Abstract
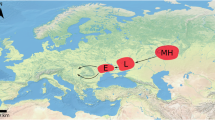
The Y chromosome plays key roles in male fertility and reflects the evolutionary history of paternal lineages. Here, we present a de novo genome assembly of the Hu sheep with the first draft assembly of ovine Y chromosome (oMSY), using nanopore sequencing and Hi-C technologies. The oMSY that we generated spans 10.6 Mb from which 775 Y-SNPs were identified by applying a large panel of whole genome sequences from worldwide sheep and wild Iranian mouflons. Three major paternal lineages (HY1a, HY1b and HY2) were defined across domestic sheep, of which HY2 was newly detected. Surprisingly, HY2 forms a monophyletic clade with the Iranian mouflons and is highly divergent from both HY1a and HY1b. Demographic analysis of Y chromosomes, mitochondrial and nuclear genomes confirmed that HY2 and the maternal counterpart of lineage C represented a distinct wild mouflon population in Iran that diverge from the direct ancestor of domestic sheep, the wild mouflons in Southeastern Anatolia. Our results suggest that wild Iranian mouflons had introgressed into domestic sheep and thereby introduced this Iranian mouflon specific lineage carrying HY2 to both East Asian and Africa sheep populations.
Similar content being viewed by others
References
Alberto, F.J., Boyer, F., Orozco-terWengel, P., Streeter, I., Servin, B., de Villemereuil, P., Benjelloun, B., Librado, P., Biscarini, F., Colli, L., et al. (2018). Convergent genomic signatures of domestication in sheep and goats. Nat Commun 9, 813.
Álvarez, I., Capote, J., Traoré, A., Fonseca, N., Pérez, K., Cuervo, M., Fernández, I., and Goyache, F. (2013). Mitochondrial analysis sheds light on the origin of hair sheep. Anim Genet 44, 344–347.
Bellott, D.W., Hughes, J.F., Skaletsky, H., Brown, L.G., Pyntikova, T., Cho, T.J., Koutseva, N., Zaghlul, S., Graves, T., Rock, S., et al. (2014). Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature 508, 494–499.
Bickhart, D.M., Rosen, B.D., Koren, S., Sayre, B.L., Hastie, A.R., Chan, S., Lee, J., Lam, E.T., Liachko, I., Sullivan, S.T., et al. (2017). Single-molecule sequencing and chromatin conformation capture enable de novo reference assembly of the domestic goat genome. Nat Genet 49, 643–650.
Bidon, T., Schreck, N., Hailer, F., Nilsson, M.A., and Janke, A. (2015). Genome-wide search identifies 1.9 Mb from the polar bear Y chromosome for evolutionary analyses. Genome Biol Evol 7, 2010–2022.
Browning, B.L., and Browning, S.R. (2016). Genotype imputation with millions of reference samples. Am J Hum Genet 98, 116–126.
Burton, J.N., Adey, A., Patwardhan, R.P., Qiu, R., Kitzman, J.O., and Shendure, J. (2013). Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat Biotechnol 31, 1119–1125.
Cai, D.W., Han, L., Zhang, X.L., Zhou, H., and Zhu, H. (2007). DNA analysis of archaeological sheep remains from China. J Archaeol Sci 34, 1347–1355.
Cai, D., Tang, Z., Yu, H., Han, L., Ren, X., Zhao, X., Zhu, H., and Zhou, H. (2011). Early history of Chinese domestic sheep indicated by ancient DNA analysis of Bronze Age individuals. J Archaeol Sci 38, 896–902.
Capella-Gutiérrez, S., Silla-Martinez, J.M., and Gabaldon, T. (2009). trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973.
Chen, L., Qiu, Q., Jiang, Y., Wang, K., Lin, Z., Li, Z., Bibi, F., Yang, Y., Wang, J., Nie, W., et al. (2019). Large-scale ruminant genome sequencing provides insights into their evolution and distinct traits. Science 364, eaav6202.
Chen, S., Zhou, Y., Chen, Y., and Gu, J. (2018). fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890.
Chen, Y., Nie, F., Xie, S., Zheng, Y., Bray, T., Dai, Q., Wang, Y., Huang, Z., Wang, D., He, L., et al. (2020). Fast and accurate assembly of Nanopore reads via progressive error correction and adaptive read selection. bioRxiv, 2020.2002.2001.930107.
Daly, K.G., Maisano Delser, P., Mullin, V.E., Scheu, A., Mattiangeli, V., Teasdale, M.D., Hare, A.J., Burger, J., Verdugo, M.P., Collins, M.J., et al. (2018). Ancient goat genomes reveal mosaic domestication in the Fertile Crescent. Science 361, 85–88.
Danecek, P., Auton, A., Abecasis, G., Albers, C.A., Banks, E., DePristo, M. A., Handsaker, R.E., Lunter, G., Marth, G.T., Sherry, S.T., et al. (2011). The variant call format and VCFtools. Bioinformatics 27, 2156–2158.
Darriba, D., Taboada, G.L., Doallo, R., and Posada, D. (2012). jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9, 772.
De Magalhães, J.P., and Costa, J. (2009). A database of vertebrate longevity records and their relation to other life-history traits. J Evolary Biol 22, 1770–1774.
Demirci, S., Baştanlar, E.K., Dağtaş, N.D., Pişkin, E., Engin, A., Özer, F., Yüncü, E., Doğan, Ş.A., and Togan, İ. (2013). Mitochondrial DNA diversity of modern, ancient and wild sheep (Ovis gmelinii anatolica) from Turkey: new insights on the evolutionary history of sheep. PLoS One 8, e81952.
Dudchenko, O., Batra, S.S., Omer, A.D., Nyquist, S.K., Hoeger, M., Durand, N.C., Shamim, M.S., Machol, I., Lander, E.S., Aiden, A.P., et al. (2017). De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95.
Durand, N.C., Robinson, J.T., Shamim, M.S., Machol, I., Mesirov, J.P., Lander, E.S., and Aiden, E.L. (2016a). Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst 3, 99–101.
Durand, N.C., Shamim, M.S., Machol, I., Rao, S.S.P., Huntley, M.H., Lander, E.S., and Aiden, E.L. (2016b). Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst 3, 95–98.
Edgar, R.C. (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32, 1792–1797.
Fan, H., Hu, Y., Shan, L., Yu, L., Wang, B., Li, M., Wu, Q., and Wei, F. (2019). Synteny search identifies carnivore Y chromosome for evolution of male specific genes. Integr Zool 14, 224–234.
Faust, G.G., and Hall, I.M. (2014). SAMBLASTER: fast duplicate marking and structural variant read extraction. Bioinformatics 30, 2503–2505.
Hughes, J.F., Skaletsky, H., Brown, L.G., Pyntikova, T., Graves, T., Fulton, R.S., Dugan, S., Ding, Y., Buhay, C.J., Kremitzki, C., et al. (2012). Strict evolutionary conservation followed rapid gene loss on human and rhesus Y chromosomes. Nature 483, 82–86.
Hughes, J.F., Skaletsky, H., Pyntikova, T., Graves, T.A., van Daalen, S.K.M., Minx, P.J., Fulton, R.S., McGrath, S.D., Locke, D.P., Friedman, C., et al. (2010). Chimpanzee and human Y chromosomes are remarkably divergent in structure and gene content. Nature 463, 536–539.
Jain, M., Koren, S., Miga, K.H., Quick, J., Rand, A.C., Sasani, T.A., Tyson, J.R., Beggs, A.D., Dilthey, A.T., Fiddes, I.T., et al. (2018a). Nanopore sequencing and assembly of a human genome with ultra-long reads. Nat Biotechnol 36, 338–345.
Jain, M., Olsen, H.E., Turner, D.J., Stoddart, D., Bulazel, K.V., Paten, B., Haussler, D., Willard, H.F., Akeson, M., and Miga, K.H. (2018b). Linear assembly of a human centromere on the Y chromosome. Nat Biotechnol 36, 321–323.
Janečka, J.E., Davis, B.W., Ghosh, S., Paria, N., Das, P.J., Orlando, L., Schubert, M., Nielsen, M.K., Stout, T.A.E., Brashear, W., et al. (2018). Horse Y chromosome assembly displays unique evolutionary features and putative stallion fertility genes. Nat Commun 9, 2945.
Jiang, Y., Xie, M., Chen, W., Talbot, R., Maddox, J.F., Faraut, T., Wu, C., Muzny, D.M., Li, Y., Zhang, W., et al. (2014). The sheep genome illuminates biology of the rumen and lipid metabolism. Science 344, 1168–1173.
Kõressaar, T., Lepamets, M., Kaplinski, L., Raime, K., Andreson, R., and Remm, M. (2018). Primer3_masker: integrating masking of template sequence with primer design software. Bioinformatics 34, 1937–1938.
Korneliussen, T.S., Albrechtsen, A., and Nielsen, R. (2014). ANGSD: analysis of next generation sequencing data. BMC Bioinf 15, 356.
Kuderna, L.F.K., Lizano, E., Julià, E., Gomez-Garrido, J., Serres-Armero, A., Kuhlwilm, M., Alandes, R.A., Alvarez-Estape, M., Juan, D., Simon, H., et al. (2019). Selective single molecule sequencing and assembly of a human Y chromosome of African origin. Nat Commun 10, 4.
Letunic, I., and Bork, P. (2019). Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Res 47, W256–W259.
Li, G., Davis, B.W., Raudsepp, T., Pearks Wilkerson, A.J., Mason, V.C., Ferguson-Smith, M., O’Brien, P.C., Waters, P.D., and Murphy, W.J. (2013). Comparative analysis of mammalian Y chromosomes illuminates ancestral structure and lineage-specific evolution. Genome Res 23, 1486–1495.
Li, H. (2018). Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100.
Li, H., and Durbin, R. (2010). Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595.
Li, H., Handsaker, B., Wysoker, A., Fennell, T., Ruan, J., Homer, N., Marth, G., Abecasis, G., and Durbin, R. (2009). The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079.
Li, R., Fu, W., Su, R., Tian, X., Du, D., Zhao, Y., Zheng, Z., Chen, Q., Gao, S., Cai, Y., et al. (2019). Towards the complete goat pan-genome by recovering missing genomic segments from the reference genome. Front Genet 10, 1169.
Lv, F.H., Peng, W.F., Yang, J., Zhao, Y.X., Li, W.R., Liu, M.J., Ma, Y.H., Zhao, Q.J., Yang, G.L., Wang, F., et al. (2015). Mitogenomic meta-analysis identifies two phases of migration in the history of eastern Eurasian sheep. Mol Biol Evol 32, 2515–2533.
Ma, Y.H., Rao, S.Q., Lu, S.J., Hou, G.Y., Guan, W.J., Li, H.B., Li, X., Zhao, Q.J., and Guo, J. (2006). Phylogeography and origin of sheep breeds in Northern China. Conserv Genet 7, 117–127.
McKenna, A., Hanna, M., Banks, E., Sivachenko, A., Cibulskis, K., Kernytsky, A., Garimella, K., Altshuler, D., Gabriel, S., Daly, M., et al. (2010). The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20, 1297–1303.
Meadows, J.R.S., Hanotte, O., Drögemüller, C., Calvo, J., Godfrey, R., Coltman, D., Maddox, J.F., Marzanov, N., Kantanen, J., and Kijas, J.W. (2006). Globally dispersed Y chromosomal haplotypes in wild and domestic sheep. Anim Genets 37, 444–453.
Meadows, J.R.S., Hawken, R.J., and Kijas, J.W. (2004). Nucleotide diversity on the ovine Y chromosome. Anim Genets 35, 379–385.
Meadows, J.R.S., and Kijas, J.W. (2009). Re-sequencing regions of the ovine Y chromosome in domestic and wild sheep reveals novel paternal haplotypes. Anim Genets 40, 119–123.
Meadows, J.R.S., Cemal, I., Karaca, O., Gootwine, E., and Kijas, J.W. (2007). five ovine mitochondrial lineages identified from sheep breeds of the near East. Genetics 175, 1371–1379.
Naval-Sanchez, M., Nguyen, Q., McWilliam, S., Porto-Neto, L.R., Tellam, R., Vuocolo, T., Reverter, A., Perez-Enciso, M., Brauning, R., Clarke, S., et al. (2018). Sheep genome functional annotation reveals proximal regulatory elements contributed to the evolution of modern breeds. Nat Commun 9, 859.
Nguyen, L.T., Schmidt, H.A., von Haeseler, A., and Minh, B.Q. (2015). IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32, 268–274.
Nijman, I.J., Rosen, B.D., Zheng, Z., Jiang, Y., Cumer, T., Daly, K.G., Bâlteanu, V.A., Berger, B., Blichfeldt, T., Boink, G., et al. (2020). Phylogeny and distribution of Y-chromosomal haplotypes in domestic, ancient and wild goats. bioRxiv, 2020.2002.2017.952051.
Pan, Z., Li, S., Liu, Q., Wang, Z., Zhou, Z., Di, R., Miao, B., Hu, W., Wang, X., Hu, X., et al. (2018). Whole-genome sequences of 89 Chinese sheep suggest role of RXFP2 in the development of unique horn phenotype as response to semi-feralization. GigaScience 7.
Paradis, E. (2010). pegas: an R package for population genetics with an integrated-modular approach. Bioinformatics 26, 419–420.
Pedrosa, S., Uzun, M., Arranz, J.J., Gutiérrez-Gil, B., San Primitivo, F., and Bayón, Y. (2005). Evidence of three maternal lineages in near eastern sheep supporting multiple domestication events. Proc R Soc B 272, 2211–2217.
Rannamäe, E., Lõugas, L., Niemi, M., Kantanen, J., Maldre, L., Kadõrova, N., and Saarma, U. (2016). Maternal and paternal genetic diversity of ancient sheep in Estonia from the Late Bronze Age to the post-medieval period and comparison with other regions in Eurasia. Anim Genet 47, 208–218.
Rezaei, H.R., Naderi, S., Chintauan-Marquier, I.C., Taberlet, P., Virk, A.T., Naghash, H.R., Rioux, D., Kaboli, M., and Pompanon, F. (2010). Evolution and taxonomy of the wild species of the genus Ovis (Mammalia, Artiodactyla, Bovidae). Mol Phylogenet Evol 54, 315–326.
Rocha, J., Chen, S., and Beja-Pereira, A. (2011). Molecular evidence for fat-tailed sheep domestication. Trop Anim Health Prod 43, 1237–1243.
Schiffels, S., and Durbin, R. (2014). Inferring human population size and separation history from multiple genome sequences. Nat Genet 46, 919–925.
Shen, W., Le, S., Li, Y., and Hu, F. (2016). SeqKit: a cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS ONE 11, e0163962.
Simão, F.A., Waterhouse, R.M., Ioannidis, P., Kriventseva, E.V., and Zdobnov, E.M. (2015). BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212.
Skinner, B.M., Sargent, C.A., Churcher, C., Hunt, T., Herrero, J., Loveland, J.E., Dunn, M., Louzada, S., Fu, B., Chow, W., et al. (2016). The pig X and Y Chromosomes: structure, sequence, and evolution. Genome Res 26, 130–139.
Soh, Y.Q.S., Alföldi, J., Pyntikova, T., Brown, L.G., Graves, T., Minx, P.J., Fulton, R.S., Kremitzki, C., Koutseva, N., Mueller, J.L., et al. (2014). Sequencing the mouse Y chromosome reveals convergent gene acquisition and amplification on both sex chromosomes. Cell 159, 800–813.
Tapio, M., Marzanov, N., Ozerov, M., Cinkulov, M., Gonzarenko, G., Kiselyova, T., Murawski, M., Viinalass, H., and Kantanen, J. (2006). Sheep mitochondrial DNA variation in European, Caucasian, and Central Asian areas. Mol Biol Evol 23, 1776–1783.
Tarasov, A., Vilella, A.J., Cuppen, E., Nijman, I.J., and Prins, P. (2015). Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034.
Tomaszkiewicz, M., Medvedev, P., and Makova, K.D. (2017). Y and W chromosome assemblies: approaches and discoveries. Trends Genets 33, 266–282.
Tomaszkiewicz, M., Rangavittal, S., Cechova, M., Sanchez, R.C., Fescemyer, H.W., Harris, R., Ye, D., O’Brien, P.C.M., Chikhi, R., Ryder, O.A., et al. (2016). A time- and cost-effective strategy to sequence mammalian Y chromosomes: an application to the de novo assembly of gorilla Y. Genome Res 26, 530–540.
Auwera, G.A., Carneiro, M.O., Hartl, C., Poplin, R., del Angel, G., Levy-Moonshine, A., Jordan, T., Shakir, K., Roazen, D., Thibault, J., et al. (2013). From FastQ data to high-confidence variant calls: the genome analysis toolkit best practices pipeline. Curr Protoc Bioinf 43, 11.10.1–11.10.33.
Vaser, R., Sović, I., Nagarajan, N., and Šikić, M. (2017). Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res 27, 737–746.
Verdugo, M.P., Mullin, V.E., Scheu, A., Mattiangeli, V., Daly, K.G., Maisano Delser, P., Hare, A.J., Burger, J., Collins, M.J., Kehati, R., et al. (2019). Ancient cattle genomics, origins, and rapid turnover in the Fertile Crescent. Science 365, 173–176.
Walker, B.J., Abeel, T., Shea, T., Priest, M., Abouelliel, A., Sakthikumar, S., Cuomo, C.A., Zeng, Q., Wortman, J., Young, S.K., et al. (2014). Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963.
Wallner, B., Palmieri, N., Vogl, C., Rigler, D., Bozlak, E., Druml, T., Jagannathan, V., Leeb, T., Fries, R., Tetens, J., et al. (2017). Y chromosome uncovers the recent oriental origin of modern stallions. Curr Biol 27, 2029–2035.e5.
Wang, X., Liu, J., Niu, Y., Li, Y., Zhou, S., Li, C., Ma, B., Kou, Q., Petersen, B., Sonstegard, T., et al. (2018). Low incidence of SNVs and indels in trio genomes of Cas9-mediated multiplex edited sheep. BMC Genomics 19, 397.
Wang, X., Zheng, Z., Cai, Y., Chen, T., Li, C., Fu, W., and Jiang, Y. (2017). CNVcaller: highly efficient and widely applicable software for detecting copy number variations in large populations. GigaScience 6.
Wong, K.H.Y., Levy-Sakin, M., and Kwok, P.Y. (2018). De novo human genome assemblies reveal spectrum of alternative haplotypes in diverse populations. Nat Commun 9, 3040.
Wu, T.D., Reeder, J., Lawrence, M., Becker, G., and Brauer, M.J. (2016). GMAP and GSNAP for genomic sequence alignment: enhancements to speed, accuracy, and functionality. Stat Genom 1418, 283–334.
Yang, J., Li, W. R., Lv, F. H., He, S. G., Tian, S. L., Peng, W. F., Sun, Y. W., Zhao, Y.X., Tu, X.L., Zhang, M., et al. (2016). Whole-genome sequencing of native sheep provides insights into rapid adaptations to extreme environments. Mol Biol Evol 33, 2576–2592.
Zeder, M.A. (2008). Domestication and early agriculture in the Mediterranean Basin: Origins, diffusion, and impact. Proc Natl Acad Sci USA 105, 11597–11604.
Zhang, G., Vahidi, S.M.F., Ma, Y.H., and Han, J.L. (2012). Limited polymorphisms of two Y-chromosomal SNPs in Chinese and Iranian sheep. Anim Genets 43, 479–480.
Zhang, M., Peng, W.F., Yang, G.L., Lv, F.H., Liu, M.J., Li, W.R., Liu, Y.G., Li, J.Q., Wang, F., Shen, Z.Q., et al. (2014). Y chromosome haplotype diversity of domestic sheep (Ovis aries) in northern Eurasia. Anim Genet 45, 903–907.
Zhao, Y.X., Yang, J., Lv, F.H., Hu, X.J., Xie, X.L., Zhang, M., Li, W.R., Liu, M.J., Wang, Y.T., Li, J.Q., et al. (2017). Genomic reconstruction of the history of native sheep reveals the peopling patterns of nomads and the expansion of early pastoralism in East Asia. Mol Biol Evol 34, 2380–2395.
Zheng, Z., Wang, X., Li, M., Li, Y., Yang, Z., Wang, X., Pan, X., Gong, M., Zhang, Y., Guo, Y., et al. (2020). The origin of domestication genes in goats. Sci Adv 6, eaaz5216.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31822052) and the National Thousand Youth Talents Plan, Natural Science Foundation of China (31802027), and Natural Science Basic Research Plan in Shaanxi Province of China (2019JQ-002). We thank the High-Performance Computing platform of Northwest A&F University for providing computing resources. We also thank members of the NextGen project for sharing their data.
Author information
Authors and Affiliations
Corresponding author
Additional information
Compliance and ethics
The author(s) declare that they have no conflict of interest. Blood samples were taken by conforming with the Helsinki Declaration of 1975 (as revised in 2008) concerning Animal Rights.
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Li, R., Yang, P., Li, M. et al. A Hu sheep genome with the first ovine Y chromosome reveal introgression history after sheep domestication. Sci. China Life Sci. 64, 1116–1130 (2021). https://doi.org/10.1007/s11427-020-1807-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-020-1807-0