Abstract
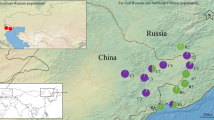
Ulva pertusa is a native species to Asia along the western coast of Pacific Ocean, with new occurrence records in the eastern coast of Pacific, the northwest coast of Atlantic and the Mediterranean Sea. However, little is known about its population genetic structure. In this study, twelve U. pertusa populations from 3 coastal areas of China: Qingdao, Yantai and Dalian, were applied to ISSR analysis. The selected 4 ISSR primers amplified 120 polymorphic bands totally. Nei’s gene diversity (H) ranged from 0.0729 to 0.1496, and Shannon’s information index (I) ranged from 0.1072 to 0.2196. Genetic diversity was greater within Qingdao populations (H = 0.2069, I = 0.3232). Analysis of molecular variance (AMOVA) showed the greatest variance within populations (68.57%), much less variance among populations (22.63%) and among areas (8.79%). Unweighted pair-group mean analysis (UPGMA) indicated that clustering of U. pertusa individuals mainly relates to their populations and geographic distances separating those populations. Genetic differentiation and limited gene flow among U. pertusa populations were indicated by ISSR analysis.
Similar content being viewed by others
References
Tseng, C K, Zhang D R, Zhang J F, et al. Manual of Chinese economic seaweeds (in Chinese). Beijing: Science Press, 1962. 43–50
Aguilar-Rosas R, Aguilar-Rosas L E, Shimada S. First record of Ulva pertusa Kjellman (Ulvales, Chlorophyta) in the Pacific coast of Mexico. Algae, 2008, 23: 201–207
López S B, Fernández I B, Lozano R B, et al. Is the cryptic alien seaweed Ulva pertusa (Ulvales, Chlorophyta) widely distributed along European Atlantic coasts? Bot Mar, 2007, 50: 267–274
Verlaque M, Belsher T, Deslous-Paoli J M. Morphology and reproduction of Asiatic Ulva pertusa (Ulvales, Chlorophyta) in Thau Lagoon (France, Mediterranean Sea). Cryptogamie Algol, 2002, 23: 301–310
Su X R, Li T W, Chang S J. A study on nutrition of Ulva pertusa Kjellm (in Chinese). Chinese J Mar Drugs, 1997, 61: 33–35
Yu P Z, Li N, Liu X G, et al. Antihyperlipidemic effects of different molecular weights sulfated-polysaccharides from Ulva pertusa. Pharmacol Res, 2003, 48: 543–549
Shimada S, Hiraoka M, Nabataet S, et al. Molecular phylogenetic analyses of the Japanese Ulva and Enteromorpha (Ulvales, Ulvophyceae), with special reference to the free-floating Ulva. Phycol Res, 2003, 51: 99–108
Slatkin M. Gene flow and the geographic structure of populations. Science, 1987, 236: 787–792
Ji J M, Ma J H, Li Y H, et al. Isomorphic alternation of generations of Ulva pertusa (in Chinese). Fishery Modernization, 2008, 35: 32–35
Hiraoka, M, Shimada S, Ohno M, et al. Asexual life history by quadriflagellate swarmers of Ulva spinulosa (Ulvales, Ulvophyceae). Phycol Res, 2003, 51: 29–34
Callow M E, Callow J A, et al. Primary adhesion of Enteromorpha (Chlorophyta, Ulvales) propagules: Quantitative settlement studies and video microscopy. J Phycol, 1997, 33: 938–947
Innes D J. Genetic structure of asexually reproducing Enteromorpha linza (Ulvales, Chlorophyta) in Long Island Sound. Mar Biol, 1987, 94: 459–467
Innes D J. Genetic differentiation in the intertidal zone in populations of the alga Enteromorpha Linza (Ulvales, Chlorophyta). Mar Biol, 1988, 97: 9–16
Wang X L, Zhao F J, Hu Z M, et al. Inter-simple sequence repeat (ISSR) analysis of genetic variation of Chondrus crispus populations from North Atlantic. Aquat Bot, 2008, 88: 154–159
Leskinen E, Pamilo P. Evolution of the ITS sequences of ribosomal DNA in Enteromorpha (Chlorophyceae). Hereditas, 1997, 126: 17–23
Ballard H E, Sytsma K J, Kowal R R. Shrinking the violets: Phylogenetic relationships of infrageneric groups in Viola (Violaceae) based on internal transcribed spacer DNA sequences. Syst Bot, 1998, 23: 439–458
Drets M E. BANDSCAN-a computer program for on-line linear scanning of human banded chromosomes. Comput Programs Biomed, 1978, 8: 283–294
Wolfe A D, Liston A. Contributions of PCR-based methods to plant systematics and evolutionary biology. In: Soltis D E, Soltis P S, Doyle J J, eds. Plant Molecular Systematics. Boston: Kluwer Academic Press, 1998. 43–86
Yeh F C, Yang R C, Boyle T. Popgene, version 1.32. Microsoft Window-based freeware for population genetic analysis. University of Alberta and Center for International Forestry Research, 1999
Nei M. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics, 1978, 89: 583–590
Miller M P. AMOVA-PREP 1.01: A program for the preparation of AMOVA input files from dominant-markers raw data. Computer software distributed by author. 1998
Huff D R, Peakall R, Smouse P E. RAPD variation within and among natural-populations of outcrossing buffalograss [Buchloë dactyloides (Nutt.) Engelm.]. Theor Appl Genet, 1993, 86: 927–934
Excoffier L, Smouse P E, Quattro J M. Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics, 1992, 131: 479–491
Meekins J F, Ballard H E, McCarthy B C. Genetic variation and molecular biogeography of a North American invasive plant species (Alliaria petiolata, Brassicaceae). Int J Plant Sci, 2001, 162: 161–169
Rohlf F J. NTSYS-pc: numerical taxonomy and multivariate analysis system, version 2.1. New York: Exeter Software. 2000
Salimath S S, Deoliveira A C, Godwin I D, et al. Assessment of genome origins and genetic diversity in the genus Eleusine with DNA markers. Genome, 1995, 38: 757–763
Nei M, Li W H. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Nati Acad Sci USA, 1979, 76: 5269–5273
Rice W R. Analyzing tables of statistical tests. Evolution, 1989, 43: 223–225
Kostamo K, Blomster J, Korpelainen H, et al. New microsatellite markers for Ulva intestinalis (Chlorophyta) and the transferability of markers across species of Ulvaceae. Phycologia, 2008, 47: 580–587
Zhao F J, Liu F L, Liu J D, et al. Genetic structure analysis of natural Sargassum muticum (Fucales, Phaeophyta) populations using RAPD and ISSR markers. J Appl Phycol, 2008, 20: 191–198
Author information
Authors and Affiliations
Corresponding authors
About this article
Cite this article
Zhao, J., Jiang, P., Li, N. et al. Analysis of genetic variation within and among Ulva pertusa (Ulvaceae, Chlorophyta) populations using ISSR markers. Chin. Sci. Bull. 55, 705–711 (2010). https://doi.org/10.1007/s11434-009-0715-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-009-0715-0