Abstract
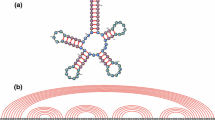
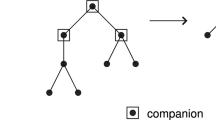
We give a Large Deviation Principle (LDP) with explicit rate function for the distribution of vertex degrees in plane trees, a combinatorial model of RNA secondary structures. We calculate the typical degree distributions based on nearest neighbor free energies, and compare our results with the branching configurations found in two sets of large RNA secondary structures. We find substantial agreement overall, with some interesting deviations which merit further study.
Similar content being viewed by others
References
Bakhtin, Y., Heitsch, C.E., 2008. Large deviations for random trees. J. Stat. Phys. 132(3), 551–560.
Cannone, J.J., Subramanian, S., Schnare, M.N., Collett, J.R., D’Souza, L.M., Du, Y., Feng, B., Lin, N., Madabusi, L.V., Müller, K.M., Pande, N., Shang, Z., Yu, N., Gutell, R.R., 2002. The Comparative RNA Web (CRW) Site: an online database of comparative sequence and structure information for ribosomal, intron, and other RNAs. BMC Bioinform. 3(1).
Clote, P., Gasieniec, L., Kolpakov, R., Kranakis, E., Krizanc, D., 2005. On realizing shapes in the theory of RNA neutral networks. J. Theor. Biol. 236(2), 216–227.
Clote, P., Kranakis, E., Krizanc, D., Stacho, L., 2007. Asymptotic expected number of base pairs in optimal secondary structure for random RNA using the Nussinov–Jacobson energy model. Discrete Appl. Math. 155(6–7), 759–787.
Dembo, A., Zeitouni, O., 1998. Large deviations techniques and applications, volume 38 of Applications of Mathematics, 2nd edn. Springer, New York.
Doshi, K.J., Cannone, J.J., Cobaugh, C.W., Gutell, R.R., 2004 Evaluation of the suitability of free-energy minimization using nearest-neighbor energy parameters for RNA secondary structure prediction. BMC Bioinform. 5(105).
Ellis, R.S., 2006. Entropy, large deviations, and statistical mechanics. In: Classics in Mathematics. Springer, Berlin. Reprint of the 1985 original.
Fields, D.S., Gutell, R.R., 1996. An analysis of large rRNA sequences folded by a thermodynamic method. Fold Des. 1(6), 419–430.
Fontana, W., Konings, D., Stadler, P.F., Schuster, P., 1993. Statistics of RNA secondary structures. Biopolymers 33(9), 1389–1404.
Heitsch, C.E., 2008a. Combinatorial insights into RNA secondary structures. In preparation.
Heitsch, C.E., 2008b. A new metric on plane trees and RNA configurations. In revision.
Hofacker, I.L., Schuster, P., Stadler, P.F., 1998. Combinatorics of RNA secondary structures. Discrete Appl. Math. 88(1–3), 207–237.
Konings, D.A.M., Gutell, R.R., 1995. A comparison of thermodynamic foldings with comparatively derived structures of 16S and 16S-like rRNAs. RNA 1(6), 559–574.
Lu, Z.J., Turner, D.H., Mathews, D.H., 2006. A set of nearest neighbor parameters for predicting the enthalpy change of RNA secondary structure formation. Nucleic Acids Res 34(17), 4912–4924.
Mathews, D.H., Sabina, J., Zuker, M., Turner, D.H., 1999. Expanded sequence dependence of thermodynamic parameters improves prediction of RNA secondary structure. J. Mol. Biol. 288(5), 911–940.
Miklós, I., Meyer, I.M., Nagy, B., 2005. Moments of the Boltzmann distribution for RNA secondary structures. Bull. Math. Biol. 67(5), 1031–1047.
Nebel, M.E., 2004a. Identifying good predictions of RNA secondary structure. In Pac. Symp. Biocomput., 423–434.
Nebel, M.E., 2004b. Investigation of the Bernoulli model for RNA secondary structures. Bull. Math. Biol. 66(5), 925–964.
Palmenberg, A.C., Sgro, J.-Y., 1997. Topological organization of picornaviral genomes: Statistical prediction of RNA structural signals. S. Virol. 8, 231–241.
Rivas, E., Eddy, S.R., 2000. Secondary structure alone is generally not statistically significant for the detection of noncoding RNAs. Bioinformatics 16(7), 583–605.
Stanley, R.P., 1999. Enumerative Combinatorics. Vol. 2, Vol. 62. Cambridge Studies in Advanced Mathematics, Cambridge University Press, Cambridge.
Tinoco, I. Jr., Bustamante, C., 1999. How RNA folds. J. Mol. Biol. 293(2), 271–281.
Workman, C., Krogh, A., 1999. No evidence that mRNAs have lower folding free energies than random sequences with the same dinucleotide distribution. Nucleic Acids Res. 27(24), 4816–4822.
Wuchty, S., Fontana, W., Hofacker, I.L., Schuster, P., 1999. Complete suboptimal folding of RNA and the stability of secondary structures. Biopolymers 49(2), 145–165.
Zuker, M., 2003. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31(13), 3406–3415.
Zuker, M., Jacobson, A.B., 1995. “Well-determined” regions in RNA secondary structure prediction: analysis of small subunit ribosomal RNA. Nucleic Acids Res. 23(14), 2791–2798.
Zuker, M., Jaeger, J.A., Turner, D.H., 1991. A comparison of optimal and suboptimal RNA secondary structures predicted by free energy minimization with structures determined by phylogenetic comparison. Nucleic Acids Res. 19(10), 2707–2714.
Zuker, M., Mathews, D., Turner, D., 1999. Algorithms and thermodynamics for RNA secondary structure prediction: A practical guide. In: Barciszewski, J., Clark, B. (Eds.), RNA Biochemistry and Biotechnology, NATO ASI Series, pp. 11–43. Kluwer Academic, Dordrecht
Author information
Authors and Affiliations
Corresponding author
Additional information
Y. Bakhtin partially supported by NSF CAREER DMS-0742424. C.E. Heitsch partially supported by a BWF CASI and NIH NIGMS R01 GM083621.
Rights and permissions
About this article
Cite this article
Bakhtin, Y., Heitsch, C.E. Large Deviations for Random Trees and the Branching of RNA Secondary Structures. Bull. Math. Biol. 71, 84–106 (2009). https://doi.org/10.1007/s11538-008-9353-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-008-9353-y