Abstract
Fusions of genetic regulatory elements with reporter genes have long been used as tools for monitoring gene expression and have become a major component in synthetic gene circuit implementation. A major limitation of many of these systems is the relatively long half-life of the reporter protein(s), which prevents monitoring both the initiation and the termination of transcription in real-time. Furthermore, when used as components in synthetic gene circuits, the long time constants associated with reporter protein decay may significantly degrade circuit performance. In this study, short half-life variants of LuxA and LuxB from Photorhabdus luminescens were constructed in Escherichia coli by inclusion of an 11-amino acid carboxy-terminal tag that is recognized by endogenous tail-specific proteases. Results indicated that the addition of the C-terminal tag affected the functional half-life of the holoenzyme when the tag was added to luxA or to both luxA and luxB, but modification of luxB alone did not have a significant effect. In addition, it was also found that alteration of the terminal three amino acid residues of the carboxy-terminal tag fused to LuxA generated variants with half-lives of intermediate length in a manner similar to that reported for GFP. This report is the first instance of the C-terminal tagging approach for the regulation of protein half-life to be applied to an enzyme or monomer of a multi-subunit enzyme complex and will extend the utility of the bacterial luciferase reporter genes for the monitoring of dynamic changes in gene expression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Reporter genes are transcriptional fusions of genetic regulatory elements to reporter proteins that provide readily detectable signal outputs. Such reporter proteins include LacZ, chloramphenicol acetyltransferase, β-lactamase (Zlokarnik et al. 1998), firefly luciferase (luc), bacterial luciferase (lux), and fluorescent proteins such as those derived from the green fluorescent protein of Aequoria victoria (e.g. GFP, EGFP, YFP) (Chalfie et al. 1994; Cormack et al. 1996; Andersen et al. 1998; Miyawaki et al. 1997; also reviewed in van der Meer et al. 2004). These gene fusions have been used in detecting xenobiotic compounds (King et al. 1990; DiGrazia et al. 1991), monitoring performance of synthetic gene circuits (Elowitz and Leibler 2000; Hasty et al. 2002; Gardner et al. 2000), in genome-wide expression arrays (Van Dyk et al. 2001), as well as in in vivo imaging studies (Xiong et al. 2005; Kadurugamuwa et al. 2003; Kuklin et al. 2003; Doyle et al. 2004).
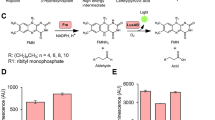
While these reporter proteins have numerous applications, all except the bacterial luciferase system require the additional input of reactants or UV light. The bioluminescent gene cassette comprising the lux system was originally isolated from Vibrio fischeri (Engebrecht et al. 1983). Later, more thermally stable versions of lux were isolated from Photorhabdus (Xenorhabus) luminescens and other species (Schmidt et al. 1989; Szittner and Meighen 1990). In these systems, the heterodimeric bacterial luciferase comprised of LuxAB oxidizes an aliphatic aldehyde substrate and FMNH2 in the presence of diatomic oxygen to release a photon of light at a wavelength of ∼490 nm (reviewed in Meighen (1991)). The remainder of the cassette consisting of the luxC, D and E genes, encode proteins that are responsible for generating and recycling the aldehyde substrate for the reaction.
Another major problem with most reporter systems are the relatively long half-lives of the reporter proteins, which complicate the detection and monitoring of dynamic changes in gene regulation at the transcriptional level. To address these issues, modifications to the protein coding sequences of some of these reporter systems have been made to include amino acids sequences that are recognized by host protein degradation machinery resulting in more rapid turnover of the reporter protein (Andersen et al. 1998; Mateus and Avery 2000; Li et al. 1998; Triccas et al. 2002). Similar tactics have been employed in the engineering of synthetic gene constructs such as the Repressilator (Elowitz and Leibler 2000).
To build on the existing utility of the bacterial lux reporter system, we modified each of the individual luxA and luxB monomers of the bacterial luciferase holoenzyme and tested them both individually and in combination. The modification employed a naturally occurring protein degradation pathway in Escherichia coli. This system, used for removing peptides synthesized from truncated mRNA, involves the addition of an 11-amino acid carboxy-terminal peptide tag (AANDENYALAA) and requires the SsrA RNA (Tu et al. 1995; Keiler et al. 1996). Cytoplasmic proteins containing the tag are rapidly degraded by the ClpP protease in conjunction with either ClpA or ClpX (Gottesman et al. 1998; Weber-Ban et al. 1999; Kim et al. 2000). In addition, the essential protease HflB may also play a role in degradation (Herman et al. 1998). Direct modification of a DNA sequence to encode the 11-amino acid SsrA tag on the 3′ end of the gene to be expressed on the carboxy-terminus of a protein has been previously shown to result in rapid degradation of proteins within the cell (Andersen et al. 1998; Elowitz and Leibler 2000; Herman et al. 1998). Furthermore, alteration of the last three amino acids on the tag has been used to alter the recognition specificity of the proteases, thereby generating a variety of protein half-lives of intermediate lengths (Andersen et al. 1998). Addition of the SsrA tag has also been shown to result in short half-life GFP variants in Pseudomonas and Mycobacterium species (Andersen et al. 1998; Triccas et al. 2002).
We have employed a similar approach for the modification of the bacterial luciferase holoenzyme and show that modification of a single monomer of the holoenzyme, LuxA, is sufficient for altering the functional half-life of the holoenzyme. Conversely, modification of the LuxB monomer alone neither impacts the performance of the holoenzyme nor results in decay of the bioluminescent signal. Based on these results, additional variants of luxA were generated containing modifications of the last three amino acids of the tag, and their half-lives were found to range between that containing the native SsrA tag (abbreviated here as -laa) and the wild type holoenzyme. Finally, the carboxy-terminal tags were cloned into an existing plasmid in a manner that should facilitate retrofitting of other P. luminescens lux-based reporters for the monitoring of rapid changes in transcription, or inclusion into synthetic gene circuits where dynamic signal performance is critical.
Materials and methods
Genetic constructs
Wild type and modified variants of the luxA and luxB genes were amplified by PCR from a vector containing the P. luminescens lux CDABE operon (pCDABE) using the primers listed in Table 1. Abbreviations for the tag (e.g. -aav) are indicated in the name of the construct after the gene being affected. Wild type luxAB and luxABaav were amplified by PCR, gel purified, and cloned into the TOPO-TA cloning vector pCR2.1 (Invitrogen) per the manufacturer’s instructions. For luxA variants, luxBaav, and wild-type luxB, the genes were independently amplified and cloned. The intergenic sequence upstream of the ribosomal binding site between luxA and luxB was modified to create a unique XmaI site and facilitate subsequent construction. The luxA variants were digested overnight with XmaI and XbaI and ligated upstream of the luxB or luxBaav genes in pCR2.1. The wild type luxAB and the newly assembled variants were then each double digested overnight from their respective TA-cloning vectors with BamHI and KpnI, gel purified, and directionally cloned in place of the GFPasv gene of the repressilator reporter plasmid (Elowitz and Leibler 2000). For insertion of the tagged luxA into an existing full lux cassette, the appropriate primary construct was subsequently digested internal to both luxA (AgeI) and luxB (BfuAI) and the tag-bearing fragment was directionally cloned into the similarly digested pCDABE. Cloning reactions were conducted at room temperature for 10 min and included: 100 ng vector DNA, a 2:1 molar ratio of insert to vector DNA, 4 U of T4 DNA ligase (Fisher Scientific, Pittsburgh, PA) and 2 × LigaFast rapid ligation buffer (Promega, Madison, WI) in 25–35 μ l reactions. Restriction enzymes were purchased from New England Biolabs (Beverly, MA), and all digestions were conducted using 5 U of each enzyme and incubated overnight at 37°C except in the case of BfuAI, in which digestion was incubated for 4 h at 50°C, followed by transfer to 37°C and the addition of AgeI.
Strains and growth conditions
Strains used in this study are listed in Table 2. Luria-Bertani media (Sambrook et al. 1989) was used for all experiments with 50 μ g/ml kanamycin as necessary, and incubated at 37°C with shaking at 100 r.p.m. when appropriate.
Bioluminescence assays
Overnight cultures of the four variants of luxAB and a negative control carrying the parent vector with the gene for destabilized green fluorescent protein, Gfpasv, were transferred into triplicate tubes of fresh LB medium containing 50μg/ml kanamycin. Cultures were then grown to an optical density at 546 nm (OD546) of ∼0.6. Initial bioluminescence measurements were made at t = 0 as described below. Subsequently, rifampicin (150 μg/ml) and tetracycline (30 μg/ml) were added to all cultures to block transcription and translation, respectively. For bioluminescence measurements, 1-ml samples were taken immediately (t = 0) and at 30 min intervals thereafter and assayed for optical density at 546 nm (OD546) and bioluminescence. OD546 was measured using a Beckman DU-640B spectrophotometer. Bioluminescence was assayed by first adding 10 μl of N-decanal (Sigma, St. Louis, MO) to the 1-ml samples, inverting 10 times, and measuring light with a SDI Deltatox handheld luminometer. Three light measurements were taken in rapid succession and these values were averaged for each sample. Error bars represent the standard deviations between the averages of three independent samples.
Variants of luxA containing the -laa, -aav and -asv tags were chosen for further analysis. Plasmids containing the constructs were digested with AgeI and BfuAI, gel purified, and cloned as described above into the similarly treated pCDABE. The latter contains the entire P. luminescens lux operon under the control of the lac promoter in the multi-copy pCR-XL-TOPO vector (Invitrogen). Resulting plasmids were then transformed into E. coli TOP10 F′ cells (Invitrogen) for IPTG-inducible expression. For induction experiments, overnight cultures of cells grown in LB with 0.2 mM IPTG were diluted 1:5 in the same, fresh, preheated medium, and incubated at 37°C until reaching an OD600 of 0.15–0.2 (approximately 4 h). Cells were subsequently collected by centrifugation, washed with fresh M9 containing 100 mM leucine medium without inducer, and incubated in M9 + leucine medium at 37°C. One milliliter samples were removed every 20 min for bioluminescent measurements using a SDI Deltatox handheld luminometer. Three measurements were taken in rapid succession for each sample and their values were averaged, and three independent replicates were performed for each variant.
Results and discussion
Bioluminescence measurements of cells harboring plasmids containing the wild type luxAB genes of P. luminescens behind a constitutive promoter were compared to those of luxA and luxB variants containing carboxy-terminal tags designed to accelerate degradation of the protein(s) within the cell. Assays using constructs containing only luxA and luxB (i.e. which lack luxC, D, and E and hence are unable to produce the substrate) were conducted by adding antibiotics to block transcription and translation (rifampicin and tetracycline, respectively), followed by monitoring bioluminescence with the addition of aldehyde over time. As shown in Fig. 1, modification of the LuxB protein did not result in a decrease of bioluminescence. Bioluminescence in this variant was found to remain relatively constant similar to the wild type over the course of the experiments. Addition of the same -aav tag (encoding AANDENYAAAV) to the luxA gene resulted in a rapid decrease in bioluminescence after time t 0. An even more dramatic decrease in bioluminescence was observed when both LuxA and LuxB included the -aav tag. Interestingly, the clone containing luxABaav (wild type luxA with modified luxB) appeared to produce more light than did the wild-type luxAB. Since this result was reproducible within the experimental parameters (Fig. 1) yet inconsistent with those generated in the absence of antibiotics (Table 3), it is believed that this result is an experimental artifact.
Total bioluminescence (arbitrary units) of clones containing wild type luxAB, and luxABaav (left axis), and luxAaavB, luxAaavBaav and the negative control (right axis) over time following the addition of tetracycline and rifampicin (t = 0). Measurements were made following the addition of N-decanal as described in the “Materials and methods”. Values are averages of triplicate samples with error bars representing standard deviations among the replicates. The negative control contains the parent plasmid without the luciferase genes
In the unregulated case where enzyme was allowed to accumulate to a steady state level dependent upon the relative half-lives of their luciferases, mid-log cultures of pLuxAaavB produced approximately 100-fold less bioluminescence than the wild-type (Table 3). The double modified holoenzyme produced bioluminescence nearly 800-fold less than the wild type (Table 3).
To determine if the drop in bioluminescence output could be attributed solely to interference of the carboxy-terminal tag to enzymatic activity, two additional constructs were made with predicted half-lives greater (-asv encoding AANDENYAASV) and lesser (-laa encoding AANDENYALAA) than those predicted for -aav. Total bioluminescent output for all of these constructs is shown in Table 3.
Variants of luxA were subsequently cloned into a vector containing the entire lux operon by restriction digestion interior to each of the luxA and B genes and replacement of the wild type fragment with fragments containing the modified luxA gene. Results of experiments on luminescent decay following the removal of inducer (isopropyl-β-d-thiogalactopyranoside; IPTG) are shown in Fig. 2.
Conclusions
The results for variants containing the modified luxA gene are consistent with those previously reported by Anderson et al. (Andersen et al. 1998) for GFP, and indicate that the single modification of this monomer is sufficient to result in decreased functional half-life of the heterodimeric holoenzyme (Fig. 1 and Table 3). This modification may impact enzymatic activity of the holoenzyme since the carboxy-terminus of LuxA is thought to be involved in substrate binding (Valkova et al. 1999). However, bioluminescence outputs for all variants were readily detectable under our test conditions. Interestingly, modification of the LuxB monomer alone neither decreased peak bioluminescence nor resulted in decay of bioluminescence over time when compared to the wild type. Modification of both luxA and luxB genes to include the -aav tag (luxAaavBaav), however, resulted in lower total bioluminescence than modification of luxA alone (Fig. 1). These results suggest that the carboxy-terminus of LuxB may be protected in the heteroduplex of the holoenzyme and inaccessible to cellular proteases, but its release following the degradation of LuxA may facilitate its subsequent degradation and account for the reduced bioluminescence of the double-modified variant.
Modification of protein half-lives in eukaryotic proteins by the carboxy-terminal additions of the PEST-rich region from the G1 cyclin Cln2 and the degradation domain of mouse ornithine decarboxylase have been used to generate destabilized variants of GFP in yeast and mammalian systems (Mateus and Avery 2000; Li et al. 1998), as well as firefly luciferase (Leclerc et al. 2000). Such an approach could also be applied to the luxA and luxB genes, which have been codon-optimized for expression in eukaryotic systems (Gupta et al. 2003; Patterson et al. 2005). Furthermore, the approach taken here in which the insertion of the luxA-tag-B fragment into an existing construct containing the entire P. luminescens lux cassette should be widely applicable to a number of existing lux-based reporter systems, and will allow relatively simple retrofitting of those constructs to generate short half-life variants for dynamic monitoring of changes in gene expression in reporter gene transcription, in vivo imaging, and synthetic biology applications.
References
Andersen J, Sternberg C, Poulsen L, Bjørn SP, Givskov M, Molin S (1998) New unstable variants of green fluorescent protein for studies of transient gene expression in bacteria. Appl Environ Microbiol 64:2240–2246
Chalfie M, Tu Y, Euskirchen G, Ward W, Prasher D (1994) Green fluorescent protein as a marker for gene expression. Science 263:802–805
Cormack B, Valdivia R, Falkow S (1996) FACS-optimized mutants of the green fluorescent protein (GFP). Gene 173:33–38
DiGrazia P, King J, Blackburn J, Applegate B, Bienkowski P, Hilton B, Sayler G (1991) Dynamic response of naphthalene biodegradation in a continuous flow soil slurry reactor. Biodegradation 2:81–91
Doyle TC, Burns SM, Contag CH (2004) In vivo bioluminescence imaging for integrated studies of infection. Cell Microbiol 6:303–317
Elowitz M, Leibler S (2000) A synthetic oscillatory network of transcriptional regulators. Nature 403:335–338
Engebrecht J, Nealson K, Silverman M (1983) Bacterial bioluminescence: isolation and genetic analysis of functions from Vibrio fischeri. Cell 32:773–781
Gardner T, Cantor C, Collins J (2000) Construction of a genetic toggle switch in Escherichia coli. Nature 403:339–342
Gottesman S, Roche E, Zhou Y, Sauer R (1998) The ClpXP and ClpAP proteases degrade proteins with carboxy-terminal peptide tails added by the SsrA-tagging system. Gene Dev 12:1338–1347
Gupta R, Patterson S, Ripp S, Simpson M, Sayler G (2003) Expression of the Photorhabdus luminescens lux genes (luxA, B, C, D, and E) in Saccharomyces cerevisiae. FEMS Yeast Res 4:305–313
Hasty J, McMillen D, Collins J (2002) Engineered gene circuits. Nature 420:224–230
Herman C, Thévenet D, Bouloc P, Walker G, D’Ari R (1998) Degradation of carboxy-terminal-tagged cytoplasmic proteins by the Escherichia coli protease HflB (FtsH). Gene Dev 12:1348–1355
Kadurugamuwa JL, Sin LV, Yu J, Francis KP, Kimura R, Purchio T, Contag PR (2003) Rapid direct method for monitoring antibiotics in a mouse model of bacterial biofilm infection. Antimicrob Agents Chemother 47:3130–3137
Keiler K, Waller P, Sauer R (1996) Role of peptide tagging system in degradation of proteins synthesized from damaged messenger RNA. Science 271:990–993
Kim Y-I, Burton R, Burton B, Sauer R, Baker T (2000) Dynamics of substrate denaturation and translocation by the ClpXP degradation machine. Mol Cell 5:639–648
King J, DiGrazia P, Applegate B, Sanseverino J, Dunbar P, Larimer F, Sayler G (1990) Rapid and sensitive bioluminescent reporter technology for naphthalene exposure and biodegradation. Science 249:778–781
Kuklin NA, Pancari GD, Tobery TW, Cope L, Jackson J, Gill C, Overbye K, Francis KP, Yu J, Montgomery D et al (2003) Real-time monitoring of bacterial infection in vivo: development of bioluminescent staphylococcal foreign-body and deep-thigh-wound mouse infection models. Antimicrob Agents Chemother 47:2740–2748
Leclerc G, Bookfor F, Faught W, Frawley L (2000) Development of a destabilized firefly luciferase enzyme for measurement of gene expression. BioTechniques 29:590–601
Li X, Zhao X, Fang Y, Jiang X, Duong T, Fan C, Huang C-C, Kain SR (1998) Generation of destabilized green fluorescent protein as a transcription reporter. J Biol Chem 273:34970–34975
Mateus C, Avery SV (2000) Destabilized green fluorescent protein for monitoring dynamic changes in yeast gene expression with flow cytometry. Yeast 16:1313–1323
van der Meer JR, Tropel D, Jaspers M (2004) Illuminating the detection chain of bacterial bioreporters. Environ Microbiol 6:1005–1020
Meighen EA (1991) Molecular biology of bacterial bioluminescence. Microbiol Rev 55:123–142
Miyawaki A, Llopis J, Heim R, McCaffery JM, Adams JA, Ikura M, Tsien RY (1997) Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature 388:882–887
Patterson S, Dionisi H, Gupta R, Sayler G (2005) Codon optimization of bacterial luciferase (lux) for expression in mammalian cells. J Ind Microbiol Biotechnol 32:115–123
Sambrook J, Fritsch E, Maniatis T (1989) Molecular cloning – a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Schmidt T, Kopecky K, Nealson K (1989) Bioluminescence of the insect pathogen Xenorhabdus luminescens. Appl Environ Microbiol 55:2607–2612
Szittner R, Meighen E (1990) Nucleotide sequence, expression, and properties of luciferase coded by lux genes from a terrestrial bacterium. J Biol Chem 265:16581–16587
Triccas J, Pinto R, Britton W (2002) Destabilized green fluorescent protein for monitoring transient changes in mycobacterial gene expression. Res Microbiol 153:379–383
Tu G, Reid G, Zhang J, Moritz R, Simpson R (1995) C-terminal extension of truncated recombinant proteins in Escherichia coli with a 10Sa RNA decapeptide. J Biol Chem 270:9322–9326
Valkova N, Szittner R, Meighen E (1999) Control of luminescence decay and flavin binding by the LuxA carboxyl-terminal regions in chimeric bacterial luciferases. Biochemistry 38:13820–13828
Van Dyk T, DeRose E, Gonye G (2001) LuxArray, a high-density, genomewide transcription analysis of Escherichia coli using bioluminescent reporter strains. J Bacteriol 183:5496–5505
Weber-Ban E, Reid B, Miranker A, Horwich A (1999) Global unfolding of a substrate protein by the Hsp100 chaperone ClpA. Nature 401:90–93
Xiong YQ, Willard J, Kadurugamuwa JL, Yu J, Francis KP, Bayer AS (2005) Real-time in vivo bioluminescent imaging for evaluating the efficacy of antibiotics in a rat Staphylococcus aureus endocarditis model. Antimicrob Agents Chemother 49:380–387
Zlokarnik G, Negulescu PA, Knapp TE, Mere L, Burres N, Feng L, Whitney M, Roemer K, Tsien RY (1998) Quantitation of transcription and clonal selection of single living cells with beta-lactamase as reporter. Science 279:84–88
Acknowledgements
We would like to thank R. Gupta and C. Cox for helpful discussions, M. Elowitz for the repressilator reporter plasmid pGFPasv, and J. Sanseverino for pCDABE. This work was supported in part by an LDRD grant from Oak Ridge National Lab and the U.S. Department of Energy and by the University of Tennessee Center for Environmental Biotechnology Research Center of Excellence.
Author information
Authors and Affiliations
Corresponding author
Additional information
Statement of competing interests: MSA and GSS have applied for patents on the materials described here.
Author’s contributions: MSA, JRW, and CSC carried out the molecular biology experiments and bioluminescence assays. MSA, GSS, and MLS conceived of the study, participated in its design and coordination, and helped to draft the manuscript. All authors read and approved the final manuscript.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License ( https://creativecommons.org/licenses/by-nc/2.0 ), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Allen, M.S., Wilgus, J.R., Chewning, C.S. et al. A destabilized bacterial luciferase for dynamic gene expression studies. Syst Synth Biol 1, 3–9 (2007). https://doi.org/10.1007/s11693-006-9001-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11693-006-9001-5