Abstract
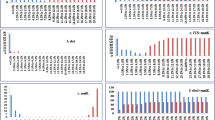
DNA barcoding is a rapidly developing frontier technology that is gaining worldwide attention. Here, seven regions (psbA-trnH, matK, ycf5, rpoC1, rbcL, ITS2, and ITS) with potential for use as DNA barcodes were tested for their ability to identify 300 samples of 192 species from 72 genera of the family Rutaceae. To evaluate each barcode’s utility for species authentication, PCR amplification efficiency, genetic divergence, and barcoding gaps were assessed. We found that the ITS2 region exhibited the highest inter-specific divergence, and that this was significantly higher than the intra-specific variation in the “DNA barcoding gap” assessment and Wilcoxon two-sample tests. The ITS2 locus had the highest identification efficiency among all tested regions. In a previous study, we found that ITS2 was able to discriminate a wide range of plant taxa, and here we confirmed that ITS2 was also able to discriminate a number of closely related species. Therefore, we propose that ITS2 is a promising candidate barcode for plant species identification.
Similar content being viewed by others
References
Hebert P D, Cywinska A, Ball S L, et al. Biological identifications through DNA barcodes. Proc R Soc Biol Sci SerB, 2003, 270: 313–321 10.1098/rspb.2002.2218, 1:CAS:528:DC%2BD3sXktVWiu7g%3D
Gregory T R. DNA barcoding does not compete with taxonomy. Nature, 2005, 434: 1067 10.1038/4341067b, 15858548, 1:CAS:528:DC%2BD2MXjsF2ltrk%3D
Miller S E. DNA barcoding and the renaissance of taxonomy. Proc Natl Acad Sci USA, 2007, 104: 4775–4776 10.1073/pnas.0700466104, 17363473, 1:CAS:528:DC%2BD2sXjvFCjsLo%3D
Hebert P D, Ratnasingham S, deWaard J R. Barcoding animal life: cytochrome c oxidase subunit 1 divergences among closely related species. Proc R Soc Biol Sci SerB, 2003, 270: S96–S99 10.1098/rsbl.2003.0025, 1:CAS:528:DC%2BD3sXns1Smsbo%3D
Vences M, Thomas M, Bonett R M, et al. Deciphering amphibian diversity through DNA barcoding: chances and challenges. Philos Trans R Soc Lond B Biol Sci, 2005, 360: 1859–1868 10.1098/rstb.2005.1717, 16221604, 1:CAS:528:DC%2BD2MXhtlSjsrjE
Janzen D H, Hajibabaei M, Burns J M, et al. Wedding biodiversity inventory of a large and complex Lepidoptera fauna with DNA barcoding. Philos Trans R Soc Lond B Biol Sci, 2005, 360: 1835–1845 10.1098/rstb.2005.1715, 16214742, 1:CAS:528:DC%2BD2MXhtlSjsrjI
Ward R D, Zemlak T S, Innes B H, et al. DNA barcoding Australia’s fish species. Philos Trans R Soc Lond B Biol Sci, 2005}, 360}: 1847–1 10.1098/rstb.2005.1716, 16214743, 1:CAS:528:DC%2BD2MXhtlSjsrjK
Kress W J, Wurdack K J, Zimmer E A, et al. Use of DNA barcodes to identify flowering plants. Proc Natl Acad Sci USA, 2005, 102: 8369–8374 10.1073/pnas.0503123102, 15928076, 1:CAS:528:DC%2BD2MXlsV2mtbY%3D
Newmaster S G, Fazekas A J, Ragupathy S. DNA barcoding in land plants: evaluation of rbcL in a multigene tiered approach. Can J Bot, 2006, 84: 335–341 10.1139/B06-047, 1:CAS:528:DC%2BD28XotFeis7c%3D
Chase M W, Cowan R S, Hollingsworth P M, et al. A proposal for a standardised protocol to barcode all land plants. Taxon, 2007, 56: 295–299
Kress W J, Erickson D L. A two-locus global DNA barcode for land plants: The coding rbcL gene complements the non-coding trnH-psbA spacer region. PLoS ONE, 2007, 2: e508 10.1371/journal.pone.0000508, 17551588, 1:CAS:528:DC%2BD2sXmvFyrs7Y%3D
Pennisi E. Wanted: a barcode for plants. Science, 2007, 318: 190–191 10.1126/science.318.5848.190, 17932267, 1:CAS:528:DC%2BD2sXhtFyrs7rE
CBOL Plant Working Group. A DNA barcode for land plants. Proc Natl Acad Sci USA, 2009, 106: 12794–12797 10.1073/pnas.0905845106
Chen S L, Yao H, Han J P, et al. Validation of the ITS2 Region as a Novel DNA Barcode for Identifying Medicinal Plant Species. PLoS ONE, 2010, 5: e8613 10.1371/journal.pone.0008613, 20062805, 1:CAS:528:DC%2BC3cXmtlertQ%3D%3D
Song J Y, Yao H, Li Y, et al. Authentication of the family Polygonaceae in Chinese pharmacopoeia by DNA barcoding technique. J Ethnopharmacol, 2009, 124: 434–439 10.1016/j.jep.2009.05.042, 19505556, 1:CAS:528:DC%2BD1MXptVSisbg%3D
Lahaye R, van der Bank M, Bogarin D, et al. DNA barcoding the floras of biodiversity hotspots. Proc Natl Acad Sci USA, 2008, 105: 2923–2928 10.1073/pnas.0709936105, 18258745
Sass C, Little D P, Stevenson D W, et al. DNA Barcoding in the Cycadales: Testing the potential of proposed barcoding markers for species identification of Cycads. PLoS ONE, 2007, 2: e1154 10.1371/journal.pone.0001154, 17987130, 1:CAS:528:DC%2BD1cXjslShsg%3D%3D
Zhu Y J, Chen S L, Yao H, et al. DNA barcoding for the identification plants of the genus Paris. Acta Pharmaceutica Sinica, 2010, 45: 376–382.
Keller A, Schleicher T, Schultz J, et al. 5.8S–28S rRNA interaction and HMM-based ITS2 annotation. Gene, 2009, 430: 50–57 10.1016/j.gene.2008.10.012, 19026726, 1:CAS:528:DC%2BD1MXovV2l
Meyer C P, Paulay G. DNA barcoding: error rates based on comprehensive sampling. PLoS Biol, 2005, 3: e422 10.1371/journal.pbio.0030422, 16336051, 1:CAS:528:DC%2BD2MXhtlWls7nL
Slabbinck B, Dawyndt P, Martens M, et al. TaxonGap: a visualization tool for intra- and inter-species variation among individual biomarkers. Bioinformatics, 2008, 24: 866–867 10.1093/bioinformatics/btn031, 18227116, 1:CAS:528:DC%2BD1cXjsVaktLw%3D
Ross H A, Murugan S, Li W L. Testing the reliability of genetic methods of species identification via simulation. Syst Biol, 2008, 57: 216–230 10.1080/10635150802032990, 18398767
Chase M W, Salamin N, Wilkinson M, et al. Land plants and DNA barcodes: short-term and long-term goals. Philos Trans R Soc Lond B Biol Sci, 2005, 360: 1889–1895 10.1098/rstb.2005.1720, 16214746, 1:CAS:528:DC%2BD2MXhtlSjsrnP
Gonzalez M A, Baraloto C, Engel J, et al. Identification of Amazonian trees with DNA barcodes. PLoS One, 2009, 4: e7483 10.1371/journal.pone.0007483, 19834612, 1:CAS:528:DC%2BD1MXhtlWqtbnI
Müller T, Philippi N, Dandekar T, et al. Distinguishing species. RNA, 2007, 13: 1469–1472 10.1261/rna.617107, 17652131, 1:CAS:528:DC%2BD2sXhtVSqt7jO
Miao M, Warren A, Song W, et al. Analysis of the internal transcribed spacer 2 (ITS2) region of scuticociliates and related taxa (Ciliophora, Oligohymenophorea) to infer their evolution and phylogeny. Protist, 2008, 159: 519–533 10.1016/j.protis.2008.05.002, 18675585, 1:CAS:528:DC%2BD1MXjs1Khsg%3D%3D
Shaw J, Lickey E B, Beck J T, et al. The tortoise and the hare. II. Relative utility of 21 noncoding chloroplast DNA sequences for phylogenetic analysis. Am J Bot, 2005, 92: 142–166 10.3732/ajb.92.1.142, 1:CAS:528:DC%2BD2MXht1Klsbc%3D
Yao H, Song J Y, Ma X Y, et al. Identification of Dendrobium species by a candidate DNA barcode sequence: The chloroplast psbA-trnH intergenic region. Planta Med, 2009, 75: 667–669 10.1055/s-0029-1185385, 19235685, 1:CAS:528:DC%2BD1MXmsFWhu7g%3D
Walter R, Herbert J W, Leon D B. The Citrus industry (Vol I). California: University of California Press, 1967. 190–430
Tanaka T. Taxonomic problem of Citrus fruits in the Orient. Bull Univ Osaka Pref Ser B, 1969, 21: 133–138
Zeng M. The knowledge comprehension and arrangement of Citrus. China Fruit, 1960, (2): 31–37
Francois L, Frederic L, Joseph M B, et al. DNA amplified fingerprinting, a useful tool for determination of genetic origin and diversity analysis in Citrus. Hort Sci, 1995, 30: 1063–1067
Chen S L, Song J Y, Yao H, et al. Strategy and key technique of identification of Chinese herbal medicine using DNA barcoding, Chin J Nat Med, 2009, 7: 322–327 10.3724/SP.J.1009.2009.00322, 1:CAS:528:DC%2BC3cXhtFKktbc%3D
Ning S P, Yan H Y, Hao G, et al. Current advances of DNA barcoding study in plants. Biodiversity Science, 2008, 16: 417–425 1:CAS:528:DC%2BD1MXotVWktLg%3D
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Luo, K., Chen, S., Chen, K. et al. Assessment of candidate plant DNA barcodes using the Rutaceae family. Sci. China Life Sci. 53, 701–708 (2010). https://doi.org/10.1007/s11427-010-4009-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-010-4009-1