Abstract
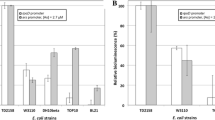
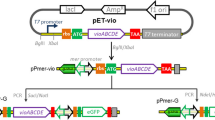
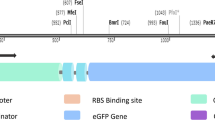
The detection of nickel in water is of great importance due to its harmfulness for living organism. A way to detect Ni is the use of whole-cell biosensors. The aim of the present work was to build a light-emitting bacterial biosensor for the detection of Ni with high specificity and low detection limit properties. For that purpose, the regulatory circuit implemented relied on the RcnR Ni/Co metallo-regulator and its rcnA natural target promoter fused to the lux reporter genes. To convert RcnR to specifically detect Ni, several mutations were tested and the C35A retained. Deleting the Ni efflux pump rcnA and introducing genes encoding several Ni-uptake systems lowered the detection thresholds. When these constructs were assayed in several Escherichia coli strains, it appeared that the detection thresholds were highly variable. The TD2158 wild-type E. coli gave rise to a biosensor ten times more active and sensitive than its W3110 E. coli K12 equivalent. This biosensor was able to confidently detect Ni concentrations as little as 80 nM (4.7 μg l−1), which makes its use compatible with the norms governing the drinking water quality.







Similar content being viewed by others
References
Bereza-Malcolm LT, Mann G, Franks AE (2014) Environmental sensing of heavy metals through whole cell microbial biosensors: a synthetic biology approach. ACS Synth Biol
Blaha D, Arous S, Bleriot C, Dorel C, Mandrand-Berthelot MA, Rodrigue A (2011) The Escherichia coli metallo-regulator RcnR represses rcnA and rcnR transcription through binding on a shared operator site: insights into regulatory specificity towards nickel and cobalt. Biochimie 93:434–439
Bleriot C, Effantin G, Lagarde F, Mandrand-Berthelot MA, Rodrigue A (2011) RcnB is a periplasmic protein essential for maintaining intracellular Ni and Co concentrations in Escherichia coli. J Bacteriol 193:3785–3793
Brutesco C et al. (2015) Influence of the genetic background and reporter in optimal light response of bacterial biosensors. Environ Sci Pollut Res Int. (in this issue)
Condee CW, Summers AO (1992) A mer-lux transcriptional fusion for real-time examination of in vivo gene expression kinetics and promoter response to altered superhelicity. J Bacteriol 174:8094–8101
Contreras M, Thiberge JM, Mandrand-Berthelot MA, Labigne A (2003) Characterization of the roles of NikR, a nickel-responsive pleiotropic autoregulator of Helicobacter pylori. Mol Microbiol 49:947–963
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci U S A 97:6640–6645
De Pina K, Desjardin V, Mandrand-Berthelot MA, Giordano G, Wu LF (1999) Isolation and characterization of the nikR gene encoding a nickel-responsive regulator in Escherichia coli. J Bacteriol 181:670–674
Dhillon TS, Poon AP, Chan D, Clark AJ (1998) General transducing phages like Salmonella phage P22 isolated using a smooth strain of Escherichia coli as host. FEMS Microbiol Lett 161:129–133
Duprey A et al (2014) “NiCo Buster”: engineering E. coli for fast and efficient capture of cobalt and nickel. J Biol Eng 8:19
Eitinger T, Mandrand-Berthelot MA (2000) Nickel transport systems in microorganisms. Arch Microbiol 173:1–9
Eitinger T, Rodionov DA, Grote M, Schneider E (2010) Canonical and ECF-type ATP-binding cassette importers in prokaryotes: diversity in modular organization and cellular functions. FEMS Microbiol Rev 35:3–67
European Drinking Water Directive (1998) Council Directive 98/83/EC. http://eur-lex.europa.eu/legal-content/EN/ALL/?uri=CELEX:31998L0083
Grass G, Fricke B, Nies DH (2005) Control of expression of a periplasmic nickel efflux pump by periplasmic nickel concentrations. Biometals 18:437–448
Iwig JS, Rowe JL, Chivers PT (2006) Nickel homeostasis in Escherichia coli—the rcnR–rcnA efflux pathway and its linkage to NikR function. Mol Microbiol 62:252–262
Iwig JS, Leitch S, Herbst RW, Maroney MJ, Chivers PT (2008) Ni(II) and Co(II) sensing by Escherichia coli RcnR. J Am Chem Soc 130:7592–7606
Jubier-Maurin V, Rodrigue A, Ouahrani-Bettache S, Layssac M, Mandrand-Berthelot MA, Kohler S, Liautard JP (2001) Identification of the nik gene cluster of Brucella suis: regulation and contribution to urease activity. J Bacteriol 183:426–434
Link AJ, Phillips D, Church GM (1997) Methods for generating precise deletions and insertions in the genome of wild-type Escherichia coli: application to open reading frame characterization. J Bacteriol 179:6228–6237
Liu T et al (2007) CsoR is a novel Mycobacterium tuberculosis copper-sensing transcriptional regulator. Nat Chem Biol 3:60–68
Lopez-Maury L, Garcia-Dominguez M, Florencio FJ, Reyes JC (2002) A two-component signal transduction system involved in nickel sensing in the cyanobacterium Synechocystis sp. PCC 6803. Mol Microbiol 43:247–256
Ma Z, Jacobsen FE, Giedroc DP (2009) Coordination chemistry of bacterial metal transport and sensing. Chem Rev 109:4644–4681
Miller JH (1992) Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor
Pal C, Bengtsson-Palme J, Rensing C, Kristiansson E, Larsson DG (2014) BacMet: antibacterial biocide and metal resistance genes database. Nucleic Acids Res 42:D737–D743
Peca L, Kos PB, Mate Z, Farsang A, Vass I (2008) Construction of bioluminescent cyanobacterial reporter strains for detection of nickel, cobalt and zinc. FEMS Microbiol Lett 289:258–264
Prévéral S, Brutesco C, Descamps E, Escoffier C, Ginet N, Pignol D, Garcia D. (2015) A bioluminescent arsenite biosensor designed for inline water analyzer. Environ Sci Pollut Res Int. (in this issue)
Rodrigue A, Effantin G, Mandrand-Berthelot MA (2005) Identification of rcnA (yohM), a nickel and cobalt resistance gene in Escherichia coli. J Bacteriol 187:2912–2916
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor
Sebbane F, Mandrand-Berthelot MA, Simonet M (2002) Genes encoding specific nickel transport systems flank the chromosomal urease locus of pathogenic yersiniae. J Bacteriol 184:5706–5713
Tibazarwa C et al (2001) A microbial biosensor to predict bioavailable nickel in soil and its transfer to plants. Environ Pollut 113:19–26
van der Meer JR, Belkin S (2010) Where microbiology meets microengineering: design and applications of reporter bacteria. Nat Rev Microbiol 8:511–522
Winson MK et al (1998) Engineering the luxCDABE genes from Photorhabdus luminescens to provide a bioluminescent reporter for constitutive and promoter probe plasmids and mini-Tn5 constructs. FEMS Microbiol Lett 163:193–202
World Health Organization (2011) Guidelines for drinking-water quality. http://www.who.int/water_sanitation_health/publications/2011/dwq_guidelines/en/
Wu LF, Navarro C, Mandrand-Berthelot MA (1991) The hydC region contains a multi-cistronic operon (nik) involved in nickel transport in Escherichia coli. Gene 107:37–42
Acknowledgments
This work was supported by the French National Agency ANR Ecotech, COMBITOX.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Philippe Garrigues
Julien Cayron and Elsa Prudent contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 422 kb)
Rights and permissions
About this article
Cite this article
Cayron, J., Prudent, E., Escoffier, C. et al. Pushing the limits of nickel detection to nanomolar range using a set of engineered bioluminescent Escherichia coli . Environ Sci Pollut Res 24, 4–14 (2017). https://doi.org/10.1007/s11356-015-5580-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-015-5580-6