Abstract
SARS-CoV M gene fragment was cloned and expressed as a recombinant protein fused with a V5 tag at the C-terminus in Vero E6 cells. In addition to un-glycosylated and glycosylated proteins, one product with smaller size initiated in-frame from the third Met residues probably through ribosomal re-initiation was also detected. Translation initiated in-frame from the third Met is unusual since the sequence around the first Met of SARS-CoV M protein contains the optimal consensus Kozak sequence. The function of this smaller translated product awaits further investigation. Similar to other N-glycosylated proteins, glycosylation of SARS-CoV M protein was occurred co-translationally in the presence of microsomes. The SARS-CoV M protein is predicted as a triple-spanning membrane protein lack of a conventional signal peptide. The second and third trans-membrane regions (a.a. 46–68 and 78–100) are predicted to be the primary type helices, which will be able to penetrate into membrane by themselves, while the first trans-membrane region (a.a. 14–36) is predicted to be the secondary type helix, which is considered to be stabilized by the interaction with other trans-membrane segments. As expected, the second and third trans-membrane regions were able to insert a cytoplasmic protein into the endoplasmic reticulum membrane more efficiently than the first one. These results should be important for the study of SARS-CoV morphogenesis.
Similar content being viewed by others
Introduction
Severe acute respiratory syndrome (SARS), a new infectious disease typically associated with fever, shortness of breath, cough, and pneumonia, first emerged in southern China in November, 2002. Within months of outbreak, SARS had spread globally, affecting over 8,000 patients in 29 countries with 774 fatalities [1]. The aetiology of SARS is associated with a newly discovered coronavirus, SARS-associated coronavirus (SARS-CoV) [2]. Subsequent studies have indicated that the SARS coronavirus is of animal origin [3], and its precursor is still present in animal populations within the region. The live-animal markets in southern China may have provided the animal–human interphase that allowed this precursor virus to adopt to human–human transmission. Therefore, even in a situation of no new infections, SARS remains a major health hazard, as new epidemics may arise. Further basic and clinical research is required to control the disease.
By May 9, 2003, 14 genomes of SARS coronavirus had been sequenced [4–8]. SARS coronavirus is phylogenetically distinct, and only distantly related to the other coronavirus clades [9, 10]. Coronaviruses are exceptionally large RNA viruses and employ complex regulatory mechanisms to express their genomes [11]. The genome structure, gene expression pattern and protein profiles of SARS-CoV are similar to those of other coronaviruses. Nine SARS-CoV specific mRNAs were synthesized in virus-infected cells [12]. These RNAs were predicted to encode two large replicative polyprotein (pp1a and pp1ab), four structural proteins (spike, membrane, envelope, and nucleocapsid proteins), and other auxiliary proteins. In the cases of other coronaviruses, the four structural proteins (S, M, E, and N) play roles in virion morphogenesis [13]. N binds to viral RNA to form nucleocapsid. Co-expression of M and E proteins together can form virus-like particles [14]. Interactions between the M and E proteins and nucleocapsids result in virus budding through cellular membrane. Through the interaction with M protein, S protein is incorporated into the viral envelope and the mature virions are released from the cells. It has also been demonstrated that virus-like particles of SARS-CoV could be formed by expressing M and E proteins in insect cells [15]. Therefore, the M protein plays a crucial role in coronavirus assembly. Characterization of SARS-CoV M protein will shed light on the studies of the morphogenesis of SARS-CoV. Here, we report the expression and membrane integration of recombinant SARS-CoV M protein in Vero E6 cells.
Materials and methods
Plasmid construction
The construction of the plasmid expressing full-length membrane protein plus V5 and His tag encoded from pcDNA3.1/V5-His A vector sequence (M-V5-His) was described previously [16].
To mutate amino acid 4 of SARS-CoV M protein from Asn to Asp, PCR primers (5′CGGAATTCATGGCAGACGACGGTACTATTACCG3′ and 5′TGCTCTAGACTGTACTAGCAAAGCAAT3′) were used to amplify the gene fragment. After PCR, the DNA fragment was digested by restriction enzymes (EcoRI/XbaI) and cloned into pcDNA3.1-V5-His A (linearized by EcoRI/XbaI) expression vector. This expression plasmid encodes full-length SARS-CoV M protein with mutation in amino acid 4 from Asn to Asp.
To mutate amino acid 83 of SARS-CoV M protein from Met to Leu, PCR primers (5′CCGGAATTCATGGCAGACAACGGTACTA3′ and 5′AATACAAGCCAGTGCAATCGCAATC3′) were used to amplify the 5′-end of the membrane gene fragment while PCR primers (5′GCGATTGCACTGGCTTGTATTGTAG3′ and 5′TGCTCTAGACTGTACTAGCAAAGCAAT3′) were used to amplify the 3′-end fragment. These two DNA fragments were linked together by PCR using primers (5′CCGGAATTCATGGCAGACAACGGTACTA3′ and 5′TGCTCTAGACTGTACTAGCAAAGCAAT3′). After PCR, the DNA fragment was digested by restriction enzymes (EcoRI/XbaI) and cloned into pcDNA3.1-V5-His A (linearized by EcoRI/XbaI) expression vector. This expression plasmid encodes full-length SARS-CoV M protein with mutation in amino acid 83 from Met to Leu. Similar approach was used to mutate amino acid 32 of SARS-CoV M protein from Met to Leu.
To clone entire SARS-CoV M gene fragment including 5′-untranslated region (starting from nt. 26361 of GI: 30027610), PCR primers (5′CCGGAATTCTGGAACTTTAACATT3′ and 5′TGCTCTAGACTGTACTAGCAAAGCAAT3′) were used to amplify the gene fragment. After PCR, the DNA fragment was digested by restriction enzymes (EcoRI/XbaI) and cloned into pcDNA3.1-V5-His A (linearized by EcoRI/XbaI) expression vector. This expression plasmid contains the full-length SARS-CoV M coding sequence and 22 nucleotides of the 5′-untranslated region.
To clone the DNA fragment with the first 115 amino acids of HCV core protein into pcDNA3.1-V5-His A, the expression plasmid (M-C-V5-His) encoding a fusion protein with full-length SARS-CoV M protein and the first 115 amino acids of HCV core protein [16] was digested with BamHI to remove the DNA fragment with full-length SARS-CoV M protein and self-ligated.
To clone the DNA fragment with the first 115 amino acids of HCV core protein and the first 50 amino acids of SARS-CoV M protein, the expression plasmid (C-M-V5-His) encoding a fusion protein with the first 115 amino acids of HCV core protein and full-length SARS-CoV M protein [16] was used as template and primers (5′CGGAATTCAGGTCTCGTAGACCG3′ and 5′TGCTCTAGAAAGCTTTATTATGTA3′) were used to perform PCR and amplify the gene fragment. After PCR, the DNA fragment was digested by restriction enzymes (EcoRI/XbaI) and cloned into pcDNA3.1-V5-His A (linearized by EcoRI/XbaI) expression vector. This expression plasmid encodes a fusion protein consisting of the first 115 amino acids of HCV core protein, plus the first 50 amino acids of SARS-CoV M protein, followed by V5 and His tag.
To link the first 115 amino acids of HCV core protein with a.a. 46–68 of SARS-CoV M protein, PCR primers (HCV-1 and 5′CGCGGATCCCCTACGCCGGGGGTCTGT3′) were used to amplify the first 115 amino acids of the core gene fragment while PCR primers (5′CGCGGATCCTACATAATAAAGCTTGT3′ and 5′TGCTCTAGAAACAGCAAGCACAAAAC3′) were used to amplify the DNA fragment encoding a.a. 46–68 of SARS-CoV M protein. After PCR reaction, these two DNA fragments were digested by restriction enzymes (EcoRI/BamHI and BamHI/XbaI separately) and cloned into pcDNA3.1-V5-HisA (Invitrogen, USA) expression vector (linearized by EcoRI/XbaI). To link the first 115 amino acids of HCV core protein with a.a. 78–100 of SARS-CoV M protein, a similar approach was performed except using primers (5′CGCGGATCCGGGATTGCGATTGCAAT3′ and 5′TGCTCTAGACCTGAAGGAAGCAACGA3′) to amplify the DNA fragment of SARS-CoV M protein a.a. 78–100.
All the expression plasmids were verified by sequencing.
Protein expression in Vero E6 cells
The Vero E6 cells were maintained in RPMI 1640 medium containing 10% fetal calf serum, 1% Glutamine (200 mM, Biological Industries, USA), and 100 μg/ml penicillin/streptomycin (Gibco BRL, USA). 2.5−2.7 × 105 cells were plated in the 35-mm dish. After an overnight incubation, cells were infected with a recombinant vaccinia virus carrying the T7 phage RNA polymerase gene [17]. Two hours after infection, cells were transfected with 0.4 μg plasmid DNA by using Effectene transfection reagent (Qiagen, Germany). 21 h after transfection, recombinant proteins in the cells were analyzed. The mRNAs transcribed from either CMV promoter or T7 promoter in this expression system use the same AUG to initiate translation.
Western blotting analysis
For Western blotting analysis, cells were dissolved in sample preparation buffers after washed by PBS twice. In our previous study [16], SARS-CoV M protein could not be detected in SDS-PAGE in regular boiling treatment. Therefore, non-heated treatments [18] were used in antigen preparations (sample buffer containing 50 mM Tris–HCl (pH 6.8), 100 mM dithiothreitol, 2% SDS, 0.1 % bromophenol blue, 10% glycerol; no boiling treatment) to detect the expression of SARS-CoV M protein. 4.5% (acrylamide percentage) gel was used as the stacking gel and 12% gel as the separating gel in this study. When proteins with smaller size were analyzed (e.g. deletion mutants of membrane protein), 15% gel was used as the separating gel. After electrophoresis, the SDS-PAGE gel was transferred to PVDF paper (Pall Corporation, USA). All procedures were then carried out at room temperature following our previous procedures [19]. The proportion of different forms of M protein (in Figs. 1a, b, 2a) was quantified using software “Quantity one” (Biorad, USA).
SARS-CoV M protein is N-glycosylated. (a) Vero E6 cells were either mock transfected (lane 1) or transfected with recombinant M protein (lanes 2–4). After transfection, cell lysates were further treated with buffer (lane 2), N-glycosidase F (lane 3) or endoglycosidase H (lane 4). After treatment, cell lysates were analyzed by Western blotting using anti-V5 mAb. (b) Vero E6 cells were mock transfected (lane 1), and transfected with either recombinant M protein (lane 2), or recombinant M protein with amino acid residue 4 changed from Asn to Asp (lane 3). After transfection, cell lysates were analyzed by Western blotting using anti-V5 mAb. Glycosylated proteins were marked by thick arrow while un-glycosylated proteins were marked by thin arrow. Two smaller proteins marked by thick or thin line were also repeatedly detected
Two smaller products were produced from SARS-CoV M gene in addition to the full-length M protein. (a) Vero E6 cells were mock transfected (lane 1), or transfected with recombinant M protein (lane 2), recombinant M protein with Met 32 changed to Leu (lane 3), recombinant M protein with Met 83 changed to Leu (lane 4), recombinant M protein with both Met 32 and Met 83 changed to Leu (lane 5). After transfection, cell lysates were analyzed by Western blotting using anti-V5 mAb. One translated product initiated from Met 83 was missing (lanes 4 and 5, marked by thin line) and one additional smaller product was produced (lanes 4 and 5, marked by double arrow)
Treatment of Endoglycosidase H or N-Glycosidase F
Vero E6 cells (1 × 106) were infected with recombinant vaccinia virus carrying the T7 phage RNA polymerase gene [17] for 2 h, then transfected with the plasmid expressing full-length membrane protein plus V5 and His tag. Cells were harvested 21 h after transfection, and lysed in RIPA buffer (150 mM NaCl, 1% NP40, 0.5% deoxychloic acid, 0.1% SDS, 50 mM Tris, pH7.5). After centrifugation for 5 min at full speed in microcentrifuge, supernatant was incubated with mouse anti-His monoclonal antibody (Santa Cruz, USA) at 4°C for overnight with shaking. The antigen-antibody complex was pulled down by pansorbin (Merck, USA). For N-glycosidase F (Roche, Germany) digestion, the immunoprecipitated pellet was boiled in 2 μl of 1% SDS for 2 min. Then, 18 μl of 20 mM NaHPO4 and 50 mM EDTA was added and boiled for another 2 min. After cooling, the enzyme (1U) was added and incubated at 37°C for overnight. For endoglycosidase H (Roche, Germany) digestion, the immunoprecipitated pellet was boiled with digestion buffer (0.1% SDS, 50 mM NaHPO4) for 4 min. After cooling, the enzyme (5 mU) was added and incubated at 37°C for overnight. After enzyme digestion, the samples were analyzed with SDS-PAGE and Western blotting.
In vitro transcription/translation
Commercially available TnT system (Promega, USA) was used to perform in vitro transcription/translation assay. The experiments were conducted following the manufacturer’s instructions. About 2 μg of DNA was used in a 50 μl reaction. Microsome was added to study the glycosylation. To stop the translation reaction, CaCl2 in a final concentration of 5 mM was added in the reaction mixture [20].
Confocal analysis
About 2.5 × 105 cells were seeded in 35 mm culture dishes. After overnight incubation, cells were transfected with 0.4 μg plasmid by using Effectene trasfection kit (Qiagen, Germany). 48 h after transfection, recombinant proteins in the cells were analyzed. Cells were fixed by acetone/methanol (1:1), at 0°C for 10 min. Fixed cells were washed with incubation buffer (0.05% NaN3, 0.02% saponin, 1% skim milk in PBS) twice for 5 min each time, then incubated with mouse anti-V5 monoclonal antibody (Invitrogen, USA), which had been diluted 200 fold, at 37°C for 30 min. Samples were washed with PBS three times (5 min each time at room temperature), then incubated with FITC-conjugated goat anti-mouse IgG antibody in 20× dilution at 37°C for 30 min. Again, samples were washed with PBS three times (5–10 min each time at room temperature). Cells were co-transfected with dsRED-ER plasmid (BD, USA) when ER localization needs to be defined. DAPI (Merck, Germany) was used to stain DNA as the localization of nucleus.
Results
SARS-CoV M protein is an N-glycosylated protein
Full-length SARS-CoV M gene fragment was cloned and expressed as a recombinant protein (221 a.a.) with a C-terminal V5-His tag (29 a.a.) in Vero E6 cells (Fig. 1a). In addition to the protein with the expected molecular weight (27.5 kDa, marked by thin arrow), several protein products of different sizes were also detected (lane 2 in Fig. 1a). One such protein with larger size (marked by thick arrow) was the glycosylated M protein since this band disappeared after treatment with either N-glycosidase F (lane 3 in Fig. 1a) or endoglycosidase H (lane 4 in Fig. 1a). The Asn-Gly-Thr (a.a. 4–6) at the N-terminal of SARS-CoV M protein is a defined consensus sequence (Asn-X-(Ser/Thr)) for N-linked glycosylation [21]. This is verified by the observation that production of the glycosylated M protein was blocked when amino acid residue 4 was changed from Asn (lane 2 in Fig. 1b) to Asp (lane 3 in Fig. 1b).
One smaller product translated in-frame from the third Met was detected
In addition to the glycosylated and un-glycosylated SARS-CoV M proteins, two smaller protein products (marked by thick line and thin line, respectively) were also detected when M gene was expressed in Vero E6 cells (Fig. 1a and b). The size of these two smaller proteins corresponds to the products translated in-frame from the second (amino acid 32) and the third (amino acid 83) Met residues. This hypothesis was supported by the expression of several M deletion mutants (Supplement Fig. 1). The protein assumed to be translated in-frame from the third Met disappeared when amino acid 83 was mutated from Met to Leu (lanes 4 and 5 in Fig. 2). Instead, one additional protein (marked by double arrow) corresponding to the product initiated in-frame from the fourth Met (amino acid 90) was detected when this mutant M protein was expressed. However, the protein assumed to be translated in-frame from the second Met still exists when amino acid 32 was mutated from Met to Leu (lanes 3 and 5 in Fig. 2). Thus, this protein should be a degraded product. The protein translated in-frame from the third Met (amino acid 83) could still be detected when the authentic 5′-untranslated region of SARS-CoV M gene was included in the expression vector (lane 3 in supplement Fig. 2). We have also quantified the proportion of these two different forms of M protein (in Figs. 1a, b and 2). Comparing with the unglycosylated form of M protein (as “100%”), the expression level of the protein with a.a. 32–221 is 40% ± 19% while the expression of the protein with a.a. 83–221 is 20% ± 6%.
To rule out the possibility that the read-out products of SARS-CoV M gene were generated by the C-terminus V5-His tag, we carried out experiments with the same tag fused to the C-terminus of a different gene (pyruvate kinase, supplement Fig. 3) No other bands are detected below molecular weight 67 kDa (574 a.a. of pyruvate kinase plus 29 a.a. of tag = 603 a.a.), although pyruvate kinase does contain several internal in-frame AUGs (e.g., a.a. 65, a.a. 107, a.a. 112, etc.). The results indicated that V5-His tag would not facilitate the processing of read-through products.
Glycosylation of SARS-CoV M protein occurs co-translationally
To determine the N-glycosylation of SARS-CoV M protein is co-translational or post-translational event, SARS-CoV M protein was in vitro translated in the reticulocyte lysate. N-glycosylated product was detected only when microsome was added before but not after of translation (Fig. 3). Thus, glycosylation of SARS-CoV M protein should occur co-translationally.
SARS-CoV M protein was in vitro translated in the reticulocyte lysate. Lane 1, no DNA was added in the lysate; Lanes 2 and 3, only the plasmid expressing SARS-CoV M protein was added; Lane 4, After the translation of SARS-CoV M protein, CaCl2 was added to stop the translation reaction and then, add the microsome. Lane 5, Micrsome and the plasmid expressing SARS-CoV M protein were added in the lysate together. After the reaction, 1 μl of the translation mixture (except 2 μl in lane 3) was used to analyze the products. Un-glycosylated M protein was marked by thin arrow while glycosylated M protein was marked by thick arrow
The second and third trans-membrane regions could insert a cytoplasmic protein into the endoplasmic reticulum membrane more efficiently than the first one
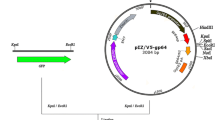
SARS-CoV M protein will go to E.R. membrane though lack of a conventional signal peptide and predominately localizes in the Golgi complex [22]. There are three predicted trans-membrane domains in the SARS-CoV M protein: a.a. 14–36, a.a. 46–68, and a.a. 78–100 (Supplement Fig. 4). The second and third trans-membrane regions (a.a. 46–68 and 78–100) are predicted to be the primary type helices while the first trans-membrane region (a.a. 14–36) is predicted to be the secondary type helix (Supplement Fig. 4). The SARS-CoV M protein mutants with deletion of anyone trans-membrane region could still enter into E.R. (data not shown). To determine which trans-membrane domain is responsible for the integration of SARS-CoV M protein into E.R., each of these three trans-membrane domains (Fig. 4) was linked with a cytoplasmic protein (a.a. 1–115 of hepatitis C virus core protein) (Fig. 5) and the recombinant fused protein was expressed in Vero E6 cells. To quantitate the average percentage of co-localization from 50 cells for each test, Image J (NIH web) program was used. The average R values of pDsRed-ER + C115, pDsRed-ER + CM50, pDsRed-ER + CM46–68 and pDsRed-ER + CM78–100 are 0.3379, 0.6632, 0.8624, and 0.8442, respectively. Thus, the second (Fig. 6) and third (Fig. 7) trans-membrane regions could insert a cytoplasmic protein into the endoplasmic reticulum membrane more efficiently than the first one (Fig. 8).
Subcellular localization of recombinant fusion protein C115-V5. The dsRED-ER plasmid was used to define the E.R. while DAPI was used to mark the localization of nucleus. Recombinant fusion protein C115-V5 was stained with FITC-conjugated antibody. Photos are Vero cells transfected with dsRED-ER and/or C115-V5 plasmid. No co-localization of C115-V5 and dsRED-ER was observed
Subcellular localization of recombinant fusion protein C-M(46–68)-V5. The dsRED-ER plasmid was used to define the E.R. while DAPI was used to mark the localization of nucleus. Recombinant fusion protein C-M(46–68)-V5 was stained with FITC-conjugated antibody. Photos of the upper panels are Vero cells transfected with dsRED-ER or C-M(46–68)-V5 plasmid while those of lower panels are Vero cells co-transfected with dsRED-ER and C-M(46–68)-V5 plasmids. Orange color (merge red color with green color) represents the localization of the recombinant protein in the E.R.
Subcellular localization of recombinant fusion protein C-M(78–100)-V5. The dsRED-ER plasmid was used to define the E.R. while DAPI was used to mark the localization of nucleus. Recombinant fusion protein C-M(78–100)-V5 was stained with FITC-conjugated antibody. Photos of the upper panels are Vero cells transfected with dsRED-ER or C-M(78–100)-V5 plasmid while those of lower panels are Vero cells co-transfected with dsRED-ER and C-M(78–100)-V5 plasmids. Orange color (merge red color with green color) represents the localization of the recombinant protein in the E.R.
Subcellular localization of recombinant fusion protein C-M(1–50)-V5. The dsRED-ER plasmid was used to define the E.R. while DAPI was used to mark the localization of nucleus. Recombinant fusion protein C-M(1–50)-V5 was stained with FITC-conjugated antibody. Photos of the upper panels are Vero cells transfected with dsRED-ER or C-M(1–50)-V5 plasmid while those of lower panels are Vero cells co-transfected with dsRED-ER and C-M(1–50)-V5 plasmids. No co-localization of C-M(1–50)-V5 and dsRED-ER was observed
Discussion
In this study, SARS-CoV M gene fragment was cloned and expressed as a recombinant protein fused with a C-terminal V5 tag in Vero E6 cells (Fig. 1). One protein larger than the expected size was the product of N-glycosylation since its size was reduced to the expected molecular weight when treated with either endo-F or endo-H (Fig. 1a), and its production was blocked when Asn 4 in the predicted glycosylation site (Asn-X-(Ser/Thr)) was mutated to Asp (Fig. 1b). In other well studied coronaviruses, the M protein is involved in the assembly and budding of virions together with the E protein [14]. Moreover, the M proteins of other coronaviruses also contain highly conserved glycosylation sequences [23], and their glycosylation may be related to the interaction between virus and host. Whether N-glycosylation of SARS-CoV M protein is related to the interaction between virus and host awaits further investigation. The N-glycosylation of SARS-CoV M protein should occur in the endoplasmic reticulum rather than Golgi complex since both endo-F and endo-H can remove the glycosylated moiety [24].
Two other expressed SARS-CoV M protein products with smaller size than the full-length one were also detected in Vero E6 cells (Figs. 1 and 2). These two smaller proteins seem to be the translated products initiated in-frame from second (amino acid 32) and third (amino acid 83) Met residues. It was verified with a Met 83 but not Met 32 mutated SARS-CoV M protein, which blocked the production of the expected product (Fig. 2). Translation initiated in-frame from the third Met could be due to ribosomal leaky scanning, re-initiation, and even internal initiation [25, 26]. It is least likely due to internal initiation since there is no conventional IRES (internal ribosomal entry site) in the SARS-CoV M gene sequence. It is less likely through ribosomal leaky scanning since neither the second nor the fourth in-frame AUG was used as the initiation codon (Figs. 1 and 2). It is probably through ribosomal re-initiation. This explains why neither the second nor the fourth in-frame AUG was used as the initiation codon (too close to the first or third in-frame AUG). And, this also explains why the protein initiated in-frame from the fourth Met was detected when the third in-frame AUG was mutated (Fig. 2). Production of this smaller product could be due to the replacement of authentic 5′-untraslated region (5′CUUAUCAUGG3′) of M gene with vector cloning sequence (5′GAAUUCAUGG3′). To rule out this possibility, expression plasmid with authentic 5′-untraslated region of SARS-CoV M gene was cloned and expressed. The protein product initiated from Met 83 was also detected in this expression construction (Supplement Fig. 2, lane 3). Translation initiated in-frame from the third Met is unusual since the sequence around the first Met (5′CUUAUCAUGG3′) of SARS-CoV M protein is the optimal consensus Kozak sequence (5′GCCA/GCCAUGG3′). Moreover, there is one out-of-frame AUG (5′GGAACAAUGG3′) with consensus Kozak sequence between the first and the second in-frame Met (GI: 30027610). Production of these smaller SARS-CoV M proteins should not be an artifact of this expression system, since another protein (pyruvate kinase) over-expressed using this system (Supplement Fig. 3) did not generate smaller products initiated from downstream in-frame AUGs. The function of this smaller translated product awaits further investigation.
Similar to other N-glycosylated proteins [24], the glycosylation of SARS-CoV M protein is occurred co-translationally but not post-translationally (Fig. 3). The M protein of mouse hepatitis virus strain A59 is also a triple-spanning membrane protein. Which of these three hydrophobic domains in A59 M protein is the insertion and anchor signal for the E.R.-integration of this protein is not conclusive [27–29]. The second and third trans-membrane regions (a.a. 46–68 and 78–100) are predicted to be the primary type helices, which will be able to penetrate into membrane by themselves, while the first trans-membrane region (a.a. 14–36) is predicted to be the secondary type helix, which is considered to be stabilized by the interaction with other trans-membrane segments [30]. To determine which trans-membrane domain is responsible for the integration of SARS-CoV M protein into E.R., each of these three trans-membrane domains was linked with a.a. 1–115 of hepatitis C virus core protein, which is suspected to be a nuclear protein since it contains the nuclear localization signal [31], and the recombinant fused protein was expressed in Vero E6 cells. Unexpectedly, the protein encoding a.a. 1–115 of hepatitis C virus core protein with V5 tag localizes in the cytoplasm (Fig. 5) but not the nucleus maybe due to the core proteins derived from different isolates. Just like the prediction, the second and third trans-membrane regions were able to insert a cytoplasmic protein into the endoplasmic reticulum membrane more efficiently than the first one (Figs. 6–8). These results suggest that the second and/or third trans-membrane regions are more likely than the first one to target SARS-CoV M protein into E.R.
In summary, when SARS-CoV M gene was expressed in Vero E6 cells, in addition to the full-length un-glycosylated M protein, glycosylated M protein and one smaller product initiated in-fame from the third Met was also detected. N-glycosylation of SARS-CoV M protein is a co-translational event. The second and third trans-membrane regions were able to insert a cytoplasmic protein into the endoplasmic reticulum membrane more efficiently than the first one. These findings should be important for the morphogenesis of SARS-CoV.
References
Poon LL, Guan Y, Nicholls JM, Yuen KY, Peiris JS (2004) The aetiology, origins, and diagnosis of severe acute respiratory syndrome. Lancet Infect Dis 4:663-671
Donnelly CA, Fisher MC, Fraser C, Ghani AC, Riley S, Ferguson NM, Anderson RM (2004) Epidemiological and genetic analysis of severe acute respiratory syndrome. Lancet Infect Dis 4:672–683
Enserink M (2003) Infectious diseases. Clues to the animal origins of SARS. Science 300:1351
Drosten C, Gunther S, Preiser W, van der Werf S, Brodt HR, Becker S, Rabenau H, Panning M, Kolesnikova L, Fouchier RA, Berger A, Burguiere AM, Cinatl J, Eickmann M, Escriou N, Grywna K, Kramme S, Manuguerra JC, Muller S, Rickerts V, Sturmer M, Vieth S, Klenk HD, Osterhaus AD, Schmitz H, Doerr HW (2003) Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N Engl J Med 348:1967–1976
Kuiken T, Fouchier R, Schutten M, Rimmelzwaan GF, van Amerongen G, van Riel D, Laman JD, de Jong T, van Doornum G, Lim W, Ling AE, Chan PK, Tam JS, Zambon MC, Gopal R, Drosten C, van der Werf S, Escriou N, Manuguerra JC, Stohr K, Peiris JS, Osterhaus AD (2003) Newly discovered coronavirus as the primary cause of severe acute respiratory syndrome. Lancet 362:263–270
Marra MA, Jones S, Astell CR, Holt RA, Brooks-Wilson A, Butterfield YS, Khattra J, Asano JK, Barber SA, Chan SY, Cloutier A, Coughlin SM, Freeman D, Girn N, Griffith OL, Leach SR, Mayo M, McDonald H, Montgomery SB, Pandoh PK, Petrescu AS, Robertson AG, Schein JE, Siddiqui A, Smailus DE, Stott JM, Yang GS, Plummer F, Andonov A, Artsob H, Bastien N, Bernard K, Booth TF, Bowness D, Czub M, Drebot M, Fernando L, Flick R, Garbutt M, Gray M, Grolla A, Jones S, Feldmann H, Meyers A, Kabani A, Li Y, Normand S, Stroher U, Tipples GA, Tyler S, Vogrig R, Ward D, Watson B, Brunham RC, Krajden M, Petric M, Skowronski DM, Upton C, Roper RL (2003) The genome sequence of the SARS-associated coronavirus. Science 300:1399–1404
Rota PA, Oberste M, Monroe SS, Nix WA, Campagnoli R, Icenogle JP, Penaranda S, Bankamp B, Maher K, Chen MH, Tong S, Tamin A, Lowe L, Frace M, DeRisi JL, Chen Q, Wang D, Erdman DD, Peret TC, Burns C, Ksiazek TG, Rollin PE, Sanchez A, Liffick S, Holloway B, Limor J, McCaustland K, Olsen-Rasmussen M, Fouchier R, Gunther S, Osterhaus AD, Drosten C, Pallansch MA, Anderson LJ, Bellini WJ (2003) Characterization of a novel coronavirus associated with severe acute respiratory syndrome. Science 300:1394–1399
Woo PC, Lau SK, Chu CM, Chan KH, Tsoi HW, Huang Y, Wong BH, Poon RW, Cai JJ, Luk WK, Poon LL, Wong SS, Guan Y, Peiris JS, Yuen KY (2005) Characterization and complete genome sequence of a novel coronavirus, coronavirus HKU1, from patients with pneumonia. J Virol 79:884–895
Holmes KV, Enjuanes L (2003) Virology. The SARS coronavirus: a postgenomic era. Science 300:1377–1388
Lai M (2003) SARS virus: the beginning of the unraveling of a new coronavirus. J Biomed Sci 10:664–675
Holmes K, Lai MM (1996) Coronaviridae: the viruses and their replication. In: Fields BN, Knipe DM, Howley PM (eds) Fields virology, vol 1. Lippincott-Raven Publishers, Philadelphia, pp 1075–1093
Thiel V, Ivanov K, Putics A, Hertzig T, Schelle B, Bayer S, Weissbrich B, Snijder EJ, Rabenau H, Doerr HW, Gorbalenya AE, Ziebuhr J (2003) Mechanisms and enzymes involved in SARS coronavirus genome expression. J Gen Virol 84:2305–2315
Nguyen VP, Hogue B (1998) Coronavirus envelope glycoprotein assembly complexes. Adv Exp Med Biol 440:361–365
Vennema H, Godeke G, Rossen JW, Voorhout WF, Horzinek MC, Opstelten DJ, Rottier PJ (1996) Nucleocapsid-independent assembly of coronavirus-like particles by co-expression of viral envelope protein genes. EMBO J 15:2020–2028
Ho Y, Lin PH, Liu CY, Lee SP, Chao YC (2004) Assembly of human severe acute respiratory syndrome coronavirus-like particles. Biochem Biophys Res Commun 318:833–838
Lee YN, Chen LK, Ma HC, Yang HH, Li HP, Lo SY (2005) Thermal aggregation of SARS-CoV membrane protein. J Virol Methods 129:152–161
Fuerst TR, Niles EG, Studier FW, Moss B (1986) Eukaryotic transient-expression system based on recombinant vaccinia virus that synthesizes bacteriophage T7 RNA polymerase. Proc Natl Acad Sci USA 83:8122–8126
Lin YL, Liao CL, Chen LK, Yeh CT, Liu CI, Ma SH, Huang YY, Huang YL, Kao CL, King CC (1998) Study of Dengue virus infection in SCID mice engrafted with human K562 cells. J Virol 72:9729–9737
Ma HC, Ke CH, Hsieh TY, Lo SY (2002) The first hydrophobic domain of the hepatitis C virus E1 protein is important for interaction with the capsid protein. J Gen Virol 83:3085–3092
Yamaga AK, Ou JH (2002) Membrane topology of the hepatitis C virus NS2 protein. J Biol Chem 277:33228–33234
Hu Y, Wen J, Tang L, Zhang H, Zhang X, Li Y, Wang J, Han Y, Li G, Shi J, Tian X, Jiang F, Zhao X, Liu S, Zeng C, Yang H (2003) The M protein of SARS-CoV: basic structural and immunological properties. Genomics Proteomics Bioinformatics 1:118–130
Nal B, Chan C, Kien F, Siu L, Tse J, Chu K, Kam J, Staropoli I, Crescenzo-Chaigne B, Escriou N, van der Werf S, Yuen KY, Altmeyer R (2005) Differential maturation and subcellular localization of severe acute respiratory syndrome coronavirus surface proteins S, M and E. J Gen Virol 86:1423–1434
de Haan CA, Roestenberg P, de Wit M, de Vries AA, Nilsson T, Vennema H, Rottier PJ (1998) Structural requirements for O-glycosylation of the mouse hepatitis virus membrane protein. J Biol Chem 273:29905–29914
Lewin B (2004) GENES VIII. Pearson Prentice Hall, Upper Saddle River
Kozak M (1999) Initiation of translation in prokaryotes and eukaryotes. Gene 234:187–208
Kozak M (2002) Pushing the limits of the scanning mechanism for initiation of translation. Gene 299:1–34
Locker JK, Rose JK, Horzinek MC, Rottier PJ (1992) Membrane assembly of the triple-spanning coronavirus M protein. Individual transmembrane domains show preferred orientation J Biol Chem 267:21911–21918
Mayer T, Tamura T, Falk M, Niemann H (1988) Membrane integration and intracellular transport of the coronavirus glycoprotein E1, a class III membrane glycoprotein. J Biol Chem 263:14956–14963
Rottier P, Armstrong J, Meyer DI (1985) Signal recognition particle-dependent insertion of coronavirus E1, an intracellular membrane glycoprotein. J Biol Chem 260:4648–4652
Hirokawa T, Boon-Chieng S, Mitaku S (1998) SOSUI: classification and secondary structure prediction system for membrane proteins. Bioinformatics 14:378–379
Chang SC, Yen JH, Kang HY, Jang MH, Chang MF (1994) Nuclear localization signals in the core protein of hepatitis C virus. Biochem Biophys Res Commun 205:1284–1290
Acknowledgements
This work has been supported by grants from Tzu Chi University (TCMRC 93017 and TCMRC-C95002–01) and from National Science Council of Taiwan (NSC 95-3112-B-320-001) to Dr. Shih-Yen Lo.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11373_2008_9235_MOESM1_ESM.tif
Two proteins suspected to be translated in-frame from the second and third Mets of SARS-CoV M protein were detected. Vero E6 cells either mock transfected (lane 1) or transfected with recombinant M protein (lane 2), recombinant M protein deleting amino acids from 47 to 60 (lane 3), recombinant M protein deleting amino acids from 61 to 90 (lane 4), recombinant M protein deleting amino acids from 91 to 100 (lane 5). After transfection, cell lysates were analyzed by Western blotting using anti-V5 mAb. Glycosylated proteins were marked by thick arrow while un-glycosylated proteins were marked by thin arrow. The protein marked by thick line is supposed to be translated from the second in-frame Met (amino acid 32) while the protein marked by thin line (and also the protein marked by the dotted line) is supposed to be translated from the third in-frame Met (amino acid 83) (TIF 37 kb)
11373_2008_9235_MOESM2_ESM.tif
SARS-CoV M gene with its authentic 5′-untranslated region can still generate smaller products in addition to the full-length M protein. Vero E6 cells were mock transfected (lane 1), or transfected with recombinant M protein without its authentic 5′-untranslated region (lane 2), recombinant M protein containing its authentic 5′-untranslated region (lane 3). After transfection, cell lysates were analyzed by Western blotting using anti-V5 mAb (TIF 126 kb)
11373_2008_9235_MOESM3_ESM.tif
No read-through products were detected when pyruvate kinase gene was expressed. Vero E6 cells were mock transfected (lane 1), or transfected with recombinant pyruvate kinase protein (lanes 2 and 3, repeat twice). After transfection, cell lysates were analyzed by Western blotting using anti-V5 mAb. The protein marked by the arrow represents the fusion protein with full-length pyruvate kinase plus V5-His tag (TIF 155 kb)
11373_2008_9235_MOESM4_ESM.tif
Topology of SARS-CoV M protein in prediction. Program (http://www.bp.nuap.nagoya-u.ac.jp/sosui/) was used to do the prediction (TIF 132 kb)
Rights and permissions
About this article
Cite this article
Ma, HC., Fang, CP., Hsieh, YC. et al. Expression and membrane integration of SARS-CoV M protein. J Biomed Sci 15, 301–310 (2008). https://doi.org/10.1007/s11373-008-9235-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11373-008-9235-1