Abstract
We propose a multiscale texture features based on Laplacian-of Gaussian (LoG) filter to predict progression free (PFS) and overall survival (OS) in patients newly diagnosed with glioblastoma (GBM). Experiments use the extracted features derived from 40 patients of GBM with T1-weighted imaging (T1-WI) and Fluid-attenuated inversion recovery (FLAIR) images that were segmented manually into areas of active tumor, necrosis, and edema. Multiscale texture features were extracted locally from each of these areas of interest using a LoG filter and the relation between features to OS and PFS was investigated using univariate (i.e., Spearman’s rank correlation coefficient, log-rank test and Kaplan-Meier estimator) and multivariate analyses (i.e., Random Forest classifier). Three and seven features were statistically correlated with PFS and OS, respectively, with absolute correlation values between 0.32 and 0.36 and p < 0.05. Three features derived from active tumor regions only were associated with OS (p < 0.05) with hazard ratios (HR) of 2.9, 3, and 3.24, respectively. Combined features showed an AUC value of 85.37 and 85.54% for predicting the PFS and OS of GBM patients, respectively, using the random forest (RF) classifier. We presented a multiscale texture features to characterize the GBM regions and predict he PFS and OS. The efficiency achievable suggests that this technique can be developed into a GBM MR analysis system suitable for clinical use after a thorough validation involving more patients.

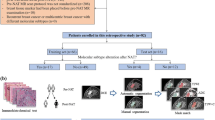
Scheme of the proposed model for characterizing the heterogeneity of GBM regions and predicting the overall survival and progression free survival of GBM patients. (1) Acquisition of pretreatment MRI images; (2) Affine registration of T1-WI image with its corresponding FLAIR images, and GBM subtype (phenotypes) labelling; (3) Extraction of nine texture features from the three texture scales fine, medium, and coarse derived from each of GBM regions; (4) Comparing heterogeneity between GBM regions by ANOVA test; Survival analysis using Univariate (Spearman rank correlation between features and survival (i.e., PFS and OS) based on each of the GBM regions, Kaplan-Meier estimator and log-rank test to predict the PFS and OS of patient groups that grouped based on median of feature), and multivariate (random forest model) for predicting the PFS and OS of patients groups that grouped based on median of PFS and OS.





Similar content being viewed by others
References
Stupp R, Hegi ME, van den Bent MJ, Mason WP, Weller M, Mirimanoff RO, Cairncross JG (2006) Changing paradigms—an update on the multidisciplinary management of malignant glioma. Oncologist 11:165–180
Zacharaki EI, Wang S, Chawla S, Soo Yoo D, Wolf R, Melhem ER, Davatzikos C (2009) Classification of brain tumor type and grade using MRI texture and shape in a machine learning scheme. Magn Reson Med 62:1609–1618
Asai A, Matsutani M, Kohno T, Nakamura O, Tanaka H, Fujimaki T, Funada N, Matsuda T, Nagata K, Takakura K (1989) Subacute brain atrophy after radiation therapy for malignant brain tumor. Cancer 63:1962–1974
Norred SE, Johnson JA (2014) Magnetic resonance-guided laser induced thermal therapy for glioblastoma multiforme: a review. Biomed Res Int 2014:761312
Patriarche JW, Erickson BJ (2007) Part 1. Automated change detection and characterization in serial MR studies of brain-tumor patients. J Digit Imaging 20:203–222
Sachdeva J, Kumar V, Gupta I, Khandelwal N, Ahuja CK (2013) Segmentation, feature extraction, and multiclass brain tumor classification. J Digit Imaging 26:1141–1150
Brown RA, Frayne R (2008) A comparison of texture quantification techniques based on the Fourier and S transforms. Med Phys 35:4998–5008
Downey K, Riches SF, Morgan VA, Giles SL, Attygalle AD, Ind TE, Barton DPJ, Shepherd JH, deSouza NM (2013) Relationship between imaging biomarkers of stage I cervical cancer and poor-prognosis histologic features: quantitative histogram analysis of diffusion-weighted MR images. AJR Am J Roentgenol 200:314–320
Holli KK, Harrison L, Dastidar P, Wäljas M, Liimatainen S, Luukkaala T, Öhman J, Soimakallio S, Eskola H (2010) Texture analysis of MR images of patients with mild traumatic brain injury. BMC Med Imaging 10:8
Schwarzmaier H-J, Eickmeyer F, von Tempelhoff W, Fiedler VU, Niehoff H, Ulrich SD, Ulrich F (2005) MR-guided laser irradiation of recurrent glioblastomas. J Magn Reson Imaging 22:799–803
Teruel JR, Heldahl MG, Goa PE, Pickles M, Lundgren S, Bathen TF, Gibbs P (2014) Dynamic contrast-enhanced MRI texture analysis for pretreatment prediction of clinical and pathological response to neoadjuvant chemotherapy in patients with locally advanced breast cancer. NMR Biomed 27:887–896
Chaddad A, Colen RR (2014) Statistical feature selection for enhanced detection of brain tumor. SPIE optical engineering+ applications. International Society for Optics and Photonics, Bellingham, pp 92170V–92170V
Chaddad A, Desrosiers C, Toews M (2016) Radiomic analysis of multi-contrast brain MRI for the prediction of survival in patients with glioblastoma multiforme. 2016 38th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC). pp 4035–4038
Eliat P-A, Olivié D, Saïkali S, Carsin B, Saint-Jalmes H, de Certaines JD (2012) Can dynamic contrast-enhanced magnetic resonance imaging combined with texture analysis differentiate malignant glioneuronal tumors from other glioblastoma? Neurol Res Int 2012:195176
Lee J, Jain R, Khalil K, Griffith B, Bosca R, Rao G, Rao A (2016) Texture feature ratios from relative CBV maps of perfusion MRI are associated with patient survival in glioblastoma. AJNR Am J Neuroradiol 37:37–43
Skogen K, Ganeshan B, Good T, Critchley G, Miles KA (2011) Imaging heterogeneity in gliomas using texture analysis. Cancer Imaging 11 Spe:S113. https://doi.org/10.1102/1470-7330.2011.9057
Kickingereder P, Götz M, Muschelli J, Wick A, Neuberger U, Shinohara RT, Sill M, Nowosielski M, Schlemmer H-P, Radbruch A (2016) Large-scale radiomic profiling of recurrent glioblastoma identifies an imaging predictor for stratifying anti-angiogenic treatment response. Clin Cancer Res 22:5765–5771
Clark K, Vendt B, Smith K, Freymann J, Kirby J, Koppel P, Moore S, Phillips S, Maffitt D, Pringle M, Tarbox L, Prior F (2013) The cancer imaging archive (TCIA): maintaining and operating a public information repository. J Digit Imaging 26:1045–1057
Gonzalez RC, Woods RE (2002) Digital Image Processing, 2nd edition. Prentice Hall, Upper Saddle River, N.J
Ganeshan B, Abaleke S, Young RCD, Chatwin CR, Miles KA (2010) Texture analysis of non-small cell lung cancer on unenhanced computed tomography: initial evidence for a relationship with tumour glucose metabolism and stage. Cancer Imaging 10:137–143
Chaddad A, Desrosiers C, Toews M (2017) Multi-scale radiomic analysis of sub-cortical regions in MRI related to autism, gender and age. Sci Rep 7:45639
Chaddad A, Desrosiers C, Bouridane A, Toews M, Hassan L, Tanougast C (2016) Multi texture analysis of colorectal cancer continuum using multispectral imagery. PLoS One 11:e0149893
Skogen K, Schulz A, Dormagen JB, Ganeshan B, Helseth E, Server A (2016) Diagnostic performance of texture analysis on MRI in grading cerebral gliomas. Eur J Radiol 85:824–829
Chen S, Yao L, Chen B (2016) A parameterized logarithmic image processing method with Laplacian of Gaussian filtering for lung nodule enhancement in chest radiographs. Med Biol Eng Comput 54:1793–1806
Cuevas A, Febrero M, Fraiman R (2004) An anova test for functional data. Comput Stat Data Anal 47:111–122
Holm S (1979) A simple sequentially rejective multiple test procedure. Scand J Stat 6:65–70
Zar JH (1972) Significance testing of the Spearman rank correlation coefficient. J Am Stat Assoc 67:578–580
Leung K-M, Elashoff RM, Afifi AA (1997) Censoring issues in survival analysis. Annu Rev Public Health 18:83–104
Kleinbaum DG, Klein M (2012) Kaplan-Meier survival curves and the log-rank test. Survival analysis. Springer, New York, pp 55–96
Strobl C, Malley J, Tutz G (2009) An introduction to recursive partitioning: rationale, application, and characteristics of classification and regression trees, bagging, and random forests. Psychol Methods 14:323
Hanley JA, McNeil BJ (1982) The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology 143:29–36
Lambin P, Leijenaar RTH, Deist TM, Peerlings J, de Jong EEC, van Timmeren J, Sanduleanu S, Larue RTHM, Even AJG, Jochems A, van Wijk Y, Woodruff H, van Soest J, Lustberg T, Roelofs E, van Elmpt W, Dekker A, Mottaghy FM, Wildberger JE, Walsh S (2017) Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol 14:749–762
Meldolesi E, Dinapoli N, Gatta R, Damiani A, Valentini V, Farchione A (2018) How Can Radiomics Improve Clinical Choices? Multidisciplinary Management of Rectal Cancer. Springer, Cham, pp 135–149
Lao J, Chen Y, Li Z-C, Li Q, Zhang J, Liu J, Zhai G (2017) A deep learning-based radiomics model for prediction of survival in glioblastoma multiforme. Sci Rep 7:10353
Chaddad A, Tanougast C (2016) Extracted magnetic resonance texture features discriminate between phenotypes and are associated with overall survival in glioblastoma multiforme patients. Med Biol Eng Comput 54:1707–1718
Herlidou-Même S, Constans JM, Carsin B, Olivie D, Eliat PA, Nadal-Desbarats L, Gondry C, Le Rumeur E, Idy-Peretti I, de Certaines JD (2003) MRI texture analysis on texture test objects, normal brain and intracranial tumors. Magn Reson Imaging 21:989–993
Inda M-M, Bonavia R, Seoane J (2014) Glioblastoma Multiforme: a look inside its heterogeneous nature. Cancers (Basel) 6:226–239
Chaddad A, Desrosiers C, Toews M (2016) Phenotypic characterization of glioblastoma identified through shape descriptors. International Society for Optics and Photonics, Bellingham, pp 97852M–97852M
Sottoriva A, Spiteri I, Piccirillo SGM, Touloumis A, Collins VP, Marioni JC, Curtis C, Watts C, Tavaré S (2013) Intratumor heterogeneity in human glioblastoma reflects cancer evolutionary dynamics. Proc Natl Acad Sci U S A 110:4009–4014
Chaddad A, Desrosiers C, Toews M (2016) GBM heterogeneity characterization by radiomic analysis of phenotype anatomical planes. Medical imaging 2016: image processing. International Society for Optics and Photonics, Bellingham, p 978424
Liu Y, Xu X, Yin L, Zhang X, Li L, Lu H (2017) Relationship between glioblastoma heterogeneity and survival time: an MR imaging texture analysis. Am J Neuroradiol 38:1695–1701
Liu Y, Zhang X, Feng N, Yin L, He Y, Xu X, Lu H (2018) The effect of glioblastoma heterogeneity on survival stratification: a multimodal MR imaging texture analysis. Acta Radiol 284185118756951
Rathore S, Akbari H, Rozycki M, Abdullah KG, Nasrallah MP, Binder ZA, Davuluri RV, Lustig RA, Dahmane N, Bilello M, O’Rourke DM, Davatzikos C (2018) Radiomic MRI signature reveals three distinct subtypes of glioblastoma with different clinical and molecular characteristics, offering prognostic value beyond IDH1. Sci Rep 8:5087
Mazurowski MA, Zhang J, Peters KB, Hobbs H (2014) Computer-extracted MR imaging features are associated with survival in glioblastoma patients. J Neuro-Oncol 120:483–488
Macyszyn L, Akbari H, Pisapia JM, Da X, Attiah M, Pigrish V, Bi Y, Pal S, Davuluri RV, Roccograndi L, Dahmane N, Martinez-Lage M, Biros G, Wolf RL, Bilello M, O’Rourke DM, Davatzikos C (2016) Imaging patterns predict patient survival and molecular subtype in glioblastoma via machine learning techniques. Neuro-Oncology 18:417–425
Zhou M, Hall L, Goldgof D, Russo R, Balagurunathan Y, Gillies R, Gatenby R (2014) Radiologically defined ecological dynamics and clinical outcomes in glioblastoma multiforme: preliminary results. Transl Oncol 7:5–13
Cui Y, Tha KK, Terasaka S, Yamaguchi S, Wang J, Kudo K, Xing L, Shirato H, Li R (2016) Prognostic imaging biomarkers in glioblastoma: development and independent validation on the basis of multiregion and quantitative analysis of MR images. Radiology 278:546–553
Chaddad A, Daniel P, Desrosiers C, Toews M, Abdulkarim B (2018) Novel radiomic features based on joint intensity matrices for predicting glioblastoma patient survival time. IEEE Journal of Biomedical and Health Informatics 1–1
Egger J, Kapur T, Fedorov A, Pieper S, Miller JV, Veeraraghavan H, Freisleben B, Golby AJ, Nimsky C, Kikinis R (2013) GBM volumetry using the 3D slicer medical image computing platform. Sci Rep 3:1364
Lambin P, Rios-Velazquez E, Leijenaar R, Carvalho S, van Stiphout RGPM, Granton P, Zegers CML, Gillies R, Boellard R, Dekker A, Aerts HJWL (2012) Radiomics: extracting more information from medical images using advanced feature analysis. Eur J Cancer 48:441–446
Ahmad Chaddad, Christian Desrosiers, Matthew Toews (2016) Radiomic analysis of multi-contrast brain MRI for the prediction of survival in patients with glioblastoma multiforme. 38th Annual International Conference of the IEEE Engineering in Medicine and Biology Society USA, p 4035:4038
Kickingereder P, Burth S, Wick A, Götz M, Eidel O, Schlemmer H-P, Maier-Hein KH, Wick W, Bendszus M, Radbruch A, Bonekamp D (2016) Radiomic profiling of glioblastoma: identifying an imaging predictor of patient survival with improved performance over established clinical and radiologic risk models. Radiology 280:880–889
Zhou M, Chaudhury B, Hall LO, Goldgof DB, Gillies RJ, Gatenby RA (2017) Identifying spatial imaging biomarkers of glioblastoma multiforme for survival group prediction. J Magn Reson Imaging 46:115–123
Ingrisch M, Schneider MJ, Nörenberg D, Negrao de Figueiredo G, Maier-Hein K, Suchorska B, Schüller U, Albert N, Brückmann H, Reiser M, Tonn J-C, Ertl-Wagner B (2017) Radiomic analysis reveals prognostic information in T1-weighted baseline magnetic resonance imaging in patients with glioblastoma. Investig Radiol 52:360–366
Prasanna P, Patel J, Partovi S, Madabhushi A, Tiwari P (2016) Radiomic features from the peritumoral brain parenchyma on treatment-naïve multi-parametric MR imaging predict long versus short-term survival in glioblastoma multiforme: preliminary findings. Eur Radiol 1–10
Kickingereder P, Götz M, Muschelli J, Wick A, Neuberger U, Shinohara RT, Sill M, Nowosielski M, Schlemmer H-P, Radbruch A, Wick W, Bendszus M, Maier-Hein KH, Bonekamp D (2016) Large-scale radiomic profiling of recurrent glioblastoma identifies an imaging predictor for stratifying anti-angiogenic treatment response. Clin Cancer Res. https://doi.org/10.1158/1078-0432.CCR-16-0702
Liu S, Wang Y, Xu K, Wang Z, Fan X, Zhang C, Li S, Qiu X, Jiang T (2017) Relationship between necrotic patterns in glioblastoma and patient survival: fractal dimension and lacunarity analyses using magnetic resonance imaging. Sci Rep 7:8302
Hu LS, Ning S, Eschbacher JM, Baxter LC, Gaw N, Ranjbar S, Plasencia J, Dueck AC, Peng S, Smith KA, Nakaji P, Karis JP, Quarles CC, Wu T, Loftus JC, Jenkins RB, Sicotte H, Kollmeyer TM, O’Neill BP, Elmquist W, Hoxworth JM, Frakes D, Sarkaria J, Swanson KR, Tran NL, Li J, Mitchell JR (2017) Radiogenomics to characterize regional genetic heterogeneity in glioblastoma. Neuro-Oncology 19:128–137
Zhang B, Chang K, Ramkissoon S, Tanguturi S, Bi WL, Reardon DA, Ligon KL, Alexander BM, Wen PY, Huang RY (2017) Multimodal MRI features predict isocitrate dehydrogenase genotype in high-grade gliomas. Neuro-Oncology 19:109–117
Sala E, Mema E, Himoto Y, Veeraraghavan H, Brenton JD, Snyder A, Weigelt B, Vargas HA (2017) Unravelling tumour heterogeneity using next-generation imaging: radiomics, radiogenomics, and habitat imaging. Clin Radiol 72:3–10
Cui Y, Ren S, Tha KK, Wu J, Shirato H, Li R (2017) Volume of high-risk intratumoral subregions at multi-parametric MR imaging predicts overall survival and complements molecular analysis of glioblastoma. Eur Radiol 27:3583–3592
Itakura H, Achrol AS, Mitchell LA, Loya JJ, Liu T, Westbroek EM, Feroze AH, Rodriguez S, Echegaray S, Azad TD, Yeom KW, Napel S, Rubin DL, Chang SD, Harsh GR, Gevaert O (2015) Magnetic resonance image features identify glioblastoma phenotypic subtypes with distinct molecular pathway activities. Sci Transl Med 7:303ra138
Funding
The study was supported by grant from Varian Medical System.
Author information
Authors and Affiliations
Contributions
A.C. designed the study, analyzed the data and wrote the paper. A.C., S.S., T.N., and B.A., reviewed and gave final approval for publication.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
For this type of study, formal consent is not required.
Electronic supplementary material
Table S1
(DOC 28 kb)
Rights and permissions
About this article
Cite this article
Chaddad, A., Sabri, S., Niazi, T. et al. Prediction of survival with multi-scale radiomic analysis in glioblastoma patients. Med Biol Eng Comput 56, 2287–2300 (2018). https://doi.org/10.1007/s11517-018-1858-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-018-1858-4