Abstract
Purpose
Accurate segmentation of left ventricle (LV) is essential for the cardiac function analysis. However, it is labor intensive and time consuming for radiologists to delineate LV boundary manually. In this paper, we present a novel self-correcting framework for the fully automatic LV segmentation.
Methods
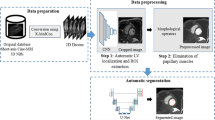
Firstly, a time-domain method is designed to extract a rectangular region of interest around the heart. Then, the simplified pulse-coupled neural network (SPCNN) is employed to locate the LV cavity. Different from the existing approaches, SPCNN can realize the self-correcting segmentation due to its parameter controllability. Subsequently, the post-processing based on the maximum gradient searching is proposed to obtain the accurate endocardium. Finally, a new external force based on the shape similarity is defined and integrated into the gradient vector flow (GVF) snake with the balloon force to segment the epicardium.
Results
We obtain encouraging segmentation results tested on the database provided by MICCAI 2009. The average percentage of good contours is 92.26 %, the average perpendicular distance is 2.38 mm, and the overlapping dice metric is 0.89. Besides, the experiment results show good correlations between the automatic segmentation and the manual delineation (for the LV ejection fraction and the LV myocardial mass, the correlation coefficients R are 0.9683 and 0.9278, respectively).
Conclusion
We propose an effective and fast method combing the SPCNN and the improved GVF for the automatic segmentation of LV.








Similar content being viewed by others
References
Petitjean C, Dacher JN (2011) A review of segmentation methods in short axis cardiac MR images. Med Image Anal 15(2):169–184. doi:10.1016/j.media.2010.12.004
Kurkure U, Pednekar A, Muthupillai R, Flamm SD, Kakadiaris IA (2009) Localization and segmentation of left ventricle in cardiac cine-MR images. IEEE Trans Biomed Eng 56(5):1360–1370. doi:10.1109/TBME.2008.2005957
Hu H, Liu H, Gao Z, Huang L (2013) Hybrid segmentation of left ventricle in cardiac MRI using gaussian-mixture model and region restricted dynamic programming. Magn Reson Imaging 31(4):575–584. doi:10.1016/j.mri.2012.10.004
Huang S, Liu J, Lee LC, Venkatesh SK, Teo LLS, Au C, Nowinski WL (2011) An image-based comprehensive approach for automatic segmentation of left ventricle from cardiac short axis cine MR images. J Digit Imaging 24(4):598–608. doi:10.1007/s10278-010-9315-4
Liu H, Hu H, Xu X, Song E (2012) Automatic left ventricle segmentation in cardiac MRI using topological stable-state thresholding and region restricted dynamic programming. Acad Radiol 19(6):723–731. doi:10.1016/j.acra.2012.02.011
Tsai I-C, Huang Y-L, Liu P-T, Chen M-C (2012) Left ventricular myocardium segmentation on delayed phase of multi-detector row computed tomography. Int J Comput Assist Radiol Surg 7(5):737–751. doi:10.1007/s11548-012-0688-3
Albà X, Figueras i Ventura RM, Lekadir K, Tobon-Gomez C, Hoogendoorn C, Frangi AF (2014) Automatic cardiac LV segmentation in MRI using modified graph cuts with smoothness and interslice constraints. Magn Reson Med 72(6):1775–1784. doi:10.1002/mrm.25079
Tufvesson J, Hedström E, Steding-ehrenborg K, Carlsson M, Arheden H, Heiberg E (2015) Validation and development of a new automatic algorithm for time-resolved segmentation of the left ventricle in magnetic resonance imaging. J Cardiovasc Magn Reson 17(Suppl 1):68–80. doi:10.1186/1532-429X-17-S1-P68
Dakua SP (2015) LV segmentation using stochastic resonance and evolutionary cellular automata. Int J Pattern Recognit Artif Intell 29:1–26. doi:10.1142/S0218001415570025
Dakua SP (2014) AnnularCut: a graph-cut design for left ventricle segmentation from magnetic resonance images. IET Image Process 8(1):1–11. doi:10.1049/iet-ipr.2013.0088
Wang Y, Jia Y (2006) Segmentation of the left ventricle from MR images via snake models incorporating shape similarities. In: Proceeding international conference on image processing ICIP, pp 213–216. doi:10.1109/ICIP.2006.312458
Grosgeorge D, Petitjean C, Caudron J, Fares J, Dacher JN (2011) Automatic cardiac ventricle segmentation in MR images: a validation study. Int J Comput Assist Radiol Surg 6(5):573–581. doi:10.1007/s11548-010-0532-6
Constantinides C, Roullot E, Lefort M, Frouin F (2012) Fully automated segmentation of the left ventricle applied to cine MR images: description and results on a database of 45 subjects. In: Proceeding annual international conference on IEEE Engineering in Medicine and Biology Society EMBS, pp 3207–3210. doi:10.1109/EMBC.2012.6346647
Wu Y, Wang Y, Jia Y (2013) Segmentation of the left ventricle in cardiac cine MRI using a shape-constrained snake model. Comput Vis Image Underst 117(9):990–1003. doi:10.1016/j.cviu.2012.12.008
Queirós S, Barbosa D, Heyde B, Morais P, Vilaça JL, Friboulet D, Bernard O, D’hooge J (2014) Fast automatic myocardial segmentation in 4D cine CMR datasets. Med Image Anal 18(7):1115–1131. doi:10.1016/j.media.2014.06.001
Faghih Roohi S, Aghaeizadeh Zoroofi R (2013) 4D statistical shape modeling of the left ventricle in cardiac MR images. Int J Comput Assist Radiol Surg 8(3):335–351. doi:10.1007/s11548-012-0787-1
Qin X, Tian Y, Yan P (2015) Feature competition and partial sparse shape modeling for cardiac image sequences segmentation. Neurocomputing 149:904–913. doi:10.1016/j.neucom.2014.07.044
Stebbing RV, Namburete AIL, Upton R, Leeson P, Noble JA (2015) Data-driven shape parameterization for segmentation of the right ventricle from 3D + t echocardiography. Med Image Anal 21(1):29–39. doi:10.1016/j.media.2014.12.002
Warfield SK, Zou KH, Wells WM (2004) Simultaneous truth and performance level estimation (STAPLE). IEEE Trans Med Imaging 23(7):903–921. doi:10.1109/TMI.2004.828354
Lebenberg J, Lalande A, Clarysse P, Buvat I, Casta C, Cochet A, Constantinidès C, Cousty J, Cesare A, Jehan-Besson S, Lefort M, Najman L, Roullot E, Sarry L, Tilmant C, Frouin F, Garreau M (2015) Improved estimation of cardiac function parameters using a combination of independent automated segmentation results in cardiovascular magnetic resonance imaging. PLoS One 10(8):e0135715-1–e0135715-16. doi:10.1371/journal.pone.0135715
Suinesiaputra A, Cowan BR, Finn JP, Fonseca CG, Kadish AH, Lee DC, Medrano-Gracia P, Warfield SK, Tao W, Young A (2012) Left ventricular segmentation challenge from cardiac MRI: a collation study. Lect Notes Comput Sci (including Subser Lect Notes Artif Intell Lect Notes Bioinformatics) 7085 LNCS:88–97. doi:10.1007/978-3-642-28326-0_9
Radau P, Lu Y, Connelly K, Paul G, Dick A.J. WGA (2009) Evaluation framework for algorithms segmenting short axis cardiac MRI. MIDAS Journal-Card MR Left Vent Segmentation Chall 2009. http://hdl.handle.net/10380/3070
Otsu N (1979) A threshold selection method from gray-level histograms. IEEE Trans Syst Man Cybern 9(1):62–66. doi:10.1109/TSMC.1979.4310076
Lu Y, Radau P, Connelly K, Dick A, Wright G (2009) Segmentation of left ventricle in cardiac cine mri: An automatic image-driven method. In: Functional imaging and modeling of the heart. Springer, Berlin, Heidelberg, pp 339–347. doi:10.1007/978-3-642-01932-6_37
Chen Y, Park SK, Ma Y, Ala R (2011) A new automatic parameter setting method of a simplified PCNN for image segmentation. IEEE Trans Neural Netw 22(6):880–892. doi:10.1109/TNN.2011.2128880
Chou N, Wu J, Bai Bingren J, Qiu A, Chuang K-H (2011) Robust automatic rodent brain extraction using 3D pulse-coupled neural networks (PCNN). IEEE Trans Image Process 20(9):2554–2564. doi:10.1109/TIP.2011.2126587
Gao C, Zhou D, Guo Y (2014) An iterative thresholding segmentation model using a modified pulse coupled neural network. Neural Process Lett 39(1):81–95. doi:10.1007/s11063-013-9291-z
Wei S, Hong Q, Hou M (2011) Automatic image segmentation based on PCNN with adaptive threshold time constant. Neurocomputing 74(9):1485–1491. doi:10.1016/j.neucom.2011.01.005
Ma YD, Liu L, Zhan K, Wu YQ (2010) Pulse-coupled neural networks and one-class support vector machines for geometry invariant texture retrieval. Image Vis Comput 28(11):1524–1529. doi:10.1016/j.imavis.2010.03.006
Stewart RD, Fermin I, Opper M (2002) Region growing with pulse-coupled neural networks: an alternative to seeded region growing. IEEE Trans Neural Netw 13(6):1557–1562. doi:10.1109/TNN.2002.804229
Xu C, Prince JL (1998) Snakes, shapes, and gradient vector flow. IEEE Trans Image Process 7(3):359–369. doi:10.1109/83.661186
Cohen LD (1991) On active contour models and balloons. CVGIP Image Underst 53(2):211–218. doi:10.1016/1049-9660(91)90028-N
Lalande A, Salve N, Comte A, Jaulent MC, Legrand L, Walker P, Cottin Y, Wolf J, Brunotte F (2004) Left ventricular ejection fraction calculation from automatically selected and processed diastolic and systolic frames in short-axis cine-MRI. J Cardiovasc Magn Reson 6(4):817–827. doi:10.1081/JCMR-200036143
Constantinides C, Chenoune Y, Kachenoura N, Roullot E, Mousseaux E, Herment A, Frouin F (2009) Semi-automated cardiac segmentation on cine magnetic resonance images using GVF-Snake deformable models. MIDAS J-Card MR Left Vent Segmentation Chall. http://hdl.handle.net/10380/3108
Cousty J, Najman L, Couprie M, Clément-Guinaudeau S, Goissen T, Garot J (2010) Segmentation of 4D cardiac MRI: automated method based on spatio-temporal watershed cuts. Image Vis Comput 28(8):1229–1243. doi:10.1016/j.imavis.2010.01.001
Schaerer J, Casta C, Pousin J, Clarysse P (2010) A dynamic elastic model for segmentation and tracking of the heart in MR image sequences. Med Image Anal 14(6):738–749. doi:10.1016/j.media.2010.05.009
Fonseca CG, Backhaus M, Bluemke DA, Britten RD, Do Chung J, Cowan BR, Dinov ID, Finn JP, Hunter PJ, Kadish AH, Lee DC, Lima JC, Medrano-Gracia P, Shivkumar K, Suinesiaputra A, Tao W, Young A (2011) The cardiac atlas project-an imaging database for computational modeling and statistical atlases of the heart. Bioinformatics 27(16):2288–2295. doi:10.1093/bioinformatics/btr360
Suinesiaputra A, Medrano-Gracia P, Cowan BR, Young A (2015) Big heart data: advancing health informatics through data sharing in cardiovascular imaging. IEEE J Biomed Health Inform 19(4):1283–1290. doi:10.1109/JBHI.2014.2370952
Lebenberg J, Buvat I, Lalande A, Clarysse P, Casta C, Cochet A, Constantinidès C, Cousty J, Cesare A, Jehan-Besson S, Lefort M, Najman L, Roullot E, Sarry L, Tilmant C, Garreau MG, Frouin F (2012) Nonsupervised ranking of different segmentation approaches: Application to the estimation of the left ventricular ejection fraction from cardiac cine MRI sequences. IEEE Trans Med Imaging 31(8):1651–1660. doi:10.1109/TMI.2012.2201737
Acknowledgments
The authors thank the editor and the reviewers for their comments that have helped improve this paper. We also thank Sunnybrook Health Sciences Centre for providing the clinical image data, ground truths and evaluation software for us. We acknowledge the whole French MEDIEVAL group for allowing the comparisons with their segmentation results (which are freely available on https://github.com/frederiquefrouin/Medieval). Besides, we used the data made available through the Cardiac Atlas Project. This work was supported in part by National Natural Science Foundation of China (No. 61175012), Natural Science Foundation of Gansu Province (No. 1208RJZA265) and Fundamental Research Funds for the Central Universities (Nos. lzujbky-2015-197 & lzujbky-2015-196).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
None.
Appendix
Appendix
As shown in Fig. 9, the single neuron structure of SPCNN model includes three parts: input field, modulation field and pulse generator field [28]. And, the discrete SPCNN was described as follows:
In the input field, each neuron \(N_{xy}\) in the position (x, y) receives information from both the feeding input \(F_{xy}[n]\) and the linking input \(L_{xy}[n]\) in iteration n. \(F_{xy}[n]\) is equal to the normalized gray intensity of the original input image \(S_{xy}\). \(L_{xy}[n]\) is the result of that \(N_{xy}\) communicates with its eight neighboring neurons \(N_{kl}\) through a constant synaptic weights W. \(Y_{kl}[n-1]\) is the output of \(N_{kl}\) in the previous iteration. In the modulation field, \(F_{xy}[n]\) and \(L_{xy}[n]\) are modulated through the linking strength \(\beta \) to obtain the internal activity \(U_{xy}[n]\) which is used to judge whether the neuron \(N_{xy}\) fires or not. In the last (pulse generator) field, \(U_{xy}[n]\) is compared with the dynamic threshold of previous iteration \(E_{xy}[n-1]\). If \(U_{xy}[n]>E_{xy}[n-1]\), \(N_{xy}\) fires (i.e., setting the output \(Y_{xy}[n]\) to one); otherwise \(N_{xy}\) does not fire (i.e., setting \(Y_{xy}[n]\) to zero). In this way, the output matrix Y[n], which consists of 0 and 1, generates a binary image in iteration n. The dynamic threshold \(E_{xy}[n]\) is updated in each iteration according to the output \(Y_{xy}[n-1]\): If \(N_{xy}\) fires in the previous iteration \((Y_{xy}[n-1]=1),E_{xy}[n]\) will increase by parameter \(V_{E;}\) if \(N_{xy}\) does not fire (\(Y_{xy}[n-1]=0\)), \(\hbox {E}_{xy}[n]\) will decay by an exponential decay coefficient \(e^{-{\alpha }_e}\) [25].
Single neuron structure of SPCNN model [28]
Rights and permissions
About this article
Cite this article
Ma, Y., Wang, L., Ma, Y. et al. An SPCNN-GVF-based approach for the automatic segmentation of left ventricle in cardiac cine MR images. Int J CARS 11, 1951–1964 (2016). https://doi.org/10.1007/s11548-016-1429-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-016-1429-9