Summary
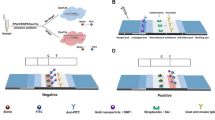
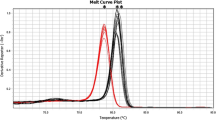
Pseudomonas aeruginosa (P. aeruginosa) is a major opportunistic pathogen in hospital-acquired infections. Thus, early diagnosis is the best strategy for fighting against these infections. In this report, we incorporated multiple cross displacement amplification (MCDA) combined with the malachite green (MG) for rapid, sensitive, specific and visual detection of P. aeruginosa by targeting the oprl gene. The MCDA-MG assay was conducted at 67°C for only 40 min during the amplification stage, and then products were directly detected by using colorimetric indicators (MG), eliminating the use of an electrophoresis instrument or amplicon analysis equipment. The entire process, including specimen processing (35 min), amplification (40 min) and detection (5 min), can be finished within 80 min. All 30 non-P. aeruginosa strains tested negative, indicating the high specificity of the MCDA primers. The analytical sensitivity of the MCDA-MG assay was 100 fg of genomic templates per reaction in pure culture, which was in complete accordance with MCDA by gel electrophoresis and real-time turbidity. The assay was also successfully applied to detecting P. aeruginosa in stool samples. Therefore, the rapidity, simplicity, and nearly equipment-free platform of the MCDA-MG technique make it possible for clinical diagnosis, and more.
Similar content being viewed by others
References
Tang Y, Ali Z, Zou J, et al. Detection of Pseudomonas aeruginosa based on magnetic enrichment and nested PCR. J Nanosci Nanotechnol, 2014,14(7):4886–4890
Manajit O, Longyant S, Sithigorngul P, et al. Development of uracil-DNA-glycosylase-supplemented loop-mediated isothermal amplification coupled with nanogold probe (UDG-LAMP-AuNP) for specific detection of Pseudomonas aeruginosa. Mol Med Rep, 2018,17(4):5734–5743
Aghamollaei H, Moghaddam MM, Kooshki H, et al. Detection of Pseudomonas aeruginosa by a triplex polymerase chain reaction assay based on lasI/R and gyrB genes. J Infect Public Health, 2015,8(4):314–322
Shi H, Chen Z, Kan J Development of loop-mediated isothermal amplification assays for genotyping of Type III Secretion System in Pseudomonas aeruginosa. Lett Appl Microbiol, 2015,61(4):361–366
Salman M, Ali A, Haque A A novel multiplex PCR for detection of Pseudomonas aeruginosa: A major cause of wound infections. Pak J Med Sci, 2013,29(4):957–961
Hery-Arnaud G, Nowak E, Caillon J, et al. Evaluation of quantitative PCR for early diagnosis of Pseudomonas aeruginosa infection in cystic fibrosis: a prospective cohort study. Clin Microbiol Infect, 2017,23(3):203–207
Lee CS, Wetzel K, Buckley T, et al. Rapid and sensitive detection of Pseudomonas aeruginosa in chlorinated water and aerosols targeting gyrB gene using real-time PCR. J Appl Microbiol, 2011,111(4):893–903
Jami Al-Ahmadi G, Zahmatkesh Roodsari R Fast and specific detection of Pseudomonas Aeruginosa from other pseudomonas species by PCR. Ann Burns Fire Disasters, 2016,29(4):264–267
Goto M, Shimada K, Sato A, et al. Rapid detection of Pseudomonas aeruginosa in mouse feces by colorimetric loop-mediated isothermal amplification. J Microbiol Methods, 2010,81(3):247–252
Zhao X, Wang L, Li Y, et al. Development and application of a loop-mediated isothermal amplification method on rapid detection of Pseudomonas aeruginosa strains. World J Microbiol Biotechnol, 2011,27:181–184
Wang Y, Wang Y, Zhang L, et al. Multiplex, Rapid, and Sensitive Isothermal Detection of Nucleic-Acid Sequence by Endonuclease Restriction-Mediated Real- Time Multiple Cross Displacement Amplification. Front Microbiol, 2016,7:753
Wang Y, Wang Y, Ma AJ, et al. Rapid and Sensitive Isothermal Detection of Nucleic-acid Sequence by Multiple Cross Displacement Amplification. Sci Rep, 2015,5:11902
Wang Y, Li H, Li D, et al. Multiple Cross Displacement Amplification Combined with Gold Nanoparticle- Based Lateral Flow Biosensor for Detection of Vibrio parahaemolyticus. Front Microbiol, 2016,7:2047
Wang Y, Wang Y, Xu J, et al. Development of Multiple Cross Displacement Amplification Label-Based Gold Nanoparticles Lateral Flow Biosensor for Detection of Shigella spp. Front Microbiol, 2016,7:1834
Wang Y, Yan W, Fu S, et al. Multiple Cross Displacement Amplification Coupled With Nanoparticles-Based Lateral Flow Biosensor for Detection of Staphylococcus aureus and Identification of Methicillin-Resistant S aureus. Front Microbiol, 2018,9:907
van der Waaij D Colonization resistance of the digestive tract: clinical consequences and implications. J Antimicrob Chemother, 1982,10(4):263–270
Hazlett LD, Rosen DD, Berk RS. Age-related susceptibility to Pseudomonas aeruginosa ocular infections in mice. Infect Immun, 1978,20(1):25–29
Ohman DE, Burns RP, Iglewski BH. Corneal infections in mice with toxin A and elastase mutants of Pseudomonas aeruginosa. J Infect Dis, 1980,142(4):547–555
Dong D, Zou D, Liu H, et al. Rapid detection of Pseudomonas aeruginosa targeting the toxA gene in intensive care unit patients from Beijing, China. Front Microbiol, 2015,6:1100
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors have declared that no competing interests exist.
Rights and permissions
About this article
Cite this article
Li, Dx., Shu, Gl., Wang, Wj. et al. Simple, Rapid and Sensitive Detection of Pseudomonas aeruginesa by Colorimetric Multiple Cross Displacement Amplification. CURR MED SCI 40, 372–379 (2020). https://doi.org/10.1007/s11596-020-2169-1
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11596-020-2169-1