis rules: insights into non-ribosomal peptide synthesis. J Pept Sci 15(12):799–807
Marahiel MA, Essen LO (2009) Chapter 13. Nonribosomal peptide synthetases mechanistic and structural aspects of essential domains. Methods Enzymol 458:337–351
Martinez del Pozo A, Pospischil MA, Ueno H, Manning JM, Tanizawa K, Nishimura K, Soda K, Ringe D, Stoddard B, Petsko GA (1989) Effects of d-serine on bacterial d-amino acid transaminase: accumulation of an intermediate and inactivation of the enzyme. Biochemistry 28(22):8798–8803
Miller WG, Brant DA, Flory PJ (1967) Random coil configurations of polypeptide copolymers. J Mol Biol 23(1):67–80
Mohanraja K, Dhanasekaran M, Kundu B, Durani S (2003) Mechanism-based protein design: attempted “nucleation-condensation” approach to a possible minimal helix-bundle protein. Biopolymers 70(3):355–363
Nanda V, DeGrado WF (2006) Computational design of heterochiral peptides against a helical target. J Am Chem Soc 128(3):809–816
Patel K, Srivastava KR, Durani S (2010) Zinc-finger hydrolase: Computational selection of a linker and a sequence towards metal activation with a synthetic alphabetabeta protein. Bioorg Med Chem 2010:27
Pauling L, Corey RB (1950) Two hydrogen-bonded spiral configurations of the polypeptide chain. J Am Chem Soc 72(11):5349. doi:10.1021/ja01167a545
Pauling L, Corey RB, Branson HR (1951) The structure of proteins; two hydrogen-bonded helical configurations of the polypeptide chain. Proc Natl Acad Sci USA 37(4):205–211
Presta LG, Rose GD (1988) Helix signals in proteins. Science 240(4859):1632–1641
Ramachandran GN, Sasisekharan V (1968) Conformation of polypeptides and proteins. Adv Protein Chem 23:283–438
Ramachandran GN, Ramakrishnan C, Sasisekharan V (1963) Stereochemistry of polypeptide chain configurations. J Mol Biol 7:95–99
Ramakrishnan V, Ranbhor R, Durani S (2005) Simulated folding in polypeptides of diversified molecular tacticity: implications for protein folding and de novo design. Biopolymers 78(2):96–105
Ramakrishnan V, Ranbhor R, Kumar A, Durani S (2006) The link between sequence and conformation in protein structures appears to be stereochemically established. J Phys Chem B 110(18):9314–9323
Rana S, Kundu B, Durani S (2004) Stereospecific peptide folds. A rationally designed molecular bracelet. Chem Commun (Camb) (21):2462–2463
Rana S, Kundu B, Durani S (2005) A small peptide stereochemically customized as a globular fold with a molecular cleft. Chem Commun (Camb) (2):207–209
Rana S, Kundu B, Durani S (2007a) A double catgrip mixed l and d mini protein only 20 residues long. Bioorg Med Chem 15(11):3874–3882
Rana S, Kundu B, Durani S (2007b) A mixed-alpha, beta miniprotein stereochemically reprogrammed to high-binding affinity for acetylcholine. Biopolymers 87(4):231–243
Ranbhor R, Ramakrishnan V, Kumar A, Durani S (2006) The interplay of sequence and stereochemistry in defining conformation in proteins and polypeptides. Biopolymers 83(5):537–545
Richardson JS, Richardson DC (1988) Amino acid preferences for specific locations at the ends of alpha helices. Science 240(4859):1648–1652
Robinson T (1976) D-Amino acids in higher plants. Life Sci 19(8):1097–1102
Rohl CA, Strauss CE, Misura KM, Baker D (2004) Protein structure prediction using Rosetta. Methods Enzymol 383:66–93
Rose GD, Gierasch LM, Smith JA (1985) Turns in peptides and proteins. Adv Protein Chem 37:1–109
Rose GD, Fleming PJ, Banavar JR, Maritan A (2006) A backbone-based theory of protein folding. Proc Natl Acad Sci USA 103(45):16623–16633
Sibanda BL, Thornton JM (1985) Beta-hairpin families in globular proteins. Nature 316(6024):170–174
Sibanda BL, Blundell TL, Thornton JM (1989) Conformation of beta-hairpins in protein structures. A systematic classification with applications to modelling by homology, electron density fitting and protein engineering. J Mol Biol 206(4):759–777
Struthers MD, Cheng RP, Imperiali B (1996) Design of a monomeric 23-residue polypeptide with defined tertiary structure. Science 271(5247):342–345
Szabo G, Urry DW (1979) N-acetyl gramicidin: single-channel properties and implications for channel structure. Science 203(4375):55–57
Tanford C (1968) Protein denaturation. Adv Protein Chem 23:121–282
Urry DW (1971) The gramicidin A transmembrane channel: a proposed pi(L, D) helix. Proc Natl Acad Sci USA 68(3):672–676
van Gunsteren WF, Billeter SR, Hünenberger PH, Krüger P, Mark AE, Scott WRP, Tironi IG (1996) Biomolecular simulation: the GROMOS96 manual and user guide. Vdf Hochschulverlag AG an der ETH Zürich, Zürich
Wallace BA (1998) Recent advances in the high resolution structures of bacterial channels: gramicidin A. J Struct Biol 121(2):123–141
Wallace BA (2000) Common structural features in gramicidin and other ion channels. Bioessays 22(3):227–234
Watson JD, Milner-White EJ (2002a) The conformations of polypeptide chains where the main-chain parts of successive residues are enantiomeric. their occurrence in cation and anion-binding regions of proteins. J Mol Biol 315(2):183–191
Watson JD, Milner-White EJ (2002b) A novel main-chain anion-binding site in proteins: the nest. A particular combination of phi, psi values in successive residues gives rise to anion-binding sites that occur commonly and are found often at functionally important regions. J Mol Biol 315(2):171–182
Weaver TM (2000) The pi-helix translates structure into function. Protein Sci 9(1):201–206
Srinivasan R RIBOSOME V1.0. http://roselab.jhu.edu/~raj/Manuals/ribosome.html
Zhang C, DeLisi C (1998) Estimating the number of protein folds. J Mol Biol 284(5):1301–1305
Acknowledgments
Authors thank Prof. Susheel Durani for the core concept of this manuscript. VR thanks IYBA program of Dept of Biotechnology, Govt. of India for funding. The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kumar, A., Ramakrishnan, V. Creating novel protein scripts beyond natural alphabets. Syst Synth Biol 4, 247–256 (2010). https://doi.org/10.1007/s11693-011-9068-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11693-011-9068-5