Abstract
Background
For patients undergoing resection of colorectal liver metastases (CLMs), the prognostic role of somatic gene alterations is increasingly recognized. F-box/WD repeat–containing protein 7 (FBXW7) is a tumor suppressor gene found in approximately 10% of patients with colorectal cancer. The aim of this study is to assess the association of FBXW7 with overall survival after CLM resection.
Methods
Patients who underwent initial CLM resection during 2001–2016 and had genetic sequencing data were studied. Risk factors for overall survival (OS) were evaluated with Cox proportional hazards models using backward elimination.
Results
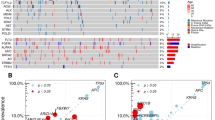
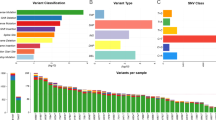
Of 2045 patients who underwent CLM resection during the study period, 476 were included. The majority (90.5%) underwent prehepatectomy chemotherapy. A total of 27 patients (5.7%) had FBXW7 alteration, along with 240 (50.4%) RAS, 337 (70.8%) TP53, 51 (10.7%) SMAD4, and 27 (5.7%) BRAF. Cox proportional hazards model analyses including 5 somatic gene alteration status and 12 clinicopathologic factors revealed FBXW7(hazard ratio [HR] 1.99, P = 0.015), BRAF (HR 2.47, P = 0.023), RAS (HR 2.42, P < 0.001), TP53 (HR 2.00, P < 0.001), and SMAD4 alterations (HR 1.90, P = 0.004) as significantly associated with OS, together with three clinicopathologic factors, prehepatectomy chemotherapy > 6 cycles (HR 1.51, P = 0.021), number of CLM (HR 1.05, P = 0.007), and largest liver metastasis diameter (HR 1.07, P = 0.023). The covariate-adjusted 5-year OS was significantly lower in patients with FBXW7 alteration than in patients with FBXW7 wild-type (40.4% vs.59.4%, P = 0.015).
Conclusions
FBXW7 alterations are associated with worse survival after CLM resection. The information on multiple somatic gene alterations is imperative for risk stratification and patient selection for CLM resection.




Similar content being viewed by others
Abbreviations
- CLM:
-
colorectal liver metastases
- FBXW7 :
-
F-box/WD repeat–containing protein 7
- OS:
-
overall survival
- HR:
-
hazard ratio
- CI:
-
confidence intervals
- VEGF:
-
vascular endothelial growth factor
- EGFR:
-
epidermal growth factor receptor
- MAPK:
-
mitogen-activated protein kinase
- TGF-β:
-
transforming growth factor-β
References
Fong Y, Fortner J, Sun RL, Brennan MF, Blumgart LH. Clinical score for predicting recurrence after hepatic resection for metastatic colorectal cancer: analysis of 1001 consecutive cases. Annals of surgery. 1999;230(3):309–18; discussion 18–21.
Rees M, Tekkis PP, Welsh FK, O’Rourke T, John TG. Evaluation of long-term survival after hepatic resection for metastatic colorectal cancer: a multifactorial model of 929 patients. Annals of surgery. 2008;247(1):125-35. https://doi.org/10.1097/SLA.0b013e31815aa2c2.
Nash GM, Gimbel M, Shia J, Nathanson DR, Ndubuisi MI, Zeng ZS et al. KRAS mutation correlates with accelerated metastatic progression in patients with colorectal liver metastases. Annals of surgical oncology. 2010;17(2):572-8. https://doi.org/10.1245/s10434-009-0605-3.
Vauthey JN, Zimmitti G, Kopetz SE, Shindoh J, Chen SS, Andreou A et al. RAS mutation status predicts survival and patterns of recurrence in patients undergoing hepatectomy for colorectal liver metastases. Annals of surgery. 2013;258(4):619–26; discussion 26–7. https://doi.org/10.1097/SLA.0b013e3182a5025a.
Brudvik KW, Kopetz SE, Li L, Conrad C, Aloia TA, Vauthey JN. Meta-analysis of KRAS mutations and survival after resection of colorectal liver metastases. The British journal of surgery. 2015;102(10):1175-83. https://doi.org/10.1002/bjs.9870.
Amikura K, Akagi K, Ogura T, Takahashi A, Sakamoto H. The RAS mutation status predicts survival in patients undergoing hepatic resection for colorectal liver metastases: The results from a genetic analysis of all-RAS. Journal of surgical oncology. 2018;117(4):745-55. https://doi.org/10.1002/jso.24910.
Wang K, Liu W, Yan XL, Li J, Xing BC. Long-term postoperative survival prediction in patients with colorectal liver metastasis. Oncotarget. 2017;8(45):79927-34. https://doi.org/10.18632/oncotarget.20322.
Chun YS, Passot G, Yamashita S, Nusrat M, Katsonis P, Loree JM et al. Deleterious Effect of RAS and Evolutionary High-risk TP53 Double Mutation in Colorectal Liver Metastases. Annals of surgery. 2019;269(5):917-23. https://doi.org/10.1097/SLA.0000000000002450.
Kawaguchi Y, Lillemoe HA, Panettieri E, Chun YS, Tzeng CD, Aloia TA et al. Conditional Recurrence-Free Survival after Resection of Colorectal Liver Metastases: Persistent Deleterious Association with RAS and TP53 Co-Mutation. Journal of the American College of Surgeons. 2019;229(3):286-94 e1. https://doi.org/10.1016/j.jamcollsurg.2019.04.027.
Mizuno T, Cloyd JM, Vicente D, Omichi K, Chun YS, Kopetz SE et al. SMAD4 gene mutation predicts poor prognosis in patients undergoing resection for colorectal liver metastases. European journal of surgical oncology : the journal of the European Society of Surgical Oncology and the British Association of Surgical Oncology. 2018;44(5):684-92. https://doi.org/10.1016/j.ejso.2018.02.247.
Lang H, Baumgart J, Heinrich S, Tripke V, Passalaqua M, Maderer A et al. Extended Molecular Profiling Improves Stratification and Prediction of Survival After Resection of Colorectal Liver Metastases. Annals of surgery. 2019;270(5):799-805. https://doi.org/10.1097/SLA.0000000000003527.
Datta J, Smith JJ, Chatila WK, McAuliffe JC, Kandoth C, Vakiani E et al. Co-Altered Ras/B-raf and TP53 is Associated with Extremes of Survivorship and Distinct Patterns of Metastasis in Metastatic Colorectal Cancer Patients. Clinical cancer research : an official journal of the American Association for Cancer Research. 2019. https://doi.org/10.1158/1078-0432.CCR-19-2390.
Gagniere J, Dupre A, Gholami SS, Pezet D, Boerner T, Gonen M et al. Is Hepatectomy Justified for BRAF Mutant Colorectal Liver Metastases?: A Multi-institutional Analysis of 1497 Patients. Annals of surgery. 2018. https://doi.org/10.1097/SLA.0000000000002968.
Margonis GA, Buettner S, Andreatos N, Kim Y, Wagner D, Sasaki K et al. Association of BRAF Mutations With Survival and Recurrence in Surgically Treated Patients With Metastatic Colorectal Liver Cancer. JAMA surgery. 2018;153(7):e180996. https://doi.org/10.1001/jamasurg.2018.0996.
Malapelle U, Pisapia P, Sgariglia R, Vigliar E, Biglietto M, Carlomagno C et al. Less frequently mutated genes in colorectal cancer: evidences from next-generation sequencing of 653 routine cases. Journal of clinical pathology. 2016;69(9):767-71. https://doi.org/10.1136/jclinpath-2015-203403.
Korphaisarn K, Morris VK, Overman MJ, Fogelman DR, Kee BK, Raghav KPS et al. FBXW7 missense mutation: a novel negative prognostic factor in metastatic colorectal adenocarcinoma. Oncotarget. 2017;8(24):39268-79. https://doi.org/10.18632/oncotarget.16848.
Brouquet A, Abdalla EK, Kopetz S, Garrett CR, Overman MJ, Eng C et al. High survival rate after two-stage resection of advanced colorectal liver metastases: response-based selection and complete resection define outcome. Journal of clinical oncology : official journal of the American Society of Clinical Oncology. 2011;29(8):1083-90. https://doi.org/10.1200/JCO.2010.32.6132.
Vauthey JN, Chaoui A, Do KA, Bilimoria MM, Fenstermacher MJ, Charnsangavej C et al. Standardized measurement of the future liver remnant prior to extended liver resection: methodology and clinical associations. Surgery. 2000;127(5):512-9. https://doi.org/10.1067/msy.2000.105294.
Kawaguchi Y, Lillemoe HA, Vauthey JN. Dealing with an insufficient future liver remnant: Portal vein embolization and two-stage hepatectomy. Journal of surgical oncology. 2019;119(5):594-603. https://doi.org/10.1002/jso.25430.
Kawaguchi Y, Kopetz S, Lillemoe HA, Hwang H, Wang X, Tzeng CD, Chun YS, Aloia TA, Vauthey JN. A new surveillance algorithm after resection of colorectal liver metastases based on changes in recurrence risk and ras mutation status. J Natl Compr Canc Netw. 2020;18(11):1500-1508. https://doi.org/10.6004/jnccn.2020.7596.
Kawaguchi Y, Kopetz S, Newhook TE, De Bellis M, Chun YS, Tzeng CD et al. Mutation Status of RAS, TP53, and SMAD4 is Superior to Mutation Status of RAS Alone for Predicting Prognosis after Resection of Colorectal Liver Metastases. Clinical cancer research : an official journal of the American Association for Cancer Research. 2019;25(19):5843-51. https://doi.org/10.1158/1078-0432.CCR-19-0863.
Singh RR, Patel KP, Routbort MJ, Reddy NG, Barkoh BA, Handal B et al. Clinical validation of a next-generation sequencing screen for mutational hotspots in 46 cancer-related genes. The Journal of molecular diagnostics : JMD. 2013;15(5):607-22. https://doi.org/10.1016/j.jmoldx.2013.05.003.
Amin MB, Greene FL, Edge SB, Compton CC, Gershenwald JE, Brookland RK et al. The Eighth Edition AJCC Cancer Staging Manual: Continuing to build a bridge from a population-based to a more “personalized” approach to cancer staging. CA: a cancer journal for clinicians. 2017;67(2):93-9. https://doi.org/10.3322/caac.21388.
Vauthey JN, Kopetz SE. From multidisciplinary to personalized treatment of colorectal liver metastases: 4 reasons to consider RAS. Cancer. 2013;119(23):4083-5. https://doi.org/10.1002/cncr.28348.
Chun YS, Passot G, Yamashita S, Nusrat M, Katsonis P, Loree JM et al. Deleterious Effect of RAS and Evolutionary High-risk TP53 Double Mutation in Colorectal Liver Metastases. Annals of surgery. 2017. https://doi.org/10.1097/SLA.0000000000002450.
Summers MG, Smith CG, Maughan TS, Kaplan R, Escott-Price V, Cheadle JP. BRAF and NRAS Locus-Specific Variants Have Different Outcomes on Survival to Colorectal Cancer. Clinical cancer research : an official journal of the American Association for Cancer Research. 2017;23(11):2742-9. https://doi.org/10.1158/1078-0432.CCR-16-1541.
Schirripa M, Cremolini C, Loupakis F, Morvillo M, Bergamo F, Zoratto F et al. Role of NRAS mutations as prognostic and predictive markers in metastatic colorectal cancer. International journal of cancer Journal international du cancer. 2015;136(1):83-90. https://doi.org/10.1002/ijc.28955.
Cercek A, Braghiroli MI, Chou JF, Hechtman JF, Kemeny N, Saltz L et al. Clinical Features and Outcomes of Patients with Colorectal Cancers Harboring NRAS Mutations. Clinical cancer research : an official journal of the American Association for Cancer Research. 2017;23(16):4753-60. https://doi.org/10.1158/1078-0432.CCR-17-0400.
Kawaguchi Y, Lillemoe HA, Vauthey JN. Gene mutation and surgical technique: Suggestion or more? Surgical oncology. 2019. https://doi.org/10.1016/j.suronc.2019.07.004.
Makuch RW. Adjusted survival curve estimation using covariates. Journal of chronic diseases. 1982;35(6):437-43.
Ghali WA, Quan H, Brant R, van Melle G, Norris CM, Faris PD et al. Comparison of 2 methods for calculating adjusted survival curves from proportional hazards models. JAMA : the journal of the American Medical Association. 2001;286(12):1494-7.
Sanchez-Vega F, Mina M, Armenia J, Chatila WK, Luna A, La KC et al. Oncogenic Signaling Pathways in The Cancer Genome Atlas. Cell. 2018;173(2):321-37 e10. https://doi.org/10.1016/j.cell.2018.03.035.
Akhoondi S, Sun D, von der Lehr N, Apostolidou S, Klotz K, Maljukova A et al. FBXW7/hCDC4 is a general tumor suppressor in human cancer. Cancer research. 2007;67(19):9006-12. https://doi.org/10.1158/0008-5472.CAN-07-1320.
Jardim DL, Wheler JJ, Hess K, Tsimberidou AM, Zinner R, Janku F et al. FBXW7 mutations in patients with advanced cancers: clinical and molecular characteristics and outcomes with mTOR inhibitors. PloS one. 2014;9(2):e89388. https://doi.org/10.1371/journal.pone.0089388.
Lobry C, Oh P, Mansour MR, Look AT, Aifantis I. Notch signaling: switching an oncogene to a tumor suppressor. Blood. 2014;123(16):2451-9. https://doi.org/10.1182/blood-2013-08-355818.
Masuda K, Ishikawa Y, Onoyama I, Unno M, de Alboran IM, Nakayama KI et al. Complex regulation of cell-cycle inhibitors by Fbxw7 in mouse embryonic fibroblasts. Oncogene. 2010;29(12):1798-809. https://doi.org/10.1038/onc.2009.469.
Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487(7407):330-7. https://doi.org/10.1038/nature11252.
Acknowledgements
The authors thank Ms. Ruth Haynes for the administrative support in the preparation of this manuscript.
Statement of Author Contribution
Substantial contributions to:
The conception or design of the work: YK, TN, JNV
The acquisition, analysis, or interpretation of data for the work: YK, TN, HT, CWT, YSH, TA, SK, JNV
Drafting the work or revising it critically for important intellectual content: all authors
Final approval of the version to be published: all authors
Agreement to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved: all authors
Funding
This research was supported in part by the National Institutes of Health through MD Anderson Cancer Center Support Grant, CA016672.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
Nothing to disclose.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Previous communication to a society or meeting: This study was presented on June 3, 2020 in the Society for Surgery of the Alimentary Tract Wednesday Webinars due to the cancellation of the 61th Annual Meeting of the Society for Surgery of the Alimentary Tract Plenary Session.
Supplementary Information
Supplementary Figure 1
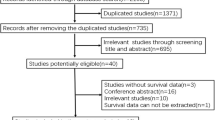
Patient selection. *In patients who completed 2-stage hepatectomy, only the second-stage hepatectomy was included. The first-stage hepatectomy was excluded not to include patients twice in our analysis. (DOCX 357 kb)
Supplementary Figure 2
Overall survival (OS) by TP53 and FBXW7 alteration status. (A) OS curves. (B) OS curves after adjustment for somatic gene alteration status (BRAF, RAS, and SMAD4), prehepatectomy chemotherapy (> 6 cycles vs. ≤ 6 cycles or no prehepatectomy chemotherapy), number of CLM, and largest liver metastasis diameter. (DOCX 380 kb)
ESM 1
(DOCX 16 kb)
Rights and permissions
About this article
Cite this article
Kawaguchi, Y., Newhook, T.E., Tran Cao, H.S. et al. Alteration of FBXW7 is Associated with Worse Survival in Patients Undergoing Resection of Colorectal Liver Metastases. J Gastrointest Surg 25, 186–194 (2021). https://doi.org/10.1007/s11605-020-04866-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11605-020-04866-2