Abstract
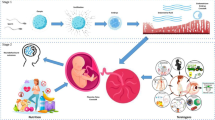
It is well established that the decline in female reproductive outcomes is related to postovulatory aging of oocytes and advanced maternal age. Poor oocyte quality is correlated with compromised genetic integrity and epigenetic changes during the oocyte aging process. Here, we review the epigenetic alterations, mainly focused on DNA methylation, histone acetylation and methylation associated with postovulatory oocyte aging as well as advanced maternal age. Furthermore, we address the underlying epigenetic mechanisms that contribute to the decline in oocyte quality during oocyte aging.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Jones K T. Meiosis in oocytes: predisposition to aneuploidy and its increased incidence with age. Hum Reprod Update, 2008, 14: 143–158
Wang Z B, Schatten H, Sun Q Y. Why is chromosome segregation error in oocytes increased with maternal aging? Physiology (Bethesda), 2011, 26: 314–325
Wang Q, Sun Q Y. Evaluation of oocyte quality: morphological, cellular and molecular predictors. Reprod Fertil Dev, 2007, 19: 1–12
Austin C R. In: Whittaher J R, ed. Concepts of Development. Sinauer Associates Inc., 1974
Kikuchi K, Naito K, Noguchi J, et al. Maturation/M-phase promoting factor: a regulator of aging in porcine oocytes. Biol Reprod, 2000, 63: 715–722
Miao Y L, Kikuchi K, Sun Q Y, et al. Oocyte aging: cellular and molecular changes, developmental potential and reversal possibility. Hum Reprod Update, 2009, 15: 573–585
Tatone C, Amicarelli F, Carbone M C, et al. Cellular and molecular aspects of ovarian follicle aging. Hum Reprod Update, 2008, 14: 131–142
Ma J Y, Liang X W, Schatten H, et al. Active DNA demethylation in mammalian preimplantation embryos: new insights and new perspectives. Mol Hum Reprod, 2012, 18: 333–340
Reik W, Dean W, Walter J. Epigenetic reprogramming in mammalian development. Science, 2001, 293: 1089–1093
Verona R I, Mann M R, Bartolomei M S. Genomic imprinting: intricacies of epigenetic regulation in clusters. Annu Rev Cell Dev Biol, 2003, 19: 237–259
Bartolomei M S, Ferguson-Smith A C. Mammalian genomic imprinting. Cold Spring Harb Perspect Biol, 2011, 3: a002592
Reik W. Stability and flexibility of epigenetic gene regulation in mammalian development. Nature, 2007, 447: 425–432
Hiura H, Obata Y, Komiyama J, et al. Oocyte growth-dependent progression of maternal imprinting in mice. Genes Cells, 2006, 11: 353–361
La Salle S, Mertineit C, Taketo T, et al. Windows for sex-specific methylation marked by DNA methyltransferase expression profiles in mouse germ cells. Dev Biol, 2004, 268: 403–415
Lucifero D, La Salle S, Bourc’his D, et al. Coordinate regulation of DNA methyltransferase expression during oogenesis. BMC Dev Biol, 2007, 7: 36
Lucifero D, Mann M R, Bartolomei M S, et al. Gene-specific timing and epigenetic memory in oocyte imprinting. Hum Mol Genet, 2004, 13: 839–849
Bestor T H. The DNA methyltransferases of mammals. Hum Mol Genet, 2000, 9: 2395–2402
Branco M R, Oda M, Reik W. Safeguarding parental identity: Dnmt1 maintains imprints during epigenetic reprogramming in early embryogenesis. Genes Dev, 2008, 22: 1567–1571
Allfrey V G, Faulkner R, Mirsky A E. Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc Natl Acad Sci USA, 1964, 51: 786–794
Gutierrez R M, Hnilica L S. Tissue specificity of histone phosphorylation. Science, 1967, 157: 1324–1325
Huletsky A, Niedergang C, Frechette A, et al. Sequential ADP-ribosylation pattern of nucleosomal histones. ADP-ribosylation of nucleosomal histones. Eur J Biochem, 1985, 146: 277–285
Nathan D, Ingvarsdottir K, Sterner D E, et al. Histone sumoylation is a negative regulator in Saccharomyces cerevisiae and shows dynamic interplay with positive-acting histone modifications. Genes Dev, 2006, 20: 966–976
Koprinarova M A, Russev G C. Dynamics of histone H4 acetylation during the cell cycle. Cell Cycle, 2008, 7: 414–416
Rada-Iglesias A, Enroth S, Andersson R, et al. Histone H3 lysine 27 trimethylation in adult differentiated colon associated to cancer DNA hypermethylation. Epigenetics, 2009, 4: 107–113
Xu D, Bai J, Duan Q, et al. Covalent modifications of histones during mitosis and meiosis. Cell Cycle, 2009, 8: 3688–3694
Kouzarides T. Chromatin modifications and their function. Cell, 2007, 128: 693–705
Strahl B D, Allis C D. The language of covalent histone modifications. Nature, 2000, 403: 41–45
Wang N, Tilly J L. Epigenetic status determines germ cell meiotic commitment in embryonic and postnatal mammalian gonads. Cell Cycle, 2010, 9: 339–349
Gu L, Wang Q, Sun Q Y. Histone modifications during mammalian oocyte maturation: dynamics, regulation and functions. Cell Cycle, 2010, 9: 1942–1950
Jiang G J, Wang K, Miao D Q, et al. Protein profile changes during porcine oocyte aging and effects of caffeine on protein expression patterns. PLoS ONE, 2011, 6: e28996
Yue M X, Fu X W, Zhou G B, et al. Abnormal DNA methylation in oocytes could be associated with a decrease in reproductive potential in old mice. J Assist Reprod Genet, 2012, 29: 643–650
Lopes F L, Fortier A L, Darricarrere N, et al. Reproductive and epigenetic outcomes associated with aging mouse oocytes. Hum Mol Genet, 2009, 18: 2032–2044
Holinka C F, Tseng Y C, Finch C E. Reproductive aging in C57BL/6J mice: plasma progesterone, viable embryos and resorption frequency throughout pregnancy. Biol Reprod, 1979, 20: 1201–1211
Manosalva I, Gonzalez A. Aging changes the chromatin configuration and histone methylation of mouse oocytes at germinal vesicle stage. Theriogenology, 2010, 74: 1539–1547
Ooga M, Inoue A, Kageyama S, et al. Changes in H3K79 methylation during preimplantation development in mice. Biol Reprod, 2008, 78: 413–424
Park K E, Magnani L, Cabot R A. Differential remodeling of mono- and trimethylated H3K27 during porcine embryo development. Mol Reprod Dev, 2009, 76: 1033–1042
Hou J, Liu L, Zhang J, et al. Epigenetic modification of histone 3 at lysine 9 in sheep zygotes and its relationship with DNA methylation. BMC Dev Biol, 2008, 8: 60
Racedo S E, Wrenzycki C, Lepikhov K, et al. Epigenetic modifications and related mRNA expression during bovine oocyte in vitro maturation. Reprod Fertil Dev, 2009, 21: 738–748
Qiao J, Chen Y, Yan L Y, et al. Changes in histone methylation during human oocyte maturation and IVF-or ICSI-derived embryo development. Fertil Steril, 2010, 93: 1628–1636
Akiyama T, Nagata M, Aoki F. Inadequate histone deacetylation during oocyte meiosis causes aneuploidy and embryo death in mice. Proc Natl Acad Sci USA, 2006, 103: 7339–7344
Manosalva I, Gonzalez A. Aging alters histone H4 acetylation and CDC2A in mouse germinal vesicle stage oocytes. Biol Reprod, 2009, 81: 1164–1171
van den Berg I M, Eleveld C, van der Hoeven M, et al. Defective deacetylation of histone 4 K12 in human oocytes is associated with advanced maternal age and chromosome misalignment. Hum Reprod, 2011, 26: 1181–1190
Lucifero D, Mertineit C, Clarke H J, et al. Methylation dynamics of imprinted genes in mouse germ cells. Genomics, 2002, 79: 530–538
Li J Y, Lees-Murdock D J, Xu G L, et al. Timing of establishment of paternal methylation imprints in the mouse. Genomics, 2004, 84: 952–960
Liang X W, Zhu J Q, Miao Y L, et al. Loss of methylation imprint of SNRPN in postovulatory aging mouse oocyte. Biochem Biophys Res Commun, 2008, 371: 16–21
Imamura T, Kerjean A, Heams T, et al. Dynamic CpG and non-CpG methylation of the Peg1/Mest gene in the mouse oocyte and preimplantation embryo. J Biol Chem, 2005, 280: 20171–20175
Tarin J J, Perez-Albala S, Aguilar A, et al. Long-term effects of postovulatory aging of mouse oocytes on offspring: a two-generational study. Biol Reprod, 1999, 61: 1347–1355
Tarin J J, Perez-Albala S, Perez-Hoyos S, et al. Postovulatory aging of oocytes decreases reproductive fitness and longevity of offspring. Biol Reprod, 2002, 66: 495–499
Liang X W, Ge Z J, Guo L, et al. Effect of postovulatory oocyte aging on DNA methylation imprinting acquisition in offspring oocytes. Fertil Steril, 2011, 96: 1479–1484
Huang J C, Yan L Y, Lei Z L, et al. Changes in histone acetylation during postovulatory aging of mouse oocyte. Biol Reprod, 2007, 77: 666–670
Liu N, Wu Y G, Lan G C, et al. Pyruvate prevents aging of mouse oocytes. Reproduction, 2009, 138: 223–234
Cui M S, Wang X L, Tang D W, et al. Acetylation of H4K12 in porcine oocytes during in vitro aging: potential role of ooplasmic reactive oxygen species. Theriogenology, 2011, 75: 638–646
Yoshida M, Kijima M, Akita M, et al. Potent and specific inhibition of mammalian histone deacetylase both in vivo and in vitro by trichostatin A. J Biol Chem, 1990, 265: 17174–17179
De La Fuente R, Viveiros M M, Wigglesworth K, et al. ATRX, a member of the SNF2 family of helicase/ATPases, is required for chromosome alignment and meiotic spindle organization in metaphase II stage mouse oocytes. Dev Biol, 2004, 272: 1–14
Wang Q, Yin S, Ai J S, et al. Histone deacetylation is required for orderly meiosis. Cell Cycle, 2006, 5: 766–774
Pan H, Ma P, Zhu W, et al. Age-associated increase in aneuploidy and changes in gene expression in mouse eggs. Dev Biol, 2008, 316: 397–407
Hamatani T, Falco G, Carter M G, et al. Age-associated alteration of gene expression patterns in mouse oocytes. Hum Mol Genet, 2004, 13: 2263–2278
Ratnam S, Mertineit C, Ding F, et al. Dynamics of Dnmt1 methyltransferase expression and intracellular localization during oogenesis and preimplantation development. Dev Biol, 2002, 245: 304–314
Hirasawa R, Chiba H, Kaneda M, et al. Maternal and zygotic Dnmt1 are necessary and sufficient for the maintenance of DNA methylation imprints during preimplantation development. Genes Dev, 2008, 22: 1607–1616
Kurihara Y, Kawamura Y, Uchijima Y, et al. Maintenance of genomic methylation patterns during preimplantation development requires the somatic form of DNA methyltransferase 1. Dev Biol, 2008, 313: 335–346
Mohan K N, Ding F, Chaillet J R. Distinct roles of DMAP1 in mouse development. Mol Cell Biol, 2011, 31: 1861–1869
Zhang L, Lu D Y, Ma W Y, et al. Age-related changes in the localization of DNA methyltransferases during meiotic maturation in mouse oocytes. Fertil Steril, 2011, 95: 1531–1534
Lees-Murdock D J, Shovlin T C, Gardiner T, et al. DNA methyltransferase expression in the mouse germ line during periods of de novo methylation. Dev Dyn, 2005, 232: 992–1002
Wood A J, Oakey R J. Genomic imprinting in mammals: emerging themes and established theories. PLoS Genet, 2006, 2: e147
Suo L, Meng Q G, Pei Y, et al. Changes in acetylation on lysine 12 of histone H4 (acH4K12) of murine oocytes during maternal aging may affect fertilization and subsequent embryo development. Fertil Steril, 2010, 93: 945–951
Yu J N, Wang M, Wang D Q, et al. Chromosome changes of aged oocytes after ovulation. Yi Chuan, 2007, 29: 225–229
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Liang, X., Ma, J., Schatten, H. et al. Epigenetic changes associated with oocyte aging. Sci. China Life Sci. 55, 670–676 (2012). https://doi.org/10.1007/s11427-012-4354-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-012-4354-3