Abstract
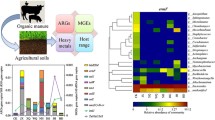
Owing to the similar mechanisms of antibiotic and metal resistance, there is a growing concern that metal contamination may select for antibiotic resistance genes (ARGs) in the environment. Here, we constructed short-term laboratory microcosms to investigate the dynamics of a wide range of ARGs and two copper (Cu) resistance genes in an agricultural soil amended with a gradient of Cu concentrations (0~1000 mg kg−1). Mobile genetic elements (MGEs) were also quantified as a proxy for the horizontal gene transfer potential of ARGs. We detected 126 unique ARGs across all the soil samples using the high-capacity quantitative PCR array, and multidrug and β-lactam resistance were the most abundant ARG categories. The copper amendments significantly enhanced the absolute and relative abundances of ARGs and MGEs, which gradually increased along the gradient of increasing Cu concentrations. The two Cu resistance genes (copA and pcoR) were highly enriched in low-level Cu treatment (50 and 100 mg kg−1), and their abundances decreased with the increasing Cu concentrations. The level of metal and antibiotic resistance gradually declined over time in all Cu-amended treatments but was still considerably higher in contaminated soils than untreated soils after 56 days’ incubation. Significant associations among ARGs and MGEs were revealed by the network analysis, suggesting the mobility potential of antibiotic resistance in Cu-amended soils. No significant positive correlations were found between ARGs and copper resistance genes, suggesting that these genes are not located in the same bacterial hosts. Taken together, our results provide empirical evidence that short-term copper stress can cause evolution of high-level antibiotic and metal resistance and significantly change the diversity, abundance, and horizontal transfer potential of soil ARGs.






Similar content being viewed by others

References
Amachawadi RG, Scott HM, Alvarado CA, Mainini TR, Vinasco J, Drouillard JS, Nagaraja TG (2013) Occurrence of the transferable copper resistance gene tcrB, among fecal enterococci of U.S. feedlot cattle fed copper supplemented diets. Appl Environ Microbiol 79:4369–4375
Aminov RI (2011) Horizontal gene exchange in environmental microbiota. Front Microbiol 2:158
Ashbolt NHJ, Amezquita A, Backhaus T, Borriello P, Brandt KK, Collignon P (2013) Human health risk assessment (HHRA) for environmental development and transfer of antibiotic resistance. Environ Health Perspect 121:993–1001
Baker-Austin C, Wright MS, Stepanauskas R, McArthur JV (2006) Co-selection of antibiotic and metal resistance. Trends Microbiol 14:176–182
Bastian M, Heymann S, Jacomy M (2009) Gephi: an open source software for exploring and manipulating networks. Icwsm 8:361–362
Berendonk TU, Manaia CM, Merlin C, Fatta-Kassinos D, Cytryn E, Walsh F, Bürgmann H, Sørum H, Norström M, Pons MN, Kreuzinger N, Huovinen P, Stefani S, Schwartz T, Kisand V, Baquero F, Martinez JL (2015) Tackling antibiotic resistance: the environmental framework. Nat Rev Microbiol 13:310–317
Berg J, Tom-Petersen A, Nybroe O (2005) Copper amendment of agricultural soil selects for bacterial antibiotic resistance in the field. Lett Appl Microbiol 40:146–151
Besaury L, Pawlak B, Quillet L (2016) Expression of copper-resistance genes in microbial communities under copper stress and oxic/anoxic conditions. Environ Sci Pollut Res Int 23:4013–4023
Burch TR, Sadowsky MJ, Lapara TM (2013) Aerobic digestion reduces the quantity of antibiotic resistance genes in residual municipal wastewater solids. Front Microbiol 4:17
Chen Q, An X, Li H, Su J, Ma Y, Zhu YG (2016) Long-term field application of sewage sludge increases the abundance of antibiotic resistance genes in soil. Environ Int 92:1–10
Chen Q, An X, Zhu YG, Su JQ, Gillings MR, Ye ZL, Cui L (2017) Application of struvite alters the antibiotic resistome in soil, rhizosphere, and phyllosphere. Environ Sci Technol 51:8149–8157
Forsberg KJ, Reyes A, Wang B, Selleck EM, Sommer MOA, Dantas G (2012) The shared antibiotic resistome of soil bacteria and human pathogen. Science 337:1107–1111
Gillings MR, Gaze WH, Pruden A, Smalla K, Tiedje JM, Zhu YG (2015) Using the class 1 integron-integrase gene as a proxy for anthropogenic pollution. ISME J 9:1269–1279
Gou M, Hu HW, Zhang YJ, Wang JT, Hayden H, Tang YQ, He JZ (2018) Aerobic composting reduces antibiotic resistance genes in cattle manure and the resistome dissemination in agricultural soils. Sci Total Environ 612:1300–1310
Han X-M, Hu H-W, Shi X-Z, Wang J-T, Han L-L, Chen D, He J-Z (2016) Impacts of reclaimed water irrigation on soil antibiotic resistome in urban parks of Victoria, Australia. Environmental Pollution 211:48-57
Hu HW, Han XM, Shi XZ, Wang JT, Han LL, Chen D, He JZ (2016a) Temporal changes of antibiotic resistance genes and bacterial communities in two contrasting soils treated with cattle manure. FEMS Microbiol Ecol 92:fiv196
Hu HW, Wang JT, Li J, Li JJ, Ma YB, Chen D, He JZ (2016b) Field-based evidence for copper contamination induced changes of antibiotic resistance in agricultural soils. Environ Microbiol 18:3896–3909
Hu HW, Wang JT, Li J, Shi XZ, Ma Y, Chen D, He JZ (2017) Long-term nickel contamination increases the occurrence of antibiotic resistance genes in agricultural soils. Environ Sci Technol 51:790–800
Hu HW, Wang JT, Singh BK, Liu YR, Chen YL, Zhang YJ, He JZ (2018) Diversity of herbaceous plants and bacterial communities regulates soil resistome across forest biomes. Environ Microbiol. https://doi.org/10.1111/1462-2920.14248
Ji X, Shen Q, Liu F, Ma J, Xu G, Wang Y, Wu M (2012) Antibiotic resistance gene abundances associated with antibiotics and heavy metals in animal manures and agricultural soils adjacent to feedlots in Shanghai; China. J Hazard Mater 235:178–185
Johnson TA, Stedtfeld RD, Wang Q, Cole JR, Hashsham SA, Looft T, Zhu YG, James MT (2016) Clusters of antibiotic resistance genes enriched together stay together in swine agriculture. mBio 7:e02214–e02215
Knapp CW, McCluskey SM, Singh BK, Campbell CD, Hudson G, Graham DW (2011) Antibiotic resistance gene abundances correlate with metal and geochemical conditions in archived Scottish soils. PLoS One 6:1–6
Li XF, Yin HB, Su JQ (2012) An attempt to quantify Cu-resistant microorganisms in a paddy soil from Jiaxing, China. Pedosphere 22:201–205
Li LG, Xia Y, Zhang T (2017) Co-occurrence of antibiotic and metal resistance genes revealed in complete genome collection. ISME J 11:651–662
Looft T, Johnson TA, Allen HK, Bayles DO, Alt DP, Stedtfeld RD, Sul WJ, Stedtfeld TM, Chai B, Cole JR (2012) In-feed antibiotic effects on the swine intestinal microbiome. Proc Natl Acad Sci U S A 109:1691–1696
Martinez JL (2009) Environmental pollution by antibiotics and by antibiotic resistance determinants. Environ Pollut 157:2893–2902
Medardus JJ, Molla BZ, Nicol M, Morrow WM, Rajala-Schultz PJ, Kazwala R, Gebreyes WA (2014) In-feed use of heavy metal micronutrients in US swine production systems and its role in persistence of multidrug-resistant salmonellae. Appl Environ Microbiol 80:2317–2325
Nesme J, Cecillon S, Delmont TO, Monier JM, Vogel TM, Simont P (2014) Large-scale metagenomic-based study of antibiotic resistance in the environment. Curr Biol 24:1096–1100
Pal C, Bengtsson-Palme J, Kristiansson E, Larsson DJ (2015) Co-occurrence of resistance genes to antibiotics, biocides and metals reveals novel insights into their co-selection potential. BMC Genomics 16:964
Poole K (2017) At the nexus of antibiotics and metals: the impact of Cu and Zn on antibiotic activity and resistance. Trends Microbiol 25:820–832
Pruden A, Pei R, Storteboom H, Carlson KH (2006) Antibiotic resistance genes as emerging contaminants: studies in Northern Colorado. Environ Sci Technol 40:7445–7450
Rizzo L, Manaia C, Merlin C, Schwartz T, Dagot C, Ploy MC, Michael I, Fatta-Kassinos D (2013) Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: a review. Sci Total Environ 447:345–360
Schmittgen TD, Livak JK (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3:1101–1108
Seiler C, Berendonk TU (2012) Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Front Miciobiol 3:399
Sentchilo V, Mayer AP, Guy L, Miyazaki R, Tringe SG, Barry K et al (2013) Community-wide plasmid gene mobilization and selection. ISME J 7:1173–1186
Silveira E, Freitas AR, Antunes P, Barros M, Campos J, Coque TM, Peixe L, Novais C (2014) Co-transfer of resistance to high concentrations of copper and first-line antibiotics among Enterococcus from different origins (humans, animals, the environment and foods) and clonal lineages. J Antimicrob Chemother 69:899–906
Soffer N, Zaneveld J, Thurber RV (2015) Phage-bacteria network analysis and its implication for the understanding of coral disease. Environ Microbiol 17:1203–1218
Stepanauskas R, Glenn TC, Jagoe CH, Tuckfield RC, Lindell AH, King CJ, McArthur JV (2006) Coselection for microbial resistance to metals and antibiotics in freshwater microcosms. Environ Microbiol 8:1510–1514
Su JQ, Wei B, Ou-Yang WY, Huang FY, Zhao Y, Xu HJ, Zhu YG (2015) Antibiotic resistome and its association with bacterial communities during sewage sludge composting. Environ Sci Technol 49:7356–7363
Suzuki MT, Taylor LT, Delong EF (2000) Quantitative analysis of small-subunit rRNA genes in mixed microbial populations via 50-nuclease assays. Appl Environ Microbiol 66:4605–4614
Trajanovska S, Britz ML, Bhave M (1997) Detection of heavy metal ion resistance in Gram-positive and Gram-negative bacteria isolated from lead-contaminated site. Biodegradation 8:113–124
Wang R, Zhang J, Sui Q, Wan H, Tong J, Chen M, Wei Y, Wei D (2016) Effect of red mud addition on tetracycline and copper resistance genes and microbial community during the full scale swine manure composting. Bioresour Technol 216:1049–1057
Yin Y, Gu J, Wang X, Song W, Zhang K, Sun W, Zhang X, Zhang Y, Li H (2017) Effects of copper addition on copper resistance, antibiotic resistance genes, and intl1 during swine manure composting. Front Microbiol 8:344
Zhang YJ, Hu HW, Gou M, Wang JT, Chen D, He JZ (2017) Temporal succession of soil antibiotic resistance genes following application of swine, cattle and poultry manures spiked with or without antibiotics. Environ Pollut 231:1621–1632
Zhang ML, Chen LH, Ye CS, Yu X (2018) Co-selection of antibiotic resistance via copper shock loading on bacteria from a drinking water bio-filter. Environ Pollut 233:132–141
Zhu YG, Johnson TA, Su JQ, Qiao M, Guo GX, Stedtfeld RD, Hashsham SA, Tiedje JM (2013) Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proc Natl Acad Sci U S A 110:3435–3440
Zhu YG, Zhao Y, Li B, Huang CL, Zhang SY, Yu S, Chen YS, Zhang T, Gillings MR, Su JQ (2017) Continental-scale pollution of estuaries with antibiotic resistance genes. Nat Microbiol 2:16270
Funding
This study was supported by Australian Research Council (DP170103628) and Australian-China Joint Research Centre (ACSRF48165).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Responsible editor: Diane Purchase
Rights and permissions
About this article
Cite this article
Kang, W., Zhang, YJ., Shi, X. et al. Short-term copper exposure as a selection pressure for antibiotic resistance and metal resistance in an agricultural soil. Environ Sci Pollut Res 25, 29314–29324 (2018). https://doi.org/10.1007/s11356-018-2978-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-018-2978-y