Abstract
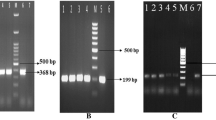
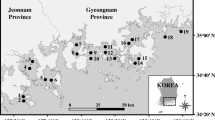
Vibrio parahaemolyticus is the most prevalent gastroenteritis-causing pathogen in Korea and in some other Asian countries. It is frequently found in oysters and other seafood. This study monitored changes in the prevalence of V. parahaemolyticus and environmental parameters in oyster aquaculture environments in Korea. From June to October 2014, we tested oysters (Crassostrea gigas) from shellfish-harvesting areas off the west coast of Korea. These 71 isolates were the sum of 16 (22.5 %), 19 (26.8 %), 23 (32.4 %), and 13 (18.3 %) isolates collected in July, August, September, and October, respectively. These 71 isolates had the following profiles of resistance against 16 antibiotics: all isolates were resistant to ampicillin and vancomycin, and 52.2, 50.7, and 50.7 % of isolates exhibited resistance to cephalothin, rifampin, and streptomycin, respectively. PCR analysis for the presence of the species-specific toxR gene confirmed that 38 (53.5 %) of the total 71 isolated strains were positive for V. parahaemolyticus. In PCR analysis for virulence of V. parahaemolyticus, of the 71 isolates tested in the present study, only 38 (53.5 %) were positive for the trh virulence gene and 71 (100 %) was negative for the tdh virulence gene.




Similar content being viewed by others
References
Akinbowale OL, Peng H, Batrton MD (2006) Antimicrobial resistance in bacteria isolated from aquaculture sources in Australia. J Appl Microbiol 100:1103–1113
Baquero F, Martinez JL, Canton R (2008) Antibiotics and antibiotic resistance in water environments. Curr Opin Biotechnol 19:260–265
Cook DW, O’Leary P, Hunsucker JC, Sloan EM, Bowers JC, Blodgett RJ, DePaola A (2002) Vibrio vulnificus and Vibrio parahaemolyticus in US retail shell oysters: a national survey from June 1998 to July 1999. J Food Prot 65:79–87
DePaola A, Ulaszek J, Kaysner CA, Tenge BJ, Nordstrom JL, Well J (2003) Molecular, serological, and virulence characteristics of Vibrio parahaemolyticus isolated from environmental, food, and clinical sources in North America and Asia. Appl Environ Microbiol 69:3999–4005
Dryselius R, Kurokawa K, Iida T (2007) Vibrionaceae, a versatile bacterial family with evolutionarily conserved variability. Res Microbiol 158(6):479–486
Ferrini AM, Mannoni V, Suffredini E, Cozzi L, Croci L (2008) Evaluation of antibacterial resistance in Vibrio strains isolated from imported seafood and Italian aquaculture settings. Food Anal Methods 1:164–170
Fuenzalida L, Armijo L, Zabala B, Hernández C, Rioseco ML, Riquelme C (2007) Vibrio parahaemolyticus strains isolated during investigation of the summer 2006 seafood related diarrhea outbreaks in two regions of Chile. Int J Food Microbiol 117:270–275
Gordon L, Giraud E, Ganiere JP, Armand F, Bouju-Albert A, de la Cotte N, Mangion C, Le Bris H (2007) Antimicrobial resistance survey in a river receiving effluents from freshwater fish farms. J Appl Microbiol 102:1167–1176
Han F, Walker RD, Janes ME, Prinyawiwatkul W, Ge B (2007) Antimicrobial susceptibilities of Vibrio parahaemolyticus and Vibrio vulnificus isolated from Louisiana Gulf and retail raw oysters. Appl Environ Microbiol 73:7096–7098
Han AR, Yoon YJ, Kim JW (2012) Antibiotic resistance and plasmid profile of Vibrio parahaemolyticus strains isolated from Kyunggi-Incheon coastal area. Kor J Microbiol 48:22–28
Honda T, Iida T (1993) The pathogenicity of Vibrio parahaemolyticus and the role of thermostable direct hemolysin and related hemolysins. Rev Med Microbiol 4:106–113
Jun JW, Kim JH, Choresca CH, Shin SP, Han JE, Han SY, Chai JY, Park SC (2012) Isolation, molecular characterization, and antibiotic susceptibility of Vibrio parahaemolyticus in Korean seafood. Foodborne Pathog Dis 9:224–231
KFDA (2010) Annual report of food-borne outbreaks. December 12, 2010, from http://fm.kfda.go.kr/
Kim SH, Sin YM, Lee MJ, Shin PK, Kim MG, Cho JS, Lee CH, Lee YJ, Chae KR (2005) Isolation of major foodborne pathogenic bacteria from ready-to-eat seafoods and its reduction strategy. J Life Sci 15:941–947
Krumperman PH (1983) Multiple antibiotic resistance indexing of Escherichia coli to identify high-risk sources of fecal contamination of foods. Appl Environ Microbiol 46:165–170
Lee JK, Jung DW, Eom SY, Oh SW, Kim Y, Kwak HS, Kim YH (2008) Occurrence of Vibrio parahaemolyticus in oysters from Korean retail outlets. Food Control 19:990–994
Lee HW, Lim SK, Kim MN (2009) Characteristics of ampicillin resistant Vibrio spp. isolated from a west coastal area of Korean Peninsula. Korean J Fish Aquat Sci 42:20–25
Mahmoud ZH, Kassu A, Mohammad A, Yamato M, Bhuiyan NA, Nair B (2006) Isolation and molecular characterization of toxigenic Vibrio parahaemolyticus from the Kii Channel, Japan. Microbiol Res 161:25–37
Martinez JL (2008) Antibiotics and antibiotic resistance genes in natural environments. Science 321:365–367
NCCLS (2003) Performance standards for antimicrobial disc susceptibility tests: approved standard, national committee for clinical laboratory standard of antimicrobial susceptibility, document M2-AS. Pennsylvania, USA
Nishibuchi M, Ishibashi M, Takeda Y, Kaper JB (1985) Detection of the thermostable direct hemolysin gene and related DNA sequences in Vibrio parahaemolyticus and other vibrio species by the DNA colony hybridization test. Am Soc Microbiol 49:481–486
Nordstrom JL, Vickery MC, Blackstone GM, Murray SL, DePaola A (2007) Development of a multiplex real-time PCR assay with an internal amplification control for the detection of total and pathogenic Vibrio parahaemolyticus bacteria in oysters. Appl Environ Microbiol 73:5840–5847
Oh EG, Son KT, Ha KS, Yoo HD, Yu HS, Shin SB, Lee HJ, Kim JH (2009) Antimicrobial resistance of Vibrio strains from brackish water on the coast of Gyeongsangnamdo. Korean J Fish Aquat Sci 42:335–343
Ottaviani D, Santarelli S, Bacchiocchi S, Masini L, Ghittino C, Bacchiocchi I (2005) Presence of pathogenic Vibrio parahaemolyticus strains in mussels from the Adriatic Sea, Italy. Food Microbiol 22:585–590
Panicker G, Call DR, Krug MJ, Bej AK (2004) Detection of pathogenic Vibrio spp. in shellfish by using multiplex PCR and DNA microarrays. Appl Environ Microbiol 70:7436–7444
Roque A, Lopez-Joven C, Lacuesta B, Elandaloussi L, Wagley S, Furones MD, Ruiz-Zarzuela I, Blas I, Rangdale R, Gomez-Gil B (2009) Detection and identification of tdh- and trh-positive Vibrio parahaemolyticus strains from four species of cultured bivalve molluscs on the Spanish Mediterranean Coast. Appl Environ Microbiol 75:7574–7577
Shimohata T, Takahashi A (2010) Diarrhea induced by infection of Vibrio parahaemolyticus. J Med Investig 57:179–182
Su YC, Liu C (2007) Vibrio parahaemolyticus: a concern of seafood safety. Food Microbiol 24:549–558
Sujeewa AK, Norrakiah AS, Laina M (2009) Prevalence of toxic genes of Vibrio parahaemolyticus in shrimps (Penaeus monodon) and culture environment. Int Food Res J 16:89–95
Tada J, Ohashi T, Nishimura N, Shirasaki Y, Ozaki H, Fukushima S, Takano J, Nishibuchi M, Takeda Y (1992) Detection of the thermostable direct hemolysin gene (tdh) and the thermostable direct hemolysin-related hemolysin gene (trh) of Vibrio parahaemolyticus by polymerase chain reaction. Mol Cell Probes 6:477–487
Vaseeharan B, Ramasamy P (2003) Abundance of potentially pathogenic micro-organisms in Penaeus monodon larvae rearing systems in India. Microbiol Res 158:299–308
Vaseeharan B, Ramasamy P, Murugan T, Chen JC (2005) In vitro susceptibility of antibiotics against Vibrio spp. and Aeromonas spp. Isolated from Penaeus monodon hatcheries and ponds. Int J Antimicrob Agents 26:285–291
Vongxay K, Wang S, Zhang X (2008) Pathogenetic characterization of Vibrio parahaemolyticus isolates from clinical and seafood sources. Int J Food Microbiol 126:71–75
Ward LN, Bej AK (2006) Detection of Vibrio parahaemolyticus in shellfish by use of multiplexed real-time PCR with TaqMan fluorescent probes. Appl Environ Microbiol 72:2031–2042
Wong MHY, Liu M, Wan HY, Chen S (2012) Characterization of extended spectrum–β lactamase–producing Vibrio parahaemolyticus. Am Soc Microbiol 56:4026–4028
Wright GD (2007) The antibiotic resistome: the nexus of chemical and genetic diversity. Nat Rev Microbiol 5:175–186
Yamamoto A, Iwahori J, Vuddhakul V, Charernjiratragul W, Vose D, Osaka K, Shigematsu M, Toyofuku H (2008) Quantitative modeling for risk assessment of Vibrio parahaemolyticus in bloody clams in southern Thailand. Int J Food Microbiol 124:70–78
Zhao F, Zhou D, Cao H, Ma L, Jiang Y (2011) Distribution, serological and molecular characterization of Vibrio parahaemolyticus from shellfish in the eastern coast of China. Food Control 22:1095–1100
Zulkifli Y, Alitheen NB, Son R, Yeap SK, Lesley MB, Raha AR (2009) Identification of Vibrio parahaemolyticus isolates by PCR targeted to the toxR gene and detection of virulence genes. Int Food Res J 16:289–296
Acknowledgments
This work was financially supported by the National Fisheries Research and Development Institute (R2015062).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Philippe Garrigues
Rights and permissions
About this article
Cite this article
Kang, CH., Shin, Y., Kim, W. et al. Prevalence and antimicrobial susceptibility of Vibrio parahaemolyticus isolated from oysters in Korea. Environ Sci Pollut Res 23, 918–926 (2016). https://doi.org/10.1007/s11356-015-5650-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-015-5650-9