Abstract
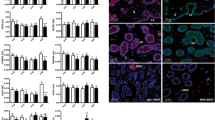
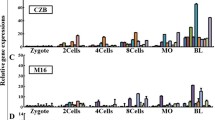
To obtain reliable results in quantitative PCR (qPCR) reactions, an endogenous control (EC) gene is needed to correct for systematic variations. In this study, a TaqMan low density array was used to quantify the expression levels of microRNA (miRNA) genes in in vivo fertilized, in vitro fertilized, parthenogenetic and somatic cell nuclear transfer blastocysts. The aim was to identify suitable EC genes for the qPCR analysis of miRNAs in porcine blastocysts. The results showed that thirty-six miRNAs were commonly expressed in the four kinds of embryos and the expression levels of eleven miRNAs were similar in the different embryo types (P-value>0.05). These 11 miRNAs were selected as candidate EC genes for further analysis and, of these, miR-16 was identified as the most stable EC gene by the GeNorm (a tool based on a pair-wise comparison model that calculates the internal control genes stability measure and determines the most reliable pair of EC genes) and NormFinder (an excel plug-in that uses an ANOVA-based model to estimate intra- and inter-group variation to indicate the single most stable EC gene) programs. In addition, a cell number normalization method validated miR-16 as a suitable EC gene for use in future qPCR analysis of miRNAs in porcine blastocysts.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Brown B D, Naldini L. Exploiting and antagonizing microRNA regulation for therapeutic and experimental applications. Nat Rev Genet, 2009, 10: 578–585
Cordes K R, Sheehy N T, White M P, et al. miR-145 and miR-143 regulate smooth muscle cell fate and plasticity. Nature, 2009, 460: 705–710
Eulalio A, Huntzinger E, Izaurralde E. Getting to the root of miRNA-mediated gene silencing. Cell, 2008, 132: 9–14
Xie X, Lu J, Kulbokas E J, et al. Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature, 2005, 434: 338–345
Foshay K M, Gallicano G I. miR-17 family miRNAs are expressed during early mammalian development and regulate stem cell differentiation. Dev Biol, 2009, 326: 431–443
Kanellopoulou C, Muljo S A, Kung A L, et al. Dicer-deficient mouse embryonic stem cells are defective in differentiation and centromeric silencing. Genes Dev, 2005, 19: 489–501
Murchison E P, Partridge J F, Tam O H, et al. Characterization of Dicer-deficient murine embryonic stem cells. Proc Natl Acad Sci USA, 2005, 102: 12135–12140
Benetti R, Gonzalo S, Jaco I, et al. A mammalian microRNA cluster controls DNA methylation and telomere recombination via Rbl2-dependent regulation of DNA methyltransferases. Nat Struct Mol Biol, 2008, 15: 268–279
Duursma A M, Kedde M, Schrier M, et al. miR-148 targets human DNMT3b protein coding region. RNA, 2008, 14: 872–877
Fabbri M, Garzon R, Cimmino A, et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc Natl Acad Sci USA, 2007, 104: 15805–15810
Bernstein E, Kim S Y, Carmell M A, et al. Dicer is essential for mouse development. Nat Genet, 2003, 35: 215–217
Tang F, Kaneda M, O’Carroll D, et al. Maternal microRNAs are essential for mouse zygotic development. Genes Dev, 2007, 21: 644–648
Davoren P A, McNeill R E, Lowery A J, et al. Identification of suitable endogenous control genes for microRNA gene expression analysis in human breast cancer. BMC Mol Biol, 2008, 9: 76
Stamova B S, Apperson M, Walker W L, et al. Identification and validation of suitable endogenous reference genes for gene expression studies in human peripheral blood. BMC Med Genomics, 2009, 2: 49
Reddy A M, Zheng Y, Jagadeeswaran G, et al. Cloning, characterization and expression analysis of porcine microRNAs. BMC Genomics, 2009, 10: 65
Wei H X, Zhang K, Ma Y F, et al. Stage-dependent effect of leptin on development of porcine embryos derived from parthenogenetic activation and transgenic somatic cell nuclear transfer. J Reprod Dev, 2009, 55: 99–104
Zhang K, Wei H X, Zhang Y H, et al. Effects of ghrelin on in vitro development of porcine in vitro fertilized and parthenogenetic embryos. J Reprod Dev, 2007, 53: 647–653
Chen C, Ridzon D A, Broomer A J, et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res, 2005, 33: e179
Tang F, Hajkova P, Barton S C, et al. MicroRNA expression profiling of single whole embryonic stem cells. Nucleic Acids Res, 2006, 34: e9
Livak K J, Schmittgen T D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods, 2001, 25: 402–408
Vandesompele J, De Preter K, Pattyn F, et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol, 2002, 3: research0034-research0034.11
Andersen C L, Jensen J L, Orntoft T F. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res, 2004, 64: 5245–5250
Wong M L, Medrano J F. Real-time PCR for mRNA quantitation. Biotechniques, 2005, 39: 75–85
Runte M, Huttenhofer A, Gross S, et al. The IC-SNURF-SNRPN transcript serves as a host for multiple small nucleolar RNA species and as an antisense RNA for UBE3A. Hum Mol Genet, 2001, 10: 2687–2700
Peltier H J, Latham G J. Normalization of microRNA expression levels in quantitative RT-PCR assays: identification of suitable reference RNA targets in normal and cancerous human solid tissues. RNA, 2008, 14: 844–852
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Li, J., Wei, H., Li, Y. et al. Identification of a suitable endogenous control gene in porcine blastocysts for use in quantitative PCR analysis of microRNAs. Sci. China Life Sci. 55, 126–131 (2012). https://doi.org/10.1007/s11427-012-4289-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-012-4289-8