Abstract
Acyl-CoA-binding proteins (ACBPs) are important for the transport of acyl groups for macromolecular biosynthesis involved in plant growth, development, and diverse stress (e.g., cold, drought, salinity, and heavy metals) responses. Here, we report the phylogeny and characteristics of the ACBP family in the woody plant Populus trichocarpa. Eight genes encoding ACBP proteins were identified, and they are distributed on eight chromosomes in P. trichocarpa. These PtACBP genes were divided into four subgroups according to gene structure, conserved motifs and phylogenetic relationship. Promoter analysis revealed that cis-elements were related to stress response, phytohormone response, and physical and reproductive growth regulation. Expression levels of PtACBP genes varied among different organs, with the highest expression in leaves and the lowest in stems. Quantitative real-time PCR (qRT-PCR) analysis showed that under salinity-alkali stresses (i.e., 200 mM NaCl, 75 mM Na2CO3, and 100 mM NaHCO3), four (PtACBP1, PtACBP3, PtACBP4 and PtACBP8) of eight PtACBP genes were significantly induced in roots and leaves. These data provide a comprehensive analysis of the ACBPs family in P. trichocarpa, which could be useful for gene function analyses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lipids are critical for maintaining cellular membrane composition and integrity and for storing energy for use in diverse cellular, developmental, and physiological events. In plants, the subcellular trafficking of lipids and their derivatives are pivotal for cellular signal perception and transduction, photosynthesis, and multiple metabolic processes (Ohlrogge and Browse 1995). Acyl-CoA-binding proteins (ACBPs) belong to an important housekeeping protein family involved in intracellular trafficking of acyl-CoA esters and in the formation and maintenance of a cytosolic acyl-CoA pool (Xiao and Chye 2011a).
On the basis of phylogenetic analysis, ACBPs have been proposed to be present in more than 66 plant species. They were experimentally identified in 27 species, ranging from algae, moss, ferns, and gymnosperms to angiosperms (Lung and Chye 2016; Wang et al. 2019). Plant ACBPs have been extensively investigated in two models, the eudicot Arabidopsis thaliana and the monocot Oryza sativa (hereafter, Arabidopsis and rice; Du et al. 2016; Lung and Chye 2016; Ye and Chye 2016). Based on the molecular mass, domain architecture, and phylogeny, plant ACBPs are differentiated into four classes including small ACBPs (class I), ankyrin-repeat ACBPs (class II), large ACBPs (class III), and kelch-ACBPs (class IV) (Lung and Chye 2016), which play nonredundant roles in different tissues and organs (Meng et al. 2011). The expression patterns of six A. thaliana ACBPs and six rice ACBPs and data from other molecular genetic analyses indicate that they have versatile functions at the cellular and subcellular levels, including transport of lipids (i.e., phospholipids, phosphatidylcholine, and acyl-CoA ester), saturated fatty acid metabolism, fatty acid β-oxidation, vesicular trafficking, and signal transduction during disparate developmental processes (e.g., pollen and seed development, seed gemination, and cuticle formation). They also participate in plant responses to diverse biotic and abiotic stresses such as ABA, indole-3-butyric acid, ethanol, heavy metal, hypoxia, cold, wounding, drought, salt, oxidative stress, and pathogen infection (Gao et al. 2009; Fan et al. 2010; Xiao and Chye 2011a; Liu et al. 2015; Du et al. 2016; Qin et al. 2016; Ye and Chye 2016; Narayanan et al. 2019, 2020; Wang et al. 2019; Zhu et al. 2021).
The diverse and specific functions of different ACBP members depend on their subcellular localization and spatiotemporal expression. AtACBPs and OsACBPs localize in a variety of subcellular components, such as the plasma membrane (PM) (Chye et al. 1999; Gao et al. 2009, 2010a, 2010b; Chen et al. 2010; Napier and Haslam 2010; Licausi et al. 2011; Du et al. 2013b), cytosol (Chen et al. 2008; Yurchenko et al. 2009, 2014; Hsiao et al. 2014; Meng et al. 2014), vesicle (Chye et al. 1999), endoplasmic reticulum (ER) reticulum/ space (Chye et al. 1999; Leung et al. 2006; Xiao et al. 2010; Xiao and Chye 2011b; Xia et al. 2012; Takato et al. 2013; Xue et al. 2014; Xie et al. 2015), endomembrane (Chye et al. 1999; Li and Chye 2003; Licausi et al. 2011; Du et al. 2013b), Golgi apparatus (Li and Chye 2003, 2004; Gao et al. 2009, 2010a; Licausi et al. 2011), chloroplast (Meng et al. 2014), peroxisome (Meng et al. 2014), nuclear periphery (Li et al. 2008), cytoskeleton (Takato et al. 2013), and apoplast (Leung et al. 2006; Xiao et al. 2010; Pastor et al. 2013; Lung and Chye 2016). Moreover, expression of plant ACBP members from Arabidopsis, rice, Cucumis sativus, Cucurbita maxima, Brassica napus, and Cocos nucifera has been detected in embryos, stem epidermis, guard cells, pollen, and phloem sap. Their expression has been linked to their specific functions in early embryogenesis (Chye et al. 1999; Chen et al. 2010; Du et al. 2013a, b; Guo et al. 2019), cuticle formation (Xue et al. 2014), drought response (Du et al. 2013a), pollen development (Hsiao et al. 2015), and systemic lipid transport (Walz et al. 2004; Suzui et al. 2006; Guelette et al. 2012). Results from model plant species have facilitated the understanding of the diverse representative functions of ACBPs.
However, key scientific issues underlying tree-specific traits, such as wood formation, long-term perennial growth, and seasonality, cannot be easily addressed using Arabidopsis or rice (Jansson and Douglas 2007). In many respects, trees are physiologically and genetically distinct from Arabidopsis due to their multiple origins throughout land plant evolution (Jansson and Douglas 2007).
Poplar (Populus spp.) is an important economic and ecological species (Bradshaw et al. 2000), a model system for tree and woody perennial plant biology, and relatively close phylogenetically to Arabidopsis in the angiosperm clade eurosid. The completion of a draft sequence of the black cottonwood (Populus trichocarpa) genome facilitated the rapid collection of genomic and molecular genetics data for Populus and the comparative analyses of Arabidopsis and Populus (Tuskan et al. 2006; Jansson and Douglas 2007). Comparative genomics and functional studies on Populus have provided important information on the mechanism of wood development, vascular cambium formation, seasonality, phenology, flowering, various stress responses, and interactions with other organisms (Jansson and Douglas 2007; Xu et al. 2021). However, the genome-wide ACBP gene family has not been comprehensively evaluated in P. trichocarpa.
In the present genome-wide analysis of ACBP genes in P. trichocarpa, we identified eight PtACBP genes and analyzed the phylogeny, gene structure, conserved motifs, chromosome locations, and expression patterns in different tissues and organs and in response to multiple abiotic stresses. These results will provide a theoretical basis for further investigation into the potential roles of PtACBP genes in growth, development, and stress response.
Materials and methods
Identification of ACBP family in P. trichocarpa
To identify ACBPs in P. trichocarpa, the protein database of Populus was downloaded from the Phytozome database (https://phytozome.jgi.doe.gov/pz/portal.html) (Goodstein et al. 2012). The updated Hidden Markov Model (HMM) profile file (ACB.hmm) for the acyl-CoA-binding domain (ACB) (PF00887) was downloaded from the Pfam database (http://pfam.xfam.org/), and then used in a searched of the poplar genome database. The amino acid sequences for six Arabidopsis acyl-CoA-binding proteins, AtACBP1 (NP_200159.1), AtACBP2 (OAP00506.1), AtACBP3 (Q9STX1.1), AtACBP4 (OAP02573.1), AtACBP5 (AED93708.1), and AtACBP6 (AEE31396.1) were used as the query sequences in a BLASTP search against the online genome database of P. trichocarpa. The ACB domain at the N-terminus, the ankyrin (ANK) domain, and the Kelch domain at the C-terminus were used as search criteria (Meng et al. 2011). The characteristics of amino acid sequences, such as molecular weight, isoelectric point (pI), number of amino acids, aliphatic index, chromosomal location, and grand average of hydropathicity GRAVY score, were determined using the online ExPASy program (http://www.expasy.org/). Cellular localization was determined using the online package Cell-PLoc 2.0 (http://www.csbio.sjtu.edu.cn/bioinf/Cell-PLoc-2/), WoLF PSORT (http://www.genscript.com/wolf-psort.html) and MultiLoc2 (http://www-bs.informatik.uni-tuebingen.de/Services/MultiLoc2).
Chromosomal locations of ACBP genes
The chromosomal location of the PtACBP genes was obtained from the Phytozome database and mapped using MapGene2Chrom web v2 (http://mg2c.iask.in/ mg2c_ v2.0/).
Structural analysis of ACBP proteins
The conserved domains of ACBPs in Arabidopsis, P. trichocarpa, and Picea sitchensis were referenced in the NCBI database (https://www.ncbi.nlm.nih.gov/protein/). IBS software (Liu et al. 2005) was used to map the protein primary structure.
Exon/intron structure and conserved motifs analysis
The Gene Structure Display Server (GSDS2.0, http://gsds.cbi.pku.edu.cn) was used to predict the distribution patterns of exons and introns in the PtACBP genes. Conserved motifs were identified in PtACBP genes by using the online MEME tool (http://meme-suite.org/tools/meme) with the options maximum distinct motifs to identify = 10, minimum width = 7, and maximum width = 50.
Analysis of cis-acting elements
Promoter sequences of the PtACBP family genes were downloaded from the Phytozome database (https://phytozome.jgi.doe.gov/pz/portal.html). The PlantCARE online database (http://bioinformatics.psb.ugent.be/webtools/plantcare/html) was used to predict regulatory elements. The GSDS2.0 website was used to visualize cis-acting elements.
Phylogenetic analysis and sequence alignments of plant ACBPs
In the NCBI protein database, the plant was searched by keyword " ACBP " to obtain ACBP protein in plants (https://www.ncbi.nlm.nih.gov/protein/), the ACBP protein sequences in woody plants were manually screened, and then these sequences had been multi-sequence alignment, After removing the same sequence in the same species (identify (A,B) > 95% & identify (B, A) > 95%), and after removed the protein sequence of the amino acid sequence length < 60, 125 sequences from 23 species were finally obtained, including eight ACBP proteins of P. trichocarpa. The sequences were phylogenetically analyzed using ClustalW (PMID: 17,846,036) with default parameters (gap opening penalty = 10, gap extension penalty = 0.2) and a tree constructed in MEGA 7 (https://www.megasoftware.net) (Larkin et al. 2007) using the maximum likelihood method (Kumar et al. 2016) and 1000 bootstrap replications. In total, 11 lant protein sequences consisting of an ACB domain were obtained using a detailed BLAST search of nonredundant protein sequences from the NCBI (http://blast.ncbi.nlm.nih.gov/Blast.cgi). Multiple sequence alignments of PtACBP and amino acid sequences from other plant species were performed using Bioedit software (Alzohairy 2011).
Plant materials, RNA extraction, cDNA synthesis, and RT-qPCR
P. trichocarpa sterile plantlets were grown in the State Key Laboratory of Tree Genetics and Breeding of Northeast.
Forestry University, Harbin, China. Sterile tissue of P. trichocarpa seedlings was cultured for 3 weeks with 16 h light/8 h dark at 25 °C. Samples from the roots, stems, and leaves were collected, immediately frozen in liquid nitrogen, and then stored at − 80 °C for further analysis. For abiotic stress assays, 3-week-old seedlings were transferred to 1/2 MS medium containing 200 mM NaCl, 100 mM NaHCO3, or 75 mM Na2CO3. The roots and leaves were harvested at 6, 12, 24, and 48 h after the start of the stress treatments, and then stored at − 80 °C for PCR analysis.
Total RNA was extracted using TRIzol reagent (Takara Bio, RNAiso Plus 9109, JPN) following the manufacturer’s protocol. RNA quality was detected using a UV spectrophotometer (BioMate 3S, Thermo Fisher Bio, USA). The PrimeScript RT Reagent Kit (Takara Bio, Perfect Real Time, RR047A, JPN) was used for cDNA synthesis. Three biological replicates of the cDNA samples were analyzed by qRT-PCR with primers for PtACBP genes designed using Primer3Plus (http://primer3plus.com/cgi-bin/dev/primer3plus.cgi) (Supplemental Table S1). The qRT-PCRs were run in an Applied Biosystems 7500 Real-Time PCR System (Applied Biosystems, Waltham, MA, USA) using SYBR Premix Ex Taq II (Takara Bio, RR820A, JPN). The total volume of the reaction is 20 μl, and the reaction conditions are as follows: 95 °C, 10 min, then 95 °C, 15 s, 57 °C, 20 s, 72 °C, 35 s for 40 cycles. Three biological replicates of each treatment were conducted.
Expression of PtACBP genes in tissues and organs of P. trichocarpa
Tissue- and organ-specific expression data of PtACBP genes in mature leaves, young leaves, roots, nodes, and internodes were derived from PopGenIE (http://popgenie.org) and used to generate visualizations (Sundell et al. 2015).
Results
Identification of ACBP proteins and chromosome localization of ACBP genes in P. trichocarpa
The HMMER searches using the ACB domain profile PF00887 as a query against the P. trichocarpa protein database (http://pfam.xfam.org/) generated 22 candidate proteins, and BLASP search using the six Arabidopsis ACBP proteins as queries resulted in 30 candidates. Then, the 22 overlapping candidates were screened for Ankyrin and Kelch domains, which generated eight ACBP proteins in P. trichocarpa. Eight PtACBPs were designated as PtACBP1 to PtACBP8, with locus numbers in the P. trichocarpa gene database of Phytozome v3.0 of Potri.003G103700.1, Potri.001G130200.1, Potri.012G017700.1, Potri.015G010200.1, Potri.002G120200.1, Potri.014G018700.1, Potri.005G026900.1, and Potri.013G018800.1, respectively (Table 1). The protein molecular weight, pI, number of amino acids, aliphatic index, chromosomal location, and grand average of hydropathicity GRAVY score are given in Table 1.
Interestingly, on the basis of Populus genome database, eight PtACBP genes were physically mapped on chromosome 1, 2, 3, 5, 12, 13, 14, and 15, respectively, but not on the other seven chromosomes of P. trichocarpa (Fig. 1). This distribution pattern implied a distinct function for PtACBP genes, which needs to be further investigated.
The PtACBP characterization and subcellular localization prediction
The eight members of PtACBPs differed considerably in the number of amino acids, ranging from 116 to 686 amino acids (aa; Table 1). Among the eight, PtACBP7 and PtACBP8 were relatively larger (677 and 686 aa, respectively), and PtACBP1 (116 aa) and PtACBP2 (117 aa) were the smallest (Table 1). Their molecular weights varied from 13.24 kDa for PtACBP1 to 75.13 kDa for PtACBP8, and their theoretical pIs ranged from 4.31 (PtACBP6) to 9.10 (PtACBP2).
The subcellular localization analysis of the PtACBPs predicted PtACBP1, PtACBP2, PtACBP3, PtACBP4, PtACBP6, PtACBP7, and PtACBP8 to be in the nucleus; PtACBP1, PtACBP3, PtACBP5, PtACBP7 and PtACBP8 in the cytoplasm; PtACBP5 and PtACBP6 in the chloroplasts; and PtACBP3, PtACBP5 and PtACBP6 in the mitochondria (Supplemental Table S2).
Architectural comparisons of ACBP domains from P. trichocarpa, Picea sitchensis, and A. thaliana
Based on domain compositions and amino acid sequence lengths, the eight PtACBPs were divided into four subfamilies (Fig. 2). All the PtACBPs had an ACB domain, and some had either an Ankyrin domain or a Kelch domain. The 10-kDa PtACBP1 (116 amino acids) and PtACBP2 (117 amino acids) had only an ACB domain and were categorized as Small-ACBPs in Class I. PtACBP3 (246 amino acids) and PtACBP4 (353 amino acids) had an ankyrin-repeat domain and were classified as ankyrin-repeat ACBPs (ANK-ACBP) in Class II. Two of the proteins with molecular weights greater than 40 kDa, PtACBP5 (364 amino acids) and PtACBP6 (427 amino acids), were categorized as Large-ACBP in Class III, and had an ACB domain in the middle. In addition, PtACBP7 (677 amino acids) and PtACBP8 (686 amino acids) had N-terminal ACB domains and at least one kelch domain and were classified as Kelch-ACBPs in Class IV (Fig. 2).
Schematic domain structures of ACBPs from Arabidopsis thaliana, Populus trichocarpa, and Picea sitchensis. Green indicates the acyl-CoA-binding domain (ACB), yellow indicates the ankyrin repeats domain (ANK), and pink indicates the kelch domain (Kelch). The “ + ” indicates that the actual number of ACBPs may be greater because sequencing of the genome is still incomplete
When comparing the PtACBPs with those from A. thaliana and P. sitchensis, we found that the amino acid sequences of PtACBPs are more similar to those of A. thaliana, and the amino acid identities in each class between P. trichocarpa and A. thaliana were as follows: classes I (73%–74%), II (66%–73%), III (40%) and IV (72%–76%). Interestingly, in P. sitchensis, PsACBP1 has only the ACB domain and is in Class 0, and there were no PsACBPs in Class III. However, A. thaliana or P. trichocarpa had no ACBPs in Class 0 (Fig. 2). ACBPs in Class 0 have an ACB domain similar to those in Class III in higher plants, but they have fewer amino acids than in Class III members. Moreover, both A. thaliana and P. trichocarpa have two ACBPs in Class II and Class IV, respectively, but P. sitchensis only has one in each (Fig. 2). In addition, A. thaliana ACBP families have only one member in class I (AtACBP6) and class III (AtACBP3), but the PtACBP family has two members in class I (PtACBP1, PtACBP2) and class III (PtACBP5 and PtACBP6. These results imply that ACBP families arose during the evolution of lower plants to higher plants (Meng et al. 2011).
Analysis of gene structure and distribution of conserved motifs
In the analysis of the structure of the eight PtACBP genes using GSDS2.0 and comparison of the mRNA and genomic DNA nucleotide sequences, exon length was found to be mostly conserved, and the number of exons was similar among each subfamily except Class I (Fig. 3). In Class I, two exons were found in PtACBP1 (122 and 220 bp) and four exons in PtACBP2 (95, 117, 62, and 79 bp). In Class II, PtACBP3 and PtACBP4 contained six exons, five of which (the 2nd to the 6th) were similar in length (105, 100–103, 91, 97, 170 bp, respectively). Similarly, in Class IV, both PtACBP7 and PtACBP8 have 18 exons. In Class III, PtACBP5 and PtACBP6 have five and six exons, respectively, and four exons were mostly conserved (778–817, 105, 100, 74 bp, respectivel). Importantly, the length and position of introns varied considerably (Fig. 3). These results indicate that there is a close relationship between the phylogeny and gene structure of PtACBP genes.
Ten conserved motifs of PtACBP family members were identified using online MEME software and named motif 1 to 10 (Fig. 4A, B). Subfamily members had similar motif types and arrangements (Fig. 4A). Class I contained only one motif; this single-motif model was exclusive to this PtACBP subfamily. Class II had three motifs, Class III had two, and Class IV had 10. Importantly, motifs 4, 5, 7, 8, 9, and 10 only appeared in Class IV; thus, these motifs can serve to identify PtACBP members in Class IV (Fig. 4A).
Predicted conserved motifs and amino acid sequences of the eight ACBPs found in Populus trichocarpa. A Classes of eight PtACBPs based on conversed motifs and corresponding p-values. Motifs are in different colors, which are defined in B. B Amino acid sequence for each motif. Motifs were predicted by MEME motif searching, and 10 were selected as the maximum number parameter
Analysis of PtACBP promoters for cis-acting element
The 2000-bp sequence upstream of the initial transcription position was analyzed to identify various cis-acting elements using the Plant CARE website. The distribution of 30 cis-acting elements in eight PtACBPs and their distinct functions are shown in Fig. 5.
Among the light-signaling elements found in the PtACBP genes were G-box, Box 4, Gap-box, GT1-motif, AE-box, TCT-motif, TCCC-motif, MRE, AT1-motif, I-box, GATA-motif, AAAC-motif, Box II, L-box, AE-box, Sp1, chs-CMA1a, and ACE (Table S3). Seven cis-elements were classified as involved in hormone responses: ABRE, P-box, GARE-motif, TGA-element, CGTCA-motif, TGACG-motif, and TCA-element (Table S3). For instance, all PtACBP genes except for PtACBP7 contained the ABA-responsive element ABRE (Fig. 5). Four PtACBP genes (i.e., PtACBP2, PtACBP3, PtACBP6, and PtACBP7) contained gibberellin-responsive P-boxes and GARE-motifs, and four genes (i.e., PtACBP2, PtACBP4, PtACBP5, and PtACBP8) contained auxin-responsive element TGAs (Fig. 5). In addition, five genes (i.e., PtACBP2, PtACBP3, PtACBP5, PtACBP7, and PtACBP8) contained MeJA responsive elements CGTCA-motifs and TGACG-motifs, and six genes (i.e., PtACBP2, PtACBP3, PtACBP5, PtACBP6, PtACBP7, and PtACBP8) contained salicylic-acid-responsive elements TCAs (Fig. 5). Also found in different PtACBP members were cis-elements such as MBS, ARE, LTR, GC-motif, and TC-rich repeats that are related to various stress responses. For example, PtACBP2, PtACBP3, PtACBP5, and PtACBP6 had drought-inducible elements (MBS), seven genes (i.e., PtACBP1 to PtACBP4, PtACBP6 − 8) had the anaerobic induction element ARE, and four genes (PtACBP1, PtACBP2, PtACBP4, PtACBP8) had a low temperature responsive (LTR) element (Fig. 5). PtACBP5 contained anoxia-specific inducibility element GC-motifs (Fig. 5). These results imply that PtACBPs are critical in P. trichocarpa for diverse stress responses.
Phylogenetic analyses of ACBPs from 23 plant species
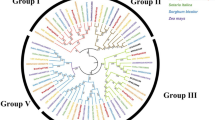
In the phylogenetic tree based on 125 ACBP sequences from 23 plant species using the maximum likelihood (ML) method, ll ACBPs clustered into five groups: Class 0, Class I, Class II, Class III, and Class IV, which contained 2, 36, 26, 32, and 29 members, respectively (Fig. 6). High-bootstrap support was obtained for classes 0, I, III, and IV, and ACBP Class I was basal to the other classes (Fig. 6). Class II was the least-supported clade with a bootstrap value of 47% (Fig. 6). PtACBP1 (XP_002303469.1) and PtACBP2 (XP_006368280.2) grouped into Class I, PtACBP3 (PNT08967.1) and PtACBP4 (XP_002321391.1) into Class II, and PtACBP5 (XP_024449804.1) and PtACBP6 (XP_006374836.1) grouped into Class III (Fig. 6). Additionally, PtACBP7 (XP_024458146.1) and PtACBP8 (XP_006375749.1) were grouped into Class IV (Fig. 6). Notably, seven of eight PtACBPs (i.e., PtACBP1, PtACBP3, PtACBP4, PtACBP5, PtACBP6, PtACBP7 and PtACBP8) except PtACBP2 were clustered closely with PeACBPs from Populus euphratica, indicating a close evolutionary relationship between PtACBPs and PeACBPs.
Sequence alignment of the ACB domain of ACBPs from different plant species
Amino acid sequence variations could influence binding affinities of ACBPs to acyl-CoA esters (Emanuelsson et al. 2000; Xie et al. 2015). Two conserved motifs of YKQA and KWDAW in ACBPs were previously considered as essential motifs for binding acyl-CoA esters (Kragelund et al. 1993; Xiao and Chye 2011a). In this study, multiple sequence alignment analysis indicated that amino acid sequences of eight PtACBPs had similarities with those from other plant species (Fig. 7). We found two conserved motifs in PtACBPs when compared with other plant species (Fig. 7). Importantly, the alignment analysis also revealed four conserved alpha-helices (H1-H4) containing five highly conserved potential binding sites for acyl-CoA esters in PtACBPs (Fig. 7). In addition, whether H4, a highly conserved site of alanine (Fig. 7), is critical for binding acyl-CoA esters still needs to be investigated.
Sequence alignments of ACB domains from the PtACBP family and other species. Blue circles show residues identical in all ACBPs. Arrowheads indicate potential binding sites for acyl-CoA esters; H1–H4 indicate the positions of four putative alpha-helices. motifs YKQA and KWDAW are underlined in red. Pt, Populus trichocarpa; At, Arabidopsis thaliana; Cr, Chlamydomonas reinhardtii; Ol, Ostreococcus lucimarinus; Pp, Physcomitrella patens; Sm, Selaginella moellendorffii; Ps, Picea stichensis; Os, Oryza sativa; Hb, Hevea brasiliensis; Pa, Populus alba; Pe, Populus euphratica
Tissue- and organ-specific expression of PtACBPs
The qRT-PCR analyses of the PtACBP genes in roots, stems, and young leaves from P. trichocarpa seedlings in sterile tissue culture showed that expression of all the PtACBP genes was highest in the leaves, lowest in the stems, and intermediate in the roots (Fig. 8A). Interestingly, expression of PtACBP5 and PtACBP2 was relatively higher than the other PtACBP genes in the leaves (Fig. 8A).
Organ-specific expression profiles of PtACBP genes in Populus trichocarpa. A Expression in roots, stems, and leaves of 49-day-old seedlings. Data showed the mean values ± SE (p < 0.05). B Diagrams to show expression levels in mature leaves, young leaves, roots, nodes, and internodes. Profiles for organ-specific expression data were derived from http://PlantGenIE.org
In addition, the tissue-specific expression patterns of PtACBP genes in mature leaves, young leaves, roots, nodes, and internodes were also predicted and visualized based on the data for PtACBP genes from the Plant Genome Integrative Explorer (http://PlantGenIE.org) (Sundell et al. 2015) (Fig. 8B). The expression data showed that the expression of PtACBP1 and PtACBP2 was highest expression in mature leaves and medium or low in roots, similar to our qRT-PCR results (Fig. 8A). Expression of PtACBP4 and PtACBP6 was high in the nodes and internodes (Fig. 8B). The expression of PtACBP5, PtACBP7, and PtACBP8 was higher in the nodes, internodes, and roots than in other organs (Fig. 8B).
Effect of salt-alkali stress on growth of P. trichocarpa
Three-week-old seedlings of P. trichocarpa were treated with 75 mM Na2CO3, 100 mM NaHCO3, or 200 mM NaCl for 0, 6, 12, 24 and 48 h (Fig. 9). Alkaline salt stresses (75 mM Na2CO3 and 100 mM NaHCO3) affected the growth of seedlings. Under 75 mM Na2CO3 for 6 h, the leaves appeared slightly curled, and old leaves turned dark brown. Under 100 mM NaHCO3 for 12 h, the old leaves were curled and/or chlorotic. After 24 h and 48 h treatments with Na2CO3 or NaHCO3, the old leaves start to turn yellow and wilt. However, seedlings treated with NaCl had no obvious differences in phenotype from the control (Fig. 9).
Expression of PtACBP genes in response to salt-alkali stresses
The qRT-PCR analysis showed that PtACBP genes differed in response to various stress treatments, including 200 mM NaCl, 75 mM Na2CO3, and 100 mM NaHCO3 treatments for 6, 12, 24, and 48 h (Fig. 10; Supplemental Fig. S1).
Salinity-alkali responsive expression profiles of PtACBP genes in roots and leaves. Color scale shows 2 − △△Ct values, which were normalized to untreated controls and log2-transformed counts. Blue and red colors indicate downregulated and upregulated, respectively, in response to 75 mM Na2CO3, 100 mM NaHCO3, or 200 mM NaCl
In roots, under 200 mM NaCl treatments, transcript levels of seven PtACBP genes (PtACBP1, PtACBP2, PtACBP3, PtACBP4, PtACBP6, PtACBP7, and PtACBP8) gradually increased from 0 to 48 h and were significantly induced by short-time stress (e.g., 1.5-fold increase after 6 h of NaCl treatments). However, PtACBP5 was downregulated threefold at 24 h (Fig. 10). In leaves under NaCl stress, the expression of PtACBP1, PtACBP3, and PtACBP4 increased similar to their expression in roots, and their differences from the control were the greatest after 24 h of NaCl stress: 4.1, 6.5, and 7.8 times higher, respectively, than the control (Fig. 10). However, the expression levels of PtACBP5 and PtACBP6 in leaves were significantly different from those in roots. PtACBP5 in leaves was downregulated more than twofold at 6 h and fourfold at 12 h, and PtACBP6 was downregulated at all sampling times (Fig. 10).
Under 75 mM Na2CO3 treatments, six genes (PtACBP1, PtACBP3, PtACBP4, PtACBP5, PtACBP7, and PtACBP8) were upregulated, then downregulated (Fig. 10). PtACBP1, PtACBP4, and PtACBP5 were upregulated after the short-time stress (6 h), with highest expression at 12 h (2.4, 6 and 2.9 times higher, respectively, than the control), then downregulated (Fig. 10). Expression of PtACBP3 increased eightfold in the roots at 24 h, whereas PtACBP2 and PtACBP6 were downregulated at 6 h, then up-regulated by 12 h (Fig. 10).
In addition, during the 100 mM NaHCO3 treatments, four genes (PtACBP1, PtACBP3, PtACBP4, and PtACBP8) were upregulated and four (PtACBP2, PtACBP5, PtACBP6, and PtACBP7) were downregulated by 6 h (Fig. 10). The expression pattern of PtACBP3 during the 100 mM NaHCO3 was similar to that under Na2CO3 treatments, which was eightfold higher than in the control. PtACBP5 and PtACBP6 in roots were repressed at all sampling times, and PtACBP5 was downregulated tenfold at 12 h.
Under 75 mM Na2CO3 treatments, the expression levels of PtACBP1, PtACBP3, PtACBP4, PtACBP5, PtACBP7, and PtACBP8 in leaves were similar to those in roots, showing a trend of upregulation, then downregulation (Fig. 10). Significantly, PtACBP3 was upregulated eightfold in leaves compared with the control after 48 h.
Under 100 mM NaHCO3 treatments, five genes (PtACBP2, PtACBP3, PtACBP4, PtACBP7, and PtACBP8) were upregulated in expression, and three (PtACBP1, PtACBP5, and PtACBP6) were downregulated at 6 h (Fig. 10). The expression of PtACBP1 and PtACBP2 in roots was opposite of that in leaves at 6 h. PtACBP5 was downregulated in roots and leaves, whereas PtACBP6 was downregulated in roots and upregulated in leaves at 24 h (Fig. 10). All these results imply that PtACBP genes act in different organs in response to various salinity-alkali conditions.
Discussion
Phylogenetic analysis reveals conservation of the PtACBP family
The search to find ACBP genes in the P. trichocarpa genome identified eight; thus, the PtACBP gene family is small, similar to the case in other plant, including six ACBP genes identified in Hevea brasiliensis (Nie et al. 2018), nine in Zea mays (Zhu et al. 2021) and five in Sorghum bicolor (Paterson et al. 2009). In the phylogenetic analysis, the PtACBP genes grouped into four subfamilies, consistent with the ACBP genes in A. thaliana and rice (Meng et al. 2011). We also compared the subfamily members from A. thaliana, rice and 21 other woody plant species (Fig. 6). As we noted earlier, Populus is relatively closely related to Arabidopsis (Janson and Douglas 2007). We found that all the PtACBP genes except PtACBP2 were evolutionarily close to the PeACBP genes form P. euphratica. This result implies that the ACBP genes in Populus are relatively conserved and have similar functions.
Cis-elements in the promoter and PtACBP expression patterns suggest crucial roles in stress responses
Cis-acting elements are essential in many biological processes and stress responses (Ibraheem et al. 2010). For example, the cis-elements G-Box, I-Box, ARE, ABRE, and CGTCA-motif respond to light, hormonal, and anaerobic induction (Guidotti et al. 1983; Busk and Pages 1998; Mattick and Gagen 2001; Hattori et al. 2002; Turner et al. 2002; Gómez-Porras et al. 2007; Staswick 2008; Yang et al. 2017; Yu et al. 2020). In the present study, several common elements that are involved in the responses to light, ABA, salicylic acid and MeJA were identified in most PtACBP promoter regions. However, only class III PtACBP5 contained an anoxia-specific inducibility element (GC-motif), implying that PtACBP5 is involved in plant abiotic stress defense and response to hypoxia, similar to class III AtACBP3 (Choi et al. 1994; Xie et al. 2015). In addition, stress-related cis-elements, including TC-rich repeats, evident in the 5’-flanking regions of PtACBP8 further indicate that the large PtACBPs might function in stress tolerance, similar to findings for rice (OsACBP4, OsACBP5 and OsACBP6) (Meng et al. 2011).
ACBPs participate in plant responses to diverse biotic and abiotic stresses, such as heavy metal, oxidative stresses, drought, hypoxia, cold, salt and pathogen infection (Gao et al. 2009; Fan et al. 2010; Liu et al. 2015; Du et al. 2016; Qin et al. 2016; Narayanan et al. 2019, 2020; Zhu et al. 2021). In A. thaliana, overexpression of AtACBP6 and AtACBP2 enhances tolerance to cold and drought stress, respectively (Du et al. 2013a, b; Liao et al. 2014). AtACBP3, AtACBP4 and AtACBP6 contribute to drought tolerance by playing a role in cuticle formation (Xia et al. 2012). Similarly, overexpression of OsACBP4 and OsACBP5 increased rice tolerance to drought and salt stresses (Meng et al. 2011). Virus-induced gene silencing reduced expression of GhACBP3 and GhACBP6 subclass genes in cotton, significantly decreasing tolerance to drought and salt stresses (Qin et al. 2016). Membrane-associated AtACBP1 and AtACBP2 bind the heavy metal Pb(II) and are also involved in response to oxidative stress (H2O2) (Gao et al. 2009, 2010b; Du et al. 2016). Interestingly, AtACBP2 interacts with a group VII ethylene response factor (ERF), RAP2.3, to regulate the hypoxia response (Li and Chye 2004). AtACBP3 also participates in the response to hypoxia by interacting with VLC acyl-CoA esters and modulating fatty acid/lipid metabolism (Xie et al. 2015). In addition, AtACBP3 and OsACBP5 participate in defense against pathogen infections. Overexpression of AtACBP3 and OsACBP5 enhance the resistance of A. thaliana and rice to the bacterial pathogen Pseudomonas syringae pv tomAto DC3000 (Narayanan et al. 2019, 2020) and to necrotrophic (Rhizoctonia solani and Cercospora oryzae) and hemibiotrophic (Magnaporthe oryzae and Fusarium graminearum) fungi and biotrophic (Xanthomonoas oryzae pv. oryzae) (Narayanan et al. 2020).
In the present study, we found that the expression patterns of eight PtACBP genes were altered in roots and leaves in response to salinity-alkali stress (Fig. 10). Two PtACBP genes (i.e., PtACBP1 and PtACBP2) were significantly induced by 75 mM NaHCO3 and three (i.e., PtACBP3, PtACBP4, and PtACBP8) by 100 mM NaHCO3 in leaves and roots. Interestingly, two class III genes (PtACBP5 and PtACBP6) were significantly induced in roots but not in leaves. However, class I PtACBP1 and PtACBP2 and class II PtACBP3 and PtACBP4 were upregulated in leaves and roots. Thus, members of the different subfamilies apparently have different roles in response to salinity and alkali stresses.
Subcellular localization of PtACBPs determines function
The subcellular localization of different ACBP members determines their function such as intracellular trafficking, protection, and pool formation of acyl-CoA esters (Lung and Chye 2016). In the eudicot A. thaliana and the monocot rice, the subcellular localization of six AtACBPs and six OsACBPs have been determined using various in silico analyses and experimental verifications, such as observations with immunoelectron microscopy and/or confocal laser-scanning microscopy after transient expression in onion/ tobacco epidermal cells and/or stable expression in transgenic plants, western blots using subcellular fractions, and subcellular proteomic analyses (Ito et al. 2011; Ye et al. 2016). Plasma-membrane-localized class II AtACBP1 and AtACBP2 bind various ligands such as heavy metals, acyl-CoA esters and phospholipids to sense early signals and convey them for downstream responses, which are critical for embryogenesis, seed germination, and diverse ABA and stress responses (Xiao et al 2008; Gao et al. 2009, 2010a; Chen et al. 2010; Du et al. 2013a). Moreover, AtACBP1 and AtACBP2 are localized to the ER or vesicles, as are class II OsACBP4 and class III OsACBP5, facilitating acyl exchange between acyl-CoA and phospholipid pools (Napier and Haslam 2010) and contributing to vesicular lipid trafficking (Chye et al. 1999), which were important for lysoPC turnover and repair of membrane structure. In addition, the apoplast-localized class III AtACBP3 binds to pathogen-secreted lipids after pathogen infection, then is rapidly degraded by apoplastic proteolysis and re-localized intracellularly to trigger the expression of defense genes (Xiao and Chye 2011b). Class IV AtACBP4 is in the cytosol and nuclear periphery and interacts with transcription factor AtEBP in response to ethylene and jasmonate (Li et al. 2008). Cytosolic class IV AtACBP4 cooperates with AtACBP5 and class I AtACBP6 during seed germination (Hsiao et al. 2014). In addition, the peroxisome-localized OsACBP6 is involved in conversion of fatty acids to carbohydrates (Meng et al. 2014). Loss of OsACBP6 not only results in the disruption of acyl-CoA homeostasis but also of peroxidase-dependent reactive oxygen species (ROS) homeostasis (Meng et al. 2020).
The subcellular location of ACBPs in woody plants is still unclear. In our silico analyses, four PtACBPs (i.e., PtACBP1, PtACBP3, PtACBP7 and PtACBP8) were localized in the cytosol and thus should be involved in intracellular acyl-CoA transport (Knudsen et al. 1999; Chen et al. 2008; Meng et al. 2014). Three ACBPs (i.e., PtACBP4, PtACBP5, and PtACBP6) were predicted to be associated with the ER and thus function in the transfer of oleoyl-CoA esters to ER for the biosynthesis of membrane phospholipids and triacylglycerol (Ohlrogge and Jaworski 1997; Xiao and Chye 2009). Besides, the predicted nuclear localization of seven PtACBPs except PtACBP6 implied that they were probably involved in regulating transcription (Li et al. 2008). In addition, the location of PtACBP3, PtACBP5 and PtACBP6 in the mitochondria and PtACBP5 in the chloroplast suggests that they are associated with membrane stability and the trafficking of lipids between these organelles and the cytosol (Li and Chye 2003; Meng et al. 2014; Block and Jouhet 2015).
Spatial expression patterns of ACBPs indicate function specialized for tissues/organs
ACBPs were expressed in a variety of A. thaliana tissues and organs, such as seeds, leaves, and roots, which function in various developmental processes (Gao et al. 2009; Du et al. 2016). During embryogenesis, AtACBP1 facilitates acyl-CoA metabolism and membrane phospholipid biogenesis and shares an overlapping role with AtACBP2 in lipid metabolism (Chen et al. 2010). Cytosolic AtACBP4, AtACBP5 and AtACBP6 facilitate acyl transfer from plastids to ER for glycerolipid synthesis during seed development (Xiao and Chye 2009; Hsiao et al. 2014). During seed germination and seedling development, AtACBP1 promots PLDα1 activity via protein–protein interaction at the plasma membrane, leading to phosphatidic acid accumulation, then triggering of ABA-mediated responses (Du et al. 2013a, b). Also, extracellular targeting of AtACBP3 mediates the modulation of VLC acyl-CoA esters, uniquely affecting membrane lipid metabolism during leaf senescence (Xiao et al. 2010; Xiao and Chye 2010). AtACBP1, AtACBP3, AtACBP4 and AtACBP6 function in cuticle development by maintaining appropriate FA/lipid levels (Xue et al. 2014). In addition, AtACBP5 and AtACBP6 have overlapping roles in pollen development (e.g., microspores, tapetal cells, and pollen grains) by transporting acyl-lipid precursors for sporopollenin biosynthesis (Hsiao et al. 2015).
In this study, we found that all eight PtACBP genes had relatively high expression in leaves, moderate expression in roots, and low expression in stems, similar to their homologs in rice (Meng et al. 2011), suggesting that the genes have similar organ specificities among different plant species.
Conclusions
ACBPs are abundant and widely distributed in various plants and are involved in important processes such as lipid metabolism, enzyme and gene expression regulation, and coping with biotic and abiotic stresses. Here, we provided genome-wide information about the ACBP family of P. trichocarpa. We identified eight P. trichocarpa ACBPs, classified into four subfamilies, which displayed considerable evolutionary divergence in P. trichocarpa and conserved domains. The organ-specific expression profiles of ACBPs indicated potentially divergent functions in P. trichocarpa growth and development. The analyses of cis-acting elements and stress-responsive gene expression profiles suggest that ACBPs are involved in responses to diverse abiotic stresses and hormonal stimuli. Our results establish a foundation for further functional characterization of P. trichocarpa ACBPs.
References
Alzohairy AM (2011) Bioedit: an important software for molecular biology. GERF Bull Biosci 1:60–61
Block MA, Jouhet J (2015) Lipid trafficking at endoplasmic reticulum chloroplast membrane contact sites. Curr Opin Cell Biol 35:21–29
Bradshaw H, Ceulemans R, Davis J, Stettler R (2000) Emerging model systems in plant biology: poplar (Populus) as a model forest tree. J Plant Growth Regul 19(3):306–313
Busk PK, Page SM (1998) Regulation of abscisic acid-induced transcription. Plant Mol Biol 37(3):425–435
Chen QF, Xiao S, Chye ML (2008) Overexpression of the Arabidopsis 10-kilodalton acyl-coenzyme a-binding protein ACBP6 enhances freezing tolerance. Plant Physiol 148(1):304–315
Chen QF, Xiao S, Qi W, Mishra G, Ma J, Wang M, Chye ML (2010) The Arabidopsis acbp1acbp2 double mutant lacking acyl-CoA-binding proteins ACBP1 and ACBP2 is embryo lethal. New Phytol 186:843–855
Choi D, Bostock RM, Avdiushko S, Hildebrand DF (1994) Lipid-derived signals that discriminate wound-and pathogen responsive isoprenoid pathways in plants: methyl jasmonate and the fungal elicitor arachidonic acid induce different 3-hydroxy-3-methylglutaryl-coenzyme A reductase genes and antimicrobial isoprenoids in Solanum tuberosum L. Proc Natl Acad Sci USA 91(6):2329–2333
Chye ML, Huang BQ, Zee SY (1999) Isolation of a gene encoding Arabidopsis membrane associated acyl-CoA binding protein and immunolocalization of its gene product. Plant J 18(2):205–214
Du ZY, Chen MX, Chen QF, Xiao S, Chye ML (2013a) Overexpression of Arabidopsis acyl-CoA-binding protein ACBP2 enhances drought tolerance. Plant Cell Environ 36:300–314
Du ZY, Chen MX, Chen QF, Xiao S, Chye ML (2013b) Arabidopsis acyl CoA-binding protein ACBP1 participates in the regulation of seed germination and seedling development. Plant J 74(2):294–309
Du ZY, Arias T, Meng W, Chye ML (2016) Plant acyl-CoA-binding proteins: an emerging family involved in plant development and stress responses. Prog Lipid Res 63:165–181
Emanuelsson O, Nielsen H, Brunak S, Heijne GV (2000) Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J Mol Biol 300(4):1005–1016
Fan J, Liu J, Culty M, Papadopoulos V (2010) Acyl-coenzyme A binding domain containing 3 (ACBD3; PAP7; GCP60): an emerging signaling molecule. Prog Lipid Res 49(3):218–234
Gao W, Xiao S, Li HY, Tsao SW, Chye ML (2009) Arabidopsis thaliana acyl-CoA-binding protein ACBP2 interacts with heavy-metalbinding farnesylated protein AtFP6. New Phytol 181:89–102
Gao W, Li HY, Xiao S, Chye ML (2010a) Acyl-CoA-binding protein 2 binds lysophospholipase 2 and lysoPC to promote tolerance to cadmium-induced oxidative stress in transgenic Arabidopsis. Plant J 62(2):989–1003
Gao W, Li HY, Xiao S, Chye ML (2010b) Protein interactors of acyl CoA-binding protein ACBP2 mediate cadmium tolerance in Arabidopsis. Plant Signal Behav 5(8):1025–1027
Gómez-Porras JL, Riaño-Pachón DM, Dreyer I, Mayer JE, Mueller-Roeber B (2007) Genome-wide analysis of ABA-responsive elements ABRE and CE3 reveals divergent patterns in Arabidopsis and rice. BMC Genomics 8(1):260
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acid Res 40(D1):D1178–D1186
Guelette BS, Benning UF, Hoffmann-Benning S (2012) Identification of lipids and lipid-binding proteins in phloem exudates from Arabidopsis thaliana. J Exp Bot 63(10):3603–3616
Guidotti A, Forchetti CM, Corda MG, Konkel D, Bennett CD, Costa E (1983) Isolation, characterization, and purification to homogeneity of an endogenous polypeptide with agonistic action on benzodiazepine receptors. P Natl Acad Sci USA 80(11):3531–3535
Guo ZH, Haslam RP, Michaelson LV, Yeung EC, Lung SC, Napier JA, Chye ML (2019) The overexpression of rice ACYL-CoA-BINDING PROTEIN2 increases grain size and bran oil content in transgenic rice. Plant J 100(6):1132–1147
Hattori T, Totsuka M, Hobo T, Kagaya Y, Yamamoto-Toyoda A (2002) Experimentally determined sequence requirement of ACGT-containing abscisic acid response element. Plant Cell Physiol 43(1):136–140
Hsiao AS, Haslam RP, Michaelson LV, Liao P, Chen QF, Sooriyaarachchi S, Mowbray SL, Napier JA, Tanner JA, Chye ML (2014) Arabidopsis cytosolic acyl-CoA-binding proteins ACBP4, ACBP5 and ACBP6 have overlapping but distinct roles in seed development. Biosci Rep 34(6):e00165
Hsiao AS, Yeung EC, Ye ZW, Chye ML (2015) The Arabidopsis cytosolic acyl-CoA-binding proteins play combinatory roles in pollen development. Plant Cell Physiol 56(2):322–333
Ibraheem O, Botha CE, Bradley G (2010) In silico analysis of cis-acting regulatory elements in 5’regulatory regions of sucrose transporter gene families in rice (Oryza sativa Japonica) and Arabidopsis thaliana. Comput Biol Chem 34(5–6):268–283
Ito J, Batth TS, Petzold CJ, Redding-Johanson AM, Mukhopadhyay A, Verboom R, Meyer EH, Millar AH, Heazlewood JL (2011) Analysis of the Arabidopsis cytosolic proteome high-lights subcellular partitioning of central plant metabolism. J Proteome Res 10(4):1571–1582
Jansson S, Douglas CJ (2007) Populus: a model system for plant biology. Annu Rev Plant Biol 58(1):435–458
Knudsen J, Jensen MV, Hansen JK, Faergeman NJ, Neergaard TB, Gaigg B (1999) Role of acylCoA binding protein in acylCoA transport, metabolism and cell signaling. Mol Cell Biochem 192(1–2):95–103
Kragelund BB, Andersen KV, Madsen JC, Knudsen J, Poulsen FM (1993) Three-dimensional structure of the complex between acyl-coenzyme A binding protein and palmitoyl-coenzyme A. J Mol Biol 230(4):1260–1277
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecul Biol Evolut 33:1870–1874
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23(21):2947–2948. https://doi.org/10.1093/bioinformatics/btm404
Leung KC, Li HY, Xiao S, Tse MH, Chye ML (2006) Arabidopsis ACBP3 is an extracellularly targeted acyl-CoA-binding protein. Planta 223(5):871–881
Li HY, Chye ML (2003) Membrane localization of Arabidopsis acyl-CoA binding protein ACBP2. Plant Mol Biol 51(4):483–492
Li HY, Chye ML (2004) Arabidopsis acyl-CoA-binding protein ACBP2 interacts with an ethylene-responsive element-binding protein, AtEBP, via its ankyrin repeats. Plant Mol Biol 54(2):233–243
Li HY, Xiao S, Chye ML (2008) Ethylene-and pathogen-inducible Arabidopsis acyl-CoA-binding protein 4 interacts with an ethylene-responsive element binding protein. J Exp Bot 59(14):3997–4006
Licausi F, Kosmacz M, Weits DA, Giuntoli B, Giorgi FM, Voesenek LACJ, Perata P, van Dongen JT (2011) Oxygen sensing in plants is mediated by an N-end rule pathway for protein destabilization. Nature 479(7373):419–422
Liu F, Zhang XB, Lu CM, Zeng XH, Li YJ, Fu DH, Wu G (2015) Non-specific lipid transfer proteins in plants: presenting new advances and an integrated functional analysis. J Exp Bot 66(19):5663–5681
Lung SC, Chye ML (2016) Deciphering the roles of acyl-CoA-binding proteins in plant cells. Protoplasma 253(5):1177–1195
Mattick JS, Gagen MJ (2001) The evolution of controlled multitasked gene networks: the role of introns and other noncoding RNAs in the development of complex organisms. Mol Biol Evol 18(9):1611–1630
Meng W, Su YC, Saunders RM, Chye ML (2011) The rice acyl-CoA-binding protein gene family: phylogeny, expression and functional analysis. New Phytol 189(4):1170–1184
Meng W, Hsiao AS, Gao CJ, Jiang LW, Chye ML (2014) Subcellular localization of rice acyl-CoA-binding proteins (ACBPs) indicates that OsACBP6::GFP is targeted to the peroxisomes. New Phytol 203(2):469–482
Meng W, Xu LJ, Du ZY, Wang F, Zhang R, Song XS, Lam SM, Shui GH, Li YH, Chye ML (2020) Rice acyl-coa-binding protein6 affects acyl-coa homeostasis and growth in rice. Rice 13(1):75
Napier JA, Haslam RP (2010) As simple as ACB-new insights into the role of acyl-CoA-binding proteins in Arabidopsis. New Phytol 186(4):781–783
Narayanan SP, Pan L, Taylor P, Lo C, Chye M (2019) Overexpression of a monocot acyl-coA-binding protein confers broad-spectrum pathogen protection in a dicot. Proteomics 19(12):e1800368
Narayanan SP, Lung SC, Liao P, Lo C, Chye ML (2020) The overexpression of OsACBP5 protects transgenic rice against necrotrophic, hemibiotrophic and biotrophic pathogens. Sci Rep-UK 10(1):14918
Nie Z, Wang YH, Wu CT, Li Y, Kang GJ, Qin HD, Zeng RZ (2018) Acyl-CoA-binding protein family members in laticifers are possibly involved in lipid and latex metabolism of Hevea brasiliensis (the Para rubber tree). BMC Genomics 19(1):5
Ohlrogge J, Browse J (1995) Lipid biosynthesis. Plant Cell 7(7):957–970
Ohlrogge JB, Jaworski JG (1997) Regulation of fatty acid syn-thesis. Annu Rev Plant Physiol Plant Mol Biol 48:109–136
Pastor S, Sethumadhavan K, Ullah AHJ, Gidda S, Cao H, Mason C, Chapital D, Scheffler B, Mullen R, Dyer J, Shockey J (2013) Molecular properties of the class III subfamily of acyl-coenzyme A binding proteins from tung tree (Vernicia fordii). Plant Sci 203–204:79–88
Paterson AH, Bowers JE, Bruggmann R, Dubchak I, Grimwood J, Gundlach H, Haberer G, Hellsten U, Mitros T, Poliakov A, Schmutz J, Spannagl M, Tang HB, Wang XY, Wicker T, Bharti AK, Chapman J, Feltus FA, Gowik U, Grigoriev IV, Lyons E, Maher CA, Martis M, Narechania A, Otillar RP, Penning BW, Salamov AA, Wang Y, Zhang LF, Carpita NC, Freeling M, Gingle AR, Hash CT, Keller B, Klein P, Kresovich S, McCann MC, Ming R, Peterson DG, Mehboob-ur-Rahman WD, Westhoff P, Mayer KF, Messing J, Rokhsar DS (2009) The Sorghum bicolor genome and the diversification of grasses. Nature 457(7229):551–556
Qin PF, Shang XG, Song J, Guo WZ (2016) Genome-wide identification of Acyl-CoA-Binding Protein (ACBP) gene family and their functional analysis in abiotic stress tolerance in cotton. Acta Agron Sin 42(11):1577–1591
Staswick PE (2008) JAZing up jasmonate signaling. Trends Plant Sci 13(2):66–71
Sundell D, Mannapperuma C, Netotea S, Delhomme N, Lin YC, Sjodin A, Van de Peer Y, Jansson S, Hvidsten TR, Street NR (2015) The plant genome integrative explorer resource: PlantGenIE.org. New Phytol 208:1149–1156
Suzui N, Nakamura S, Fujiwara T, Hayashi H, Yoneyama T (2006) A putative acyl-CoA-binding protein is a major phloem sap protein in rice (Oryza sAtiva L.). J Exp Bot 57:2571–2576
Takato H, Shimidzu M, Ashizawa Y, Takei H, Suzuki S (2013) An acylCoA-binding protein from grape that is induced through ER stress confers morphological changes and disease resistance in Arabidopsis. J Plant Physiol 170(6):591–600
Turner JG, Ellis C, Devoto A (2002) The jasmonate signal pathway. Plant Cell 14:S153-164
Tuskan GA, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N, Ralph S, Rombauts S, Salamov A, Schein J, Sterck L, Aerts A, Bhalerao RR, Bhalerao RP, Blaudez D, Boerjan W, Brun A, Brunner A, Busov V, Campbell M, Carlson J, Chalot M, Chapman J, Chen GL, Cooper D, Coutinho PM, Couturier J, Covert S, Cronk Q, Cunningham R, Davis J, Degroeve S, Dejardin A, Depamphilis C, Detter J, Dirks B, Dubchak I, Duplessis S, Ehlting J, Ellis B, Gendler K, Goodstein D, Gribskov M, Grimwood J, Groover A, Gunter L, Hamberger B, Heinze B, Helariutta Y, Henrissat B, Holligan D, Holt R, Huang W, Islam-Faridi N, Jones S, Jones-Rhoades M, Jorgensen R, Joshi C, Kangasjarvi J, Karlsson J, Kelleher C, Kirkpatrick R, Kirst M, Kohler A, Kalluri U, Larimer F, Leebens-Mack J, Leple JC, Locascio P, Lou Y, Lucas S, Martin F, Montanini B, Napoli C, Nelson DR, Nelson C, Nieminen K, Nilsson O, Pereda V, Peter G, Philippe R, Pilate G, Poliakov A, Razumovskaya J, Richardson P, Rinaldi C, Ritland K, Rouze P, Ryaboy D, Schmutz J, Schrader J, Segerman B, Shin H, Siddiqui A, Sterky F, Terry A, Tsai CJ, Uberbacher E, Unneberg P, Vahala J, Wall K, Wessler S, Yang G, Yin T, Douglas C, Marra M, Sandberg G, Van de Peer Y, Rokhsar D (2006) The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313:1596–1604
Walz C, Giavalisco P, Schad M, Juenger M, Klose J, Kehr J (2004) Proteomics of curcurbit phloem exudate reveals a network of defence proteins. Phytochemistry 65(12):1795–1804
Wang SZ, Zhang X, Dai SJ, Li Y (2019) Advances in research regarding the function of the ACBP family in plants. Pratacultural Science 36(10):2535–2548
Xia Y, Yu KS, Gao QM, Wilson EV, Navarre D, Kachroo P, Kachroo A (2012) Acyl CoA binding proteins are required for cuticle formation and plant responses to microbes. Front Plant Sci 3:224
Xiao S, Chye ML (2009) An Arabidopsis family of six acyl-CoA-binding proteins has three cytosolic members. Plant Physiol Biochem 47(6):479–484
Xiao S, Chye ML (2011a) New roles for acyl-CoA-binding proteins (ACBPs) in plant development, stress responses and lipid metabolism. Prog Lipid Res 50(2):141–151
Xiao S, Chye ML (2011b) Overexpression of Arabidopsis ACBP3 enhances NPR1-dependent plant resistance to Pseudomonas syringae pv tomAto DC3000. Plant Physiol 156(4):2069–2081
Xiao S, Li HY, Zhang JP, Chan SW, Chye ML (2008) Arabidopsis acyl-CoA-binding proteins ACBP4 and ACBP5 are subcellularly localizedto the cytosol and ACBP4 depletion affects membrane lipid composition. Plant Mol Biol 68(6):571–583
Xiao S, Gao W, Chen QF, Chan SW, Zheng SX, Ma J, Wang M, Welti R, Chye ML (2010) Overexpression of Arabidopsis acyl-CoA binding protein ACBP3 promotes starvation-induced and age-dependent leaf senescence. Plant Cell 22(6):1463–1482
Xie LJ, Yu LJ, Chen QF, Wang FZ, Huang L, Xia FN, Zhu TR, Wu JX, Yin J, Liao B, Yao N, Shu WS, Xiao S (2015) Arabidopsis acyl-CoA-binding protein ACBP3 participates in plant response to hypoxia by modulating very-long-chain fatty acid metabolism. Plant J 81(1):53–67
Xu WJ, Cheng H, Zhu SR, Cheng JY, Ji HH, Zhang BC, Cao SQ, Wang C, Tong GM, Zhen C, Mu LQ, Zhou YH, Cheng YX (2021) Functional understanding of secondary cell wall cellulose synthases in Populus trichocarpa via the Cas9/gRNA-induced gene knockouts. New Phytol 231(4):1478–1495
Xue Y, Xiao S, Kim J, Lung SC, Chen L, Tanner JA, Suh MC, Chye ML (2014) Arabidopsis membrane-associated acyl-CoA-binding protein ACBP1 is involved in stem cuticle formation. J Exp Bot 65(18):5473–5483
Yang WQ, Zhang W, Wang XX (2017) Post-translational control of ABA signalling: the roles of protein phosphorylation and ubiquitination. Plant Biotechnol J 15(1):4–14
Ye ZW, Chye ML (2016) Plant cytosolic acyl-CoA-binding proteins. Lipids 51(1):1–13
Yu ZP, Duan XB, Luo L, Dai SJ, Ding ZJ, Xia GM (2020) How plant hormones mediate salt stress responses. Trends Plant Sci 25(11):1117–1130
Yurchenko OP, Nykiforuk CL, Moloney MM, Ståhl U, Banaś A, Stymne S, Weselake RJ (2009) A 10-kDa acyl-CoA-binding protein (ACBP) from Brassica napus enhances acyl exchange between acyl-CoA and phosphatidylcholine. Plant Biotechnol J 7(7):602–610
Yurchenko O, Singer SD, Nykiforuk CL, Gidda S, Mullen RT, Moloney MM, Weselake RJ (2014) Production of a Brassica napus lowmolecular mass acyl-coenzyme A-binding protein in Arabidopsis alters the acyl-coenzyme A pool and acyl composition of oil in seeds. Plant Physiol 165(2):550–560
Zhu JT, Li WJ, Zhou YY, Pei LM, Liu JJ, Xia XY, Che RH, Li H (2021) Molecular characterization, expression and functional analysis of acyl-CoA-binding protein gene family in maize (Zea mays). BMC Plant Biol 21(1):94
Author information
Authors and Affiliations
Contributions
Yu Chang and Xinru Xu contributed equally to this work.
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Project funding: The work was supported by grants from the Fundamental Research Funds for the Central Universities (Nos. 2572018B03), the College Students Innovations Special Project funded by NEFU (No. 202010225173), the Fundamental Research Funds for the Central Universities (Nos. 2572019CT03), and the Natural Science Foundation of Heilongjiang Province (No. ZD2019C003).
The online version is available at http://www.springerlink.com.
Corresponding editor: Yu Lei
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chang, Y., Xu, X., Zheng, H. et al. Genome-wide identification and characterization of ACBP gene family in Populus reveal salinity alkali-responsive profiles. J. For. Res. 34, 481–496 (2023). https://doi.org/10.1007/s11676-022-01485-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11676-022-01485-2